Mouse Development
Introduction
The mouse (taxon-mus) has always been a good embryological model, generating easily (litters 8-20) and quickly (21d). Mouse embryology really expanded when molecular biologists used mice for gene knockouts. Suddenly it was necessary to understand development in order to understand the effect of knocking out the gene. There are over 450 different strains of inbred research mice, and these strains have recently been organized into a chart. While being an ideal model organism, only a relatively small amount (1.5%) of the total mouse genome has been sequenced. Those interested in the mouse reproductive cycle should also look at the mouse estrous cycle.
There are several systems for staging mouse development. The original and most widely used is the Theiler Stages system, which divides mouse development into 26 prenatal and 2 postnatal stages. [1]
- Mouse Stages: E1 | E2.5 | E3.0 | E3.5 | E4.5 | E5.0 | E5.5 | E6.0 | E7.0 | E7.5 | E8.0 | E8.5 | E9.0 | E9.5 | E10 | E10.5 | E11 | E11.5 | E12 | E12.5 | E13 | E13.5 | E14 | E14.5 | E15 | E15.5 | E16 | E16.5 | E17 | E17.5 | E18 | E18.5 | E19 | E20 | Timeline | About timed pregnancy
| Carnegie | Stage | |||||||||||||||||||||||
| Human | Days | 1 | 2-3 | 4-5 | 5-6 | 7-12 | 13-15 | 15-17 | 17-19 | 20 | 22 | 24 | 28 | 30 | 33 | 36 | 40 | 42 | 44 | 48 | 52 | 54 | 55 | 58 |
| Mouse | Days | 1 | 2 | 3 | E4.5 | E5.0 | E6.0 | E7.0 | E8.0 | E9.0 | E9.5 | E10 | E10.5 | E11 | E11.5 | E12 | E12.5 | E13 | E13.5 | E14 | E14.5 | E15 | E15.5 | E16 |
| Rat | Days | 1 | 3.5 | 4-5 | 5 | 6 | 7.5 | 8.5 | 9 | 10.5 | 11 | 11.5 | 12 | 12.5 | 13 | 13.5 | 14 | 14.5 | 15 | 15.5 | 16 | 16.5 | 17 | 17.5 |
| Note these Carnegie stages are only approximate day timings for average of embryos. Links: Carnegie Stage Comparison | ||||||||||||||||||||||||
| ||||||||||||||||||||||||
| Timeline Links: human timeline | mouse timeline | mouse detailed timeline | chicken timeline | rat timeline | Medaka | Category:Timeline |
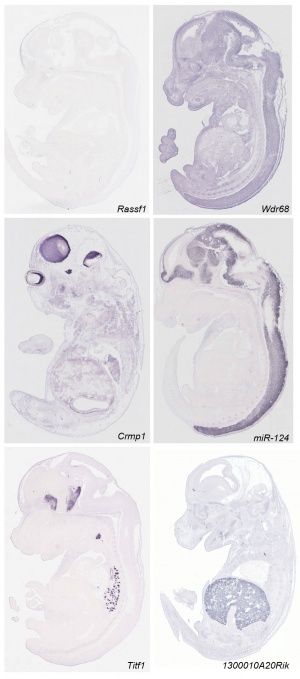
Some Recent Findings
|
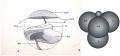
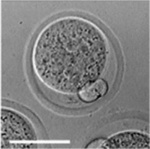
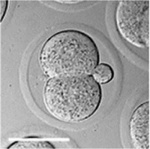
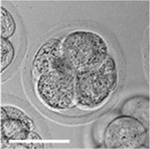
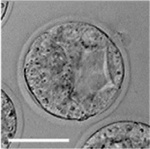
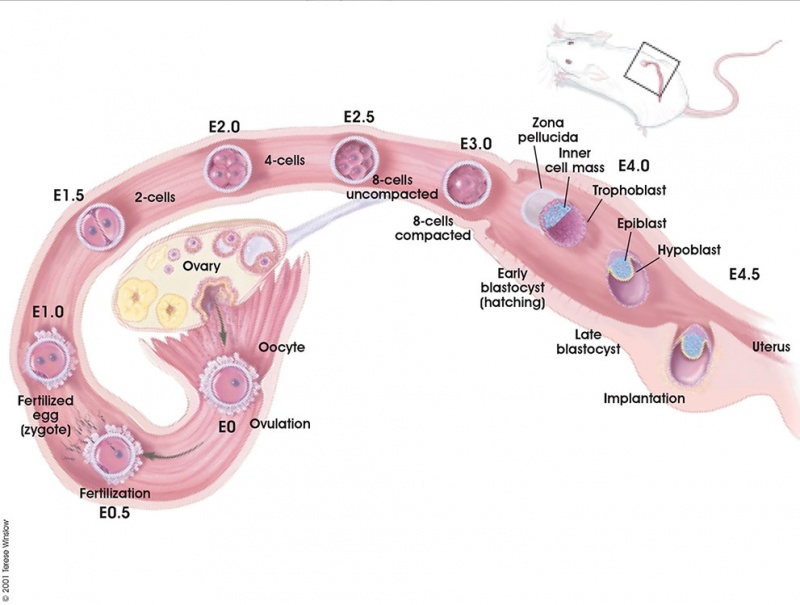
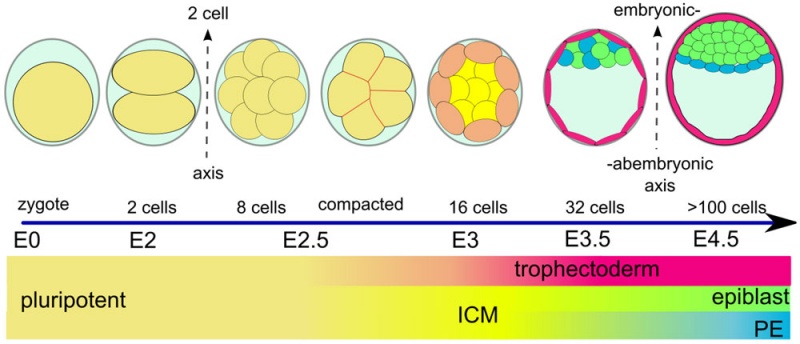
Early Mouse Development
|
|
|
|
|
| Mouse 1 cell | Mouse 2 cell | Mouse morula | Mouse early blastocyst | Mouse hatched blastocyst |
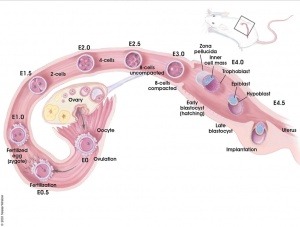
Early mouse development model[5]
|
|
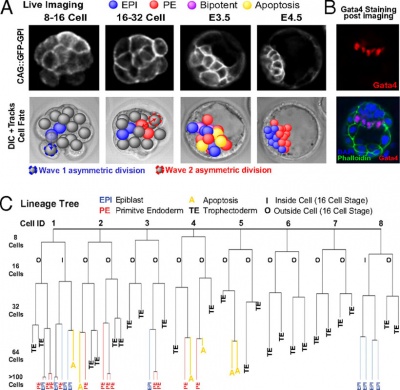
Live cell imaging and tracking[6]
|
Simulation of Zygote to Blastocyst[5]
|
- Links: Mouse Stages
Later Mouse Development
Mouse E8.5[7]
Mouse E9.5[8]
Mouse E10.5[8]
Mouse E11.5[8]
Mouse E11.5[8]
Mouse E12.5[8]
Mouse E14.5[9]
- Links: Mouse Stages | Mouse Timeline Detailed
Carnegie Stages Comparison
The table below gives an approximate comparison of human, mouse and rat embryos based upon Carnegie staging.
| Carnegie | Stage | |||||||||||||||||||||||
| Human | Days | 1 | 2-3 | 4-5 | 5-6 | 7-12 | 13-15 | 15-17 | 17-19 | 20 | 22 | 24 | 28 | 30 | 33 | 36 | 40 | 42 | 44 | 48 | 52 | 54 | 55 | 58 |
| Mouse | Days | 1 | 2 | 3 | E4.5 | E5.0 | E6.0 | E7.0 | E8.0 | E9.0 | E9.5 | E10 | E10.5 | E11 | E11.5 | E12 | E12.5 | E13 | E13.5 | E14 | E14.5 | E15 | E15.5 | E16 |
| Rat | Days | 1 | 3.5 | 4-5 | 5 | 6 | 7.5 | 8.5 | 9 | 10.5 | 11 | 11.5 | 12 | 12.5 | 13 | 13.5 | 14 | 14.5 | 15 | 15.5 | 16 | 16.5 | 17 | 17.5 |
| Note these Carnegie stages are only approximate day timings for average of embryos. Links: Carnegie Stage Comparison | ||||||||||||||||||||||||
| ||||||||||||||||||||||||
Spermatozoa Development
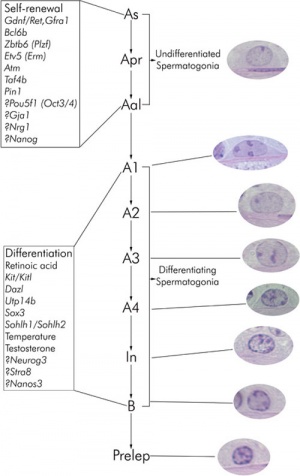
The process of spermatogenesis takes approximately 35 days:
- mitotic phase (11 days)
- meiotic phase (10 days)
- post-meiotic phase (14 days)
Spermatogonial stem cells (SSCs)
The diploid germ cells, spermatogonial stem cells (SSCs), are located on the basement membrane of the seminiferous tubules
- adult mouse testis about 30,000 SSCs
- either divides into two new single cells
- or into a pair of spermatogonia (Apr)
- that do not complete cytokinesis and stay connected by an intercellular bridge
Primitive spermatogonia subset
- Asingle (As, single isolated spermatogonia)
- Apaired (Apr, interconnected spermatogonial pairs)
- Aaligned (Aal, interconnected 4, 8, or 16 spermatogonia)
- specifically termed Aal-4, Aal-8, and Aal-16
Primitive spermatogonia cells transform without cell division into more differentiating A1 spermatogonia that undergo 6 mitotic and 2 meiotic divisions to eventually form haploid spermatids.
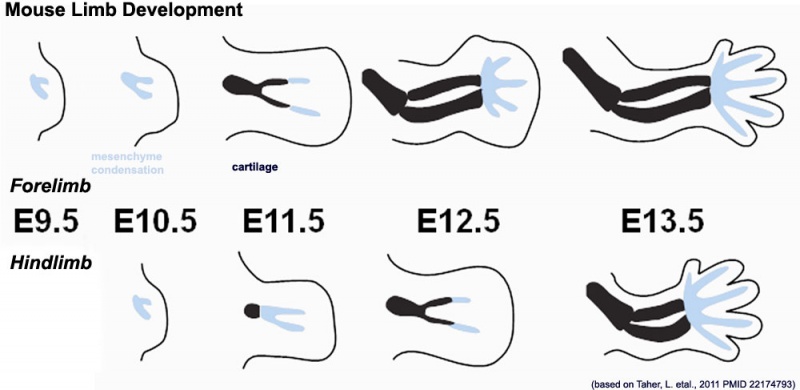
Limb Development
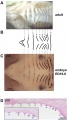
Mouse limb skeleton cartoon[11]
Fore-limb and hind-limb buds for stages E9.5 to E13.5. Hindlimbs are morphologically delayed by about half a day.
- Light blue - indicate mesenchymal condensations.
- Thick black lines - indicate cartilage as determined by alcian blue staining.
Change in cell types and tissue formation as a function of mouse developmental stage.[11]
- Links: Limb Development
Neural Development
The data below is summarised from a study of early neural development in the mouse.[12]
- initial fusion of apposing neural folds occurred at the level of the intermediate point between the third and fourth somites (caudal myelencephalon) both rostrally and caudally
- second fusion - at the original rostral end of the neural plate (rostrodorsally).
- third fusion - in the caudal diencephalon (rostrally and caudally)
- followed by complete closure of the telencephalic neuropore at the midpoint of the telencephalic roof
- then complete closure of the metencephalic neuropore at the rostral part of the metencephalic roof
- fourth fusion - at the original caudal end of the neural plate (rostrally)
- caudal neuropore completely closed at the level of the future 33rd somite
See also these 1980's papers.[13] [14]
Urogenital Development
A high-resolution description of the developing murine genitourinary tract from Theiler stage (TS) 17 (E10.5) through to TS27 (E19.5) and then to postnatal day 3 was published in 2007.[15]
The GenitoUrinary Development Molecular Anatomy Project (GUDMAP) is a consortium of laboratories working to provide the scientific and medical community with tools to facilitate research. They intend to develop:
- a molecular atlas of gene expression for the developing organs of the GenitoUrinary (GU) tract
- a high resolution molecular anatomy that highlights development of the GU system
- mouse strains to facilitate developmental and functional studies within the GU system
- tutorials describing GU organogenesis
- rapid access to primary data via the GUDMAP database
Mouse Knockouts
Knowledge about mouse development has rapidly expanded as it has become the model animal system for genetic "knock out " studies. This technology actually requires development of defined breeding programs, pseudo-pregnancy, in vitro fertilization, molecular biology, and good old fashioned histology. Without understanding normal development the molecular biologists don't stand a hope of understanding what their gene knock out has done. There is a database of all existing mouse knockouts and their consequences.
Murine Development Control Genes
Kessel, M. and Gruss, P. Science 249 374-379 (1990)
An early review of the genes, and method of identifying them, involved in early mouse development. In particular discusses Homeobox genes. (homeobox is 183bp encoding a 61 amino acid DNA-binding domain)
- Gene families
- Hox
- Pax
- POU
The Genealogy Chart of Inbred Strains
This chart shows the origins and relationships of inbred mouse strains. The chart is available as a PDF document [../pdf/mouse_genealogy.pdf Locally] or from JAX Labs and was originally published by Beck etal., 2000.
- Beck JA, Lloyd S, Hafezparast M, Lennon-Pierce M, Eppig JT, Festing MF, Fisher EM. Genealogies of mouse inbred strains. Nat Genet. 2000 Jan;24(1):23-5.
Mouse Genome
Mouse Genome completed December 2002, a draft sequence and analysis of the genome of the C57BL/6J mouse strain.
- less than 30,000 genes
- estimated size is 2.5 Gb, smaller than the human genome
- about 40% of the human and mouse genomes can be directly aligned
- about 80% of human genes have one corresponding gene in the mouse genome
- Links: Mouse Genome Sequencing: Mus musculus | Mouse Genome Informatics | Mouse Genome Project | Nature - Mouse Genome
References
- ↑ The House Mouse: Atlas of Mouse Development by Theiler Springer-Verlag, NY (1972, 1989). | online book
- ↑ 2.0 2.1 <pubmed>21267068</pubmed>| PLoS Biol. | Eurexpress transcriptome atlas
- ↑ <pubmed>22147950</pubmed>
- ↑ <pubmed>21677750</pubmed>
- ↑ 5.0 5.1 <pubmed>21573197</pubmed>| PMC3088645 | PLoS Comput Biol.
- ↑ <pubmed>20308546</pubmed>| PMC2852013 | PNAS
- ↑ <pubmed>20704721</pubmed>| BMC Dev Biol.
- ↑ 8.0 8.1 8.2 8.3 8.4 <pubmed>16683035</pubmed>| PLoS Genetics
- ↑ <pubmed>18713865</pubmed>| PNAS
- ↑ Zhou Q, Griswold MD. Regulation of spermatogonia. StemBook [Internet]. Cambridge (MA): Harvard Stem Cell Institute. PMID20614596 | http://www.ncbi.nlm.nih.gov/books/NBK27035/ Bookshelf]
- ↑ 11.0 11.1 <pubmed>22174793</pubmed>
- ↑ Neurulation in the mouse: manner and timing of neural tube closure. Sakai Y. Anat Rec. 1989 Feb;223(2):194-203. PMID: 2712345
- ↑ The histogenetic potential of neural plate cells of early-somite-stage mouse embryos. Chan WY, Tam PP. J Embryol Exp Morphol. 1986 Jul;96:183-93. PMID: 3805982
- ↑ Neurulation in the mouse. I. The ontogenesis of neural segments and the determination of topographical regions in a central nervous system. Sakai Y. Anat Rec. 1987 Aug;218(4):450-7. PMID: 3662046
- ↑ <pubmed>17452023</pubmed>| PMC2117077
Search Pubmed
Search Pubmed: Mouse Development | Mouse Embryology
Additional Images
Movies
| Fertilization mouse |
Parental genome mouse |
Nodal cilia rotation mouse |
Somitogenesis mouse |
| Migration 1 | Migration 2 | Migration 3 | microCT E11.5 |
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
Mouse Embryology
- Manipulating the Mouse Embryo (Cold Spring Harbor Laboratory)
- The e-Mouse Atlas Project EMAP
Mouse Gene Expression
- EMAGE is a database of in situ gene expression data in the mouse embryo and an accompanying suite of tools to search and analyse the data. (All site content, except where otherwise noted, is licensed under a Creative Commons Attribution License.)
- Eurexpress transcriptome atlas a High-Resolution Anatomical Atlas of the Transcriptome in the Mouse Embryo.
- GenePaint.org is a digital atlas of gene expression patterns in the mouse.
Mouse Genome
- NCBI- Mouse Genome Sequencing
- Mouse Genome Informatics
- EBI- Mouse Genome Sequencing
- ANEX Rat and Mouse Database - (University of Tokushima)
- Genetic and Physical Maps of the Mouse Genome (Whitehead Institute/MIT)
- Genetic Location Database (University of Southampton)
- Koop Human and Mouse Genome Project Research (University of Victoria)
- Genome Systems, Inc.
Mouse Jackson Laboratory
- Jackson Laboratory
- Jackson Laboratory Induced Mutant Resource
- Jackson Laboratory BioInformatics Electronic Bulletin Board System
- Mouse Inbred Strains (Jackson Labs)
- Mouse Linkage Map
- Mouse Nomenclature Rules and Guidelines (Jackson Labs)
Mouse Diseases
- Helicobacter hepaticus, a Recently Recognized Bacterial Pathogen, Associated with Chronic Hepatitis and Hepatocellular Neoplasia in Laboratory Mice (CDC)
- Helicobacter Infection in Laboratory Mice: History, Significance, Detection and Management (Charles River)
- Infectious Diseases of Mice and Rats (NAS-ILAR)
- Infectious Diseases of Mice and Rats Companion Guide (NAS-ILAR)
- Infectious Diseases of Mice and Rats (AFIP-POLA)
- Murine Helicobacters (NCI)
Mouse Transgenics
- Archives of Transgenic-List
- Mammary Transgene Database (Baylor College of Medicine)
- Targeted Mutagenesis in Mice: A Video Guide
- TBASE - Transgenic/Targeted Mutation Database
- B Universal Ltd.
- BIG BLUE (Big Blue Transgenic Mouse Mailing List Archives)
- Big Blue lacI Transgenic Mouse Page
- Charles River Laboratories, Inc.
- Chrysalis DNX Transgenic Services
Mouse Transgenic Facilities
- Transgenic Animal Facility (University of Connecticut Biotechnology Center)
- Transgenic Animal Facility (University of Illinois Biotechnology Center)
- Transgenic Animal Facility (University of Iowa)
- Transgenic Animal Facility (Karolinska Institute)
- Transgenic Animal Facility (University of Western Australia)
- Transgenic Animal Facility (University of Wisconsin Biotechnology Center)
- Transgenic Animal Model Core (University of Michigan)
- Transgenic Core (Baylor College of Medicine SPORE)
- Transgenic Facility (University of Colorado Health Science Center)
- Transgenic Facility (University of Kentucky)
- Transgenic & Knockout Mouse Facility (University of Maryland at Baltimore)
- Transgenic-List Archives
- Transgenic Models of Skin Disease Core (Harvard Skin Disease Research Center)
- Transgenic Mouse Core Facilities (Mount Sinai School of Medicine)
- Transgenic Mouse Core Facility (University of Virginia)
- Transgenic Mouse Facility (Columbia-Presbyterian Cancer Center)
- Transgenic Mouse Facility (Duke Comprehensive Cancer Center)
- Transgenic Mouse Facility (SUNY-Stony Brook)
- Transgenic Mouse Facility (University of California-Irvine)
- Transgenic Mouse Facility (University of South Carolina)
- Transgenic Mouse Facility (Yale University Cancer Center)
- Transgenic Mouse Program (University of California-Davis)
- Transgenic Mouse Research Facility (New York University Kaplan Comprehensive Cancer Center)
- Transgenics (Berlin-Buch GmbH)
Mouse Urogenital
- GenitoUrinary Development Molecular Anatomy Project (GUDMAP) Renal Development Tutorial | Genital Development Tutorial
Mouse Unsorted Links
- Covance Research Products
- Edison Biotechnology Institute (Ohio University)
- Encyclopedia of the Mouse Genome (Jackson Laboratory)
- EUCIB Mouse Backcross Database (MBx)
- Harlan Sprague Dawley, Inc.
- It's a Knockout
- Jackson Laboratory Gopher Server
- Lane List of Named Mutations and Polymorphic Loci
- Lexicon Genetics, Inc.
- Lymphocytic Choriomeningitis Virus: An Unrecognized Teratogenic Pathogen (CDC)
- M & B Breeding and Research Centre Ltd.
- Map Manager Software
- Mice and Men: Making the Most of Our Similarities (ORNL)
- Microinjection Workshop
- Mouse Atlas and Gene Expression Database Project
- Mouse Cytogenetic Map Image
- Mouse Idiograms
- Mouse Linkage Map
- Mouse and Rat Research Home Page
- MouseUp HyperCard Transgenic Mouse Database (Macintosh)
- MRC Mouse Genome Centre
- National Institute of Genetics Mouse Genetic Resources (Japan)
- NIH Animal Genetic Resource
- NIH Directory of Transgenic Mice
- NIH Mouse Club
- Of Mice and Men (HHMI's Blazing a Genetic Trail)
- Online Mendelian Inheritance in Animals (University of Sydney)
- Pathology of Genetically-altered Mice
- Portable Dictionary of the Mouse Genome
- Review of Pasteurella pneumotropica (Charles River)
- Taconic
- The Mouse in Science: Cancer Research
- The Mouse in Science: Monoclonal Antibodies
- The Mouse in Science: Vaccines
- The Mouse in Science: Why Mice?
| Animal Development: axolotl | bat | cat | chicken | cow | dog | dolphin | echidna | fly | frog | goat | grasshopper | guinea pig | hamster | horse | kangaroo | koala | lizard | medaka | mouse | opossum | pig | platypus | rabbit | rat | salamander | sea squirt | sea urchin | sheep | worm | zebrafish | life cycles | development timetable | development models | K12 |
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, June 18) Embryology Mouse Development. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Mouse_Development
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G

![Mouse E8.5[7]](/embryology/images/thumb/9/9f/Mouse-E8.5_dorsal_view.jpg/120px-Mouse-E8.5_dorsal_view.jpg)
![Mouse E9.5[8]](/embryology/images/thumb/b/b2/Mouse-E9.5.jpg/90px-Mouse-E9.5.jpg)
![Mouse E10.5[8]](/embryology/images/thumb/8/89/Mouse_CT_E10.5.jpg/90px-Mouse_CT_E10.5.jpg)
![Mouse E11.5[8]](/embryology/images/thumb/b/b2/Mouse_CT_E11.5.jpg/90px-Mouse_CT_E11.5.jpg)
![Mouse E12.5[8]](/embryology/images/thumb/d/da/Mouse_CT_E12.5.jpg/90px-Mouse_CT_E12.5.jpg)
![Mouse E14.5[9]](/embryology/images/thumb/f/fe/Mouse_CT_E14.5.jpg/77px-Mouse_CT_E14.5.jpg)