Molecular Development: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 190: | Line 190: | ||
:'''Links:''' [[Molecular_Development_-_Epigenetics|Epigenetics]] | :'''Links:''' [[Molecular_Development_-_Epigenetics|Epigenetics]] | ||
| [[File:Epigenetics cartoon.jpg| | | [[File:Epigenetics cartoon.jpg|400px|Epigenetics mechanisms]] | ||
|} | |} | ||
Revision as of 11:35, 15 July 2016
| Embryology - 21 Jun 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
This page is a link to many different resources related to molecular development. I am 2.85 billion nucleotides of DNA, but so is a chimpanzee, and all this DNA encodes only about 20,000-25,000 protein-coding genes. In development, I am not that different from a mouse or a fly and many of the signals that regulate development are used time and time again.
We have come a long way from just observing development to now wanting to understand how the complex program of development is controlled. Using new research tools and some excellent animal models researchers have discovered common themes and mechanisms that tie all embryonic development together.
What is remarkable, given our biological diversity, is the strong evolutionary conservation of developmental mechanisms. This has been a boon in allowing the use of many (easier) model systems such as the genetist's tool the fruitfly, and the worm, frog, chicken, zebrafish and mouse (see Animal Development page).
A continuing theme also seems to be the reuse of signals at different times and places within the embryo, for diiferent jobs. This has given rise to the concept of "switches" which by themselves may contain no "information" but to activate other genes or switches. Finally, you can imagine that of our 20,000-25,000 protein-coding genes, a large number of these may only be expressed during development or if reused, have a completely different role in the mature animal.
In terms of molecular mechanisms, the field of epigenetics has begun to florish with some recent important findings.
Molecular mechanisms of development is an exciting area and requires a variety of different skills. This page introduces only a few examples and should give you a feel for the topic. Note that each section of system notes has a page covering molecular development in that system. I have included information about the basic building blocks of proteins (amino acids).
| Hox Genes | |
|---|---|
|

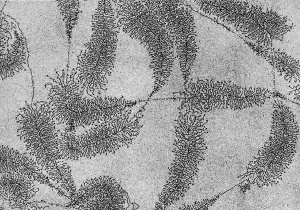
|
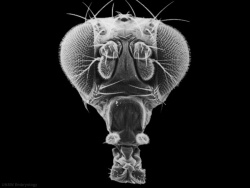
| Fly wild-type head[2] | Fly antennapedia mutant head[2] |
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
| Mechanism Links: mitosis | cell migration | cell junctions |epithelial invagination | epithelial mesenchymal transition | mesenchymal epithelial transition | epithelial mesenchymal interaction | morphodynamics | tube formation | apoptosis | autophagy | axes formation | time | molecular |
Some Recent Findings
|
Starting Out
There are several different ways to begin to look at molecular development.
- look at the earliest molecular events involved in patterning the differentiation of cells- axis formation.
- or look at early events in patterning with differentiation and migration of cells to form the trilaminar embryo (gastrulation).
- or look at the signaling mechanisms and the different types of factors involved.
- or look at their role in the development of specific systems.
Signaling during development, though complex, can also be grouped into a few specific classes. These mechanisms have also been listed and described briefly on Signaling Mechanisms page.
For this section, Molecular Biology of the Cell (now indexed and available online from PubMed) and the extracted Cell Biology version are good reference texts. Also the key journals in this area are: Development, Genes and Development, PNAS, EMBOJ.
There is also detailed molecular information available from the abnormalities pages where there are links to the OMIM Database entry for specific genetic disorders.
Gene Expression
DNA -> RNA -> Protein
Our genetic information is stored within each cell in cell organelles of the nucleus and mitochondria.
- In the nucleus as chromosomes.
- In the mitochondria as circular DNA.
Gene expression can often simply be described by the pathway: DNA -> mRNA -> Protein. While each cell contains the same genetic material, not all genes are ever expressed and the subset of genes expressed at any one time affects cellular differentiation and mature cell function. This can be impacted by both signaling and epigenetic mechanisms.
The human chromosomes consist of proteins and nucleic acids, the common nucleic acids or bases that forms DNA can be represented by a standardised IUPAC single letter code.
Deoxyribonucleic acid
Deoxyribonucleic acid (DNA) consists of 4 nucleotides (bases):
- Adenine (A)
- Cytosine (C)
- Guanine (G)
- Thymine (T)
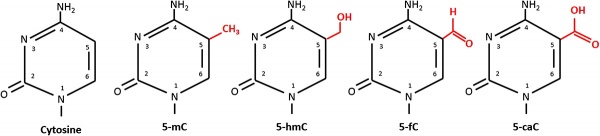
An additional amino acid, 5-Formylcytosine (5fC), a rare nucleotide has recently been identified as a stable DNA modification occurring in mammals.[5] The function of this additional base was thought to be involved only in active DNA demethylation, but may also have roles in gene regulation.
Ribonucleic acid
Ribonucleic acid (RNA) consists of 4 nucleotides:
- Adenine (A)
- Cytosine (C)
- Guanine (G)
- Uracil (U)
The original "RNA family" consisted of just 3 main members; transfer RNA (tRNA), ribosomal RNA (rRNA) and messenger RNAs (mRNA). Involved in gene expression through protein translation (synthesis).
In recent years this RNA family, shown in the table below, has been expanded to include newly identified members: small nuclear RNA (snRNA) , small nucleolar RNA (snoRNA), and short regulatory RNA (piwi-associated RNA (piRNA), endogenous short-interfering RNA (endo-siRNA) and microRNA (miRNA) and now long non-coding RNA (lncRNA). Each of these new family members has a range of potential roles in development and differentiation.
| RNA Class | Acronym | Roles | Examples | Reviews |
|---|---|---|---|---|
| transfer RNA | tRNA | carry amino acid in cell cytoplasm to ribosome | PMID 24966867 | |
| messenger RNA | mRNA | transcribed from DNA in cell nucleus and relocate to cytoplasm for translation on the ribosome | Example | |
| ribosomal RNA | rRNA | structural RNA that allow the assembly of ribosomal proteins in the cytoplasm into ribosomal subunits required for protein translation | Example | |
| small nuclear RNA | snRNA | involved in cell nucleus RNA splicing | PMID 23980890 | |
| small nucleolar RNA | snoRNA | modify other small RNAs (rRNAs and tRNAs) | box C/D snoRNAs and the box H/ACA | PMID 21664409 PMID 19446021 |
| microRNA | miRNA | post-transcriptional regulator of gene expression | PMID 25128264 | |
| endogenous short-interfering RNA | endo-siRNA | short regulatory RNA | mainly characterised in plants | PMID 22578318 |
| piwi-associated RNA | piRNA | short regulatory RNA | PMID 18032451 PMID 22103557 | |
| long non-coding RNA | ncRNA | non-coding RNA greater than 200bp in length may have different roles in signalling, protein processing and differentiation | PMID 24829860 |
Protein
The individual amino acids that form all proteins can be represented by a standardised single letter code, or three letter code or by their entire name.
Single Letter Code
A - Alanine (Ala) | C - Cysteine (Cys) | D - Aspartic Acid (Asp) | E - Glutamic Acid (Glu) | F - Phenylalanine (Phe) | G - Glycine (Gly) | H - Histidine (His) | I - Isoleucine (Ile) | K - Lysine (Lys) |L - Leucine (Leu) | M - Methionine (Met) | N - Asparagine (Asn) | P - Proline (Pro) | Q - Glutamine (Gln) | R - Arginine (Arg) | S - Serine (Ser) | T - Threonine (Thr) | V - Valine (Val) | W - Tryptophan (Trp) | Y - Tyrosine (Tyr)
| Amino Acid | 3-Letter | 1-Letter | Side chain polarity | Side chain acidity or basicity of neutral species | Hydropathy index |
|---|---|---|---|---|---|
| Alanine | Ala | A | nonpolar | neutral | 1.8 |
| Arginine | Arg | R | polar | basic (strongly) | -4.5 |
| Asparagine | Asn | N | polar | neutral | -3.5 |
| Aspartic acid | Asp | D | polar | acidic | -3.5 |
| Cysteine | Cys | C | polar | neutral | 2.5 |
| Glutamic acid | Glu | E | polar | acidic | -3.5 |
| Glutamine | Gln | Q | polar | neutral | -3.5 |
| Glycine | Gly | G | nonpolar | neutral | -0.4 |
| Histidine | His | H | polar | basic (weakly) | -3.2 |
| Isoleucine | Ile | I | nonpolar | neutral | 4.5 |
| Leucine | Leu | L | nonpolar | neutral | 3.8 |
| Lysine | Lys | K | polar | basic | -3.9 |
| Methionine | Met | M | nonpolar | neutral | 1.9 |
| Phenylalanine | Phe | F | nonpolar | neutral | 2.8 |
| Proline | Pro | P | nonpolar | neutral | -1.6 |
| Serine | Ser | S | polar | neutral | -0.8 |
| Threonine | Thr | T | polar | neutral | -0.7 |
| Tryptophan | Trp | W | nonpolar | neutral | -0.9 |
| Tyrosine | Tyr | Y | polar | neutral | -1.3 |
| Valine | Val | V | nonpolar | neutral | 4.2 |
Hydropathy Index[6]
- a number representing the hydrophobic or hydrophilic properties of the amino acid sidechain.
- larger the number the more hydrophobic the amino acid.
- most hydrophobic amino acids are isoleucine (4.5) and valine (4.2), hydrophobic amino acids tend to be internal
- most hydrophilic amino acids are arginine (-4.5) and lysine (-3.9), hydrophilic amino acids are more commonly found towards the protein surface.
- Links: Protein
Signaling Factors
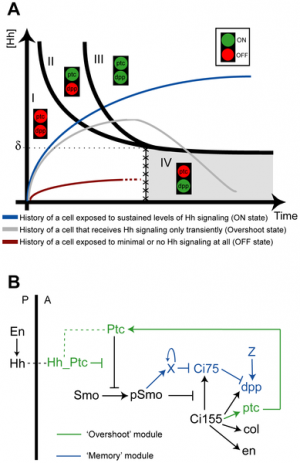
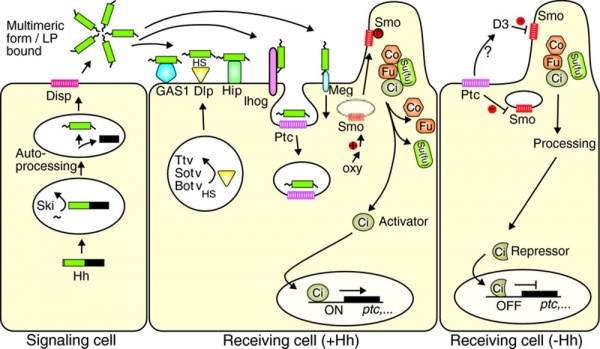
Hedgehog signaling pathway[7]
Note spelling difference: USA signaling, UK signalling.
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Axis Formation
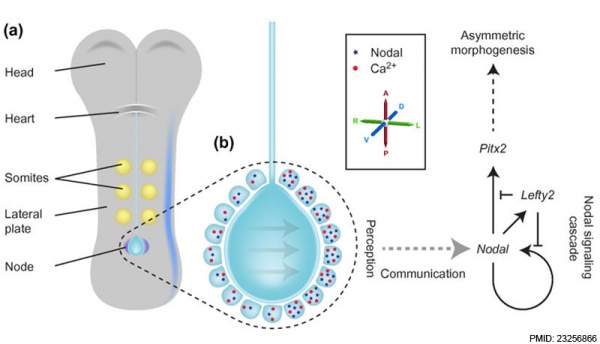
Embryo left-right asymmetry pathway[8]
The mechanisms that specify location within the embryo, therefore establishing axes, appears to be controlled by similar signaling mechanisms in different species.
- Links: Axis Formation
Transcription Factors
Transcription factors act by activating or inhibiting transcription either a single gene or more commonly acting on a family of genes. The transcription factor binding sites can be upstream of affected genes or some distance away from the initiation codon and many of these individual factors belong to families of many factors that share common DNA binding motifs.
- forkhead box (FOX) superfamily contain a DNA-binding motif known as the forkhead box or winged-helix domain.
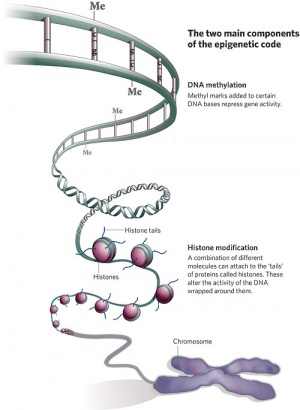
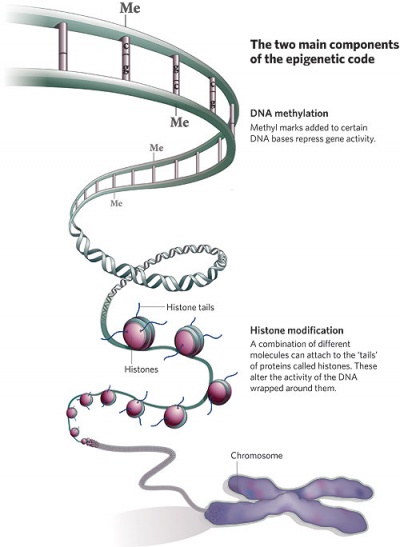
Epigenetics
| Epigenetics as the name implies, is the inheritance mechanisms that lie outside the DNA sequence of our genes.
One of the initial discoveries was the effects of DNA methylation upon gene expression and then modifications of nucleosomal histones. This DNA methylation, usually associated with 5-methylcytosine (m5C), leads to transcriptional silencing in vertebrates.
|
|
References
- ↑ <pubmed>16688142</pubmed>
- ↑ 2.0 2.1 <pubmed>108157</pubmed>
- ↑ <pubmed>26067255</pubmed>| Science.
- ↑ <pubmed>19104053</pubmed>| PNAS
- ↑ Martin Bachman, Santiago Uribe-Lewis, Xiaoping Yang, Heather E Burgess, Mario Iurlaro, Wolf Reik, Adele Murrell and Shankar Balasubramanian. 5-Formylcytosine can be a stable DNA modification in mammals Nature Chemical Biology (2015) doi:10.1038/nchembio.1848.
- ↑ 6.0 6.1 <pubmed>7108955</pubmed>
- ↑ <pubmed>19040769</pubmed>| PMC2614485 | Genome Biology
- ↑ <pubmed>23256866</pubmed>| BMC Biology
Terms
| Genetic Terms (expand to view) | ||
|---|---|---|
genetic abnormalities | Molecular Development | meiosis | mitosis
| ||
|
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- NHGR- The ENCODE Project ENCyclopedia Of DNA Elements
- Foldit - Amino Acids
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, June 21) Embryology Molecular Development. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Molecular_Development
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G