Developmental Signals - Fibroblast Growth Factor: Difference between revisions
mNo edit summary |
m (→Introduction) |
||
| (5 intermediate revisions by the same user not shown) | |||
| Line 3: | Line 3: | ||
[[File:Fgf_gene_family_evolution.jpg|thumb|300px|Fgf gene family evolution{{#pmid:20730630|PMID20730630}}]] | [[File:Fgf_gene_family_evolution.jpg|thumb|300px|Fgf gene family evolution{{#pmid:20730630|PMID20730630}}]] | ||
[[File:Mouse somitogenesis genes.jpg|thumb|Mouse somitogenesis genes{{#pmid:24304493|PMID24304493}}]] | [[File:Mouse somitogenesis genes.jpg|thumb|Mouse somitogenesis genes{{#pmid:24304493|PMID24304493}}]] | ||
Fibroblast Growth Factors ({{FGF}}) were originally identified by their ability to stimulate fibroblast cell proliferation but have a role in a | Fibroblast Growth Factors ({{FGF}}) were originally identified by their ability to stimulate fibroblast cell proliferation{{#pmid:4364816|PMID4364816}}{{#pmid:1170180|PMID1170180}} but have since been identified as having a role in a growth of a number of different tissues development and differentiation and continue to have a role in the adult. | ||
The first two identified factors were originally given the nomenclature of acidic or basic. We now know there to be at least 22 different human FGFs (Fgf1–Fgf23). These protein growth factors are bound by 4 different cell membrane receptors (FGFR1-4). FGFRs belong to the tyrosine kinase receptor family. | The first two identified factors were originally given the simple nomenclature of "acidic" or "basic". We now know there to be at least 22 different human FGFs (Fgf1–Fgf23). These protein growth factors are bound by 4 different cell membrane receptors (FGFR1-4). FGFRs belong to the tyrosine kinase receptor family. | ||
The mammalian Fgf family can be divided into the intracellular Fgf11/12/13/14 subfamily (iFGFs), the endocrine hormone-like Fgf15/21/23 subfamily (hFGFs), and the paracrine canonical Fgf subfamilies, including Fgf1/2/5, Fgf3/4/6, Fgf7/10/22, Fgf8/17/18, and Fgf9/16/20. | The mammalian Fgf family can be divided into the intracellular Fgf11/12/13/14 subfamily (iFGFs), the endocrine hormone-like Fgf15/21/23 subfamily (hFGFs), and the paracrine canonical Fgf subfamilies, including Fgf1/2/5, Fgf3/4/6, Fgf7/10/22, Fgf8/17/18, and Fgf9/16/20. | ||
| Line 32: | Line 31: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* '''Ovotesticular Disorders of Sex Development (DSD) in FGF9 Mouse Models of Human Synostosis Syndromes'''{{#pmid:32452519|PMID32452519}} "In {{mice}}, male sex determination depends on {{FGF}}9 signalling via FGFR2c in the bipotential gonads to maintain expression of the key testis gene SOX9. In humans however, while FGFR2 mutations have been linked to 46,XY disorders of sex development ({{DSD}}), the role of FGF9 is unresolved. The only reported pathogenic mutations in human FGF9; FGF9S99N and FGF9R62G, are dominant, and result in craniosynostosis (fusion of cranial sutures) or multiple synostoses (fusion of limb {{joint}}s). Whether these synostosis-causing FGF9 mutations impact upon gonadal development and DSD etiology has not been explored. We therefore examined embryonic gonads in the well-characterised Fgf9 missense mouse mutants; Fgf9S99N and Fgf9N143T, which phenocopy the skeletal defects of FGF9S99N and FGF9R62G variants respectively. XY Fgf9S99N/S99N and XY Fgf9N143T/N143T fetal mouse gonads showed severely disorganised testis cords and partial XY sex reversal at {{ME12.5}} days postcoitum (dpc), suggesting loss of FGF9 function. By {{ME15.5}} dpc, testis development in both mutants had partly recovered. Mitotic analysis in vivo and in vitro suggested that the testicular phenotypes in these mutants arise in part through reduced proliferation of the gonadal supporting cells. These data raise the possibility that human FGF9 mutations causative for dominant skeletal conditions can also lead to loss of FGF9 function in the developing {{testis}}, at least in mice. Our data suggest that in humans, testis development is largely tolerant of deleterious FGF9 mutations which lead to skeletal defects, thus offering an explanation as to why XY DSDs are rare in patients with pathogenic FGF9 variants." | |||
* '''The Warburg Effect and lactate signaling augment Fgf-MAPK to promote sensory-neural development in the otic vesicle'''{{#pmid:32338604|PMID32338604}} "Recent studies indicate that many developing tissues modify glycolysis to favor lactate synthesis, but how this promotes development is unclear. Using forward and reverse genetics in zebrafish, we show that disrupting the glycolytic gene phosphoglycerate kinase-1 (pgk1) impairs Fgf-dependent development of hair cells and neurons in the otic vesicle and other neurons in the CNS/PNS. Fgf-MAPK signaling underperforms in pgk1-/- mutants even when Fgf is transiently overexpressed. Wild-type embryos treated with drugs that block synthesis or secretion of lactate mimic the pgk1-/- phenotype, whereas pgk1-/- mutants are rescued by treatment with exogenous lactate. Lactate treatment of wild-type embryos elevates expression of Etv5b/Erm even when Fgf signaling is blocked. However, lactate's ability to stimulate neurogenesis is reversed by blocking MAPK. Thus, lactate raises basal levels of MAPK and Etv5b (a critical effector of the Fgf pathway), rendering cells more responsive to dynamic changes in Fgf signaling required by many developing tissues." | |||
* '''ERK Activity Dynamics during {{Zebrafish}} Embryonic Development'''{{#pmid:30597912|PMID30597912}} "During vertebrate development, extracellular signal-regulated kinase (ERK) is activated by growth factors such as fibroblast growth factor (FGF), and it regulates the formation of tissues/organs including eyes, brains, somites, limbs, and inner ears. However, an experimental system to monitor ERK activity dynamics in the entire body of the vertebrate embryo is lacking. We recently studied ERK activity dynamics in the pre-somitic mesoderm of living zebrafish embryos injected with mRNAs encoding a Förster resonance energy transfer (FRET)-based ERK biosensor. ... A spatiotemporal map of ERK activity in the entire body during zebrafish embryogenesis was generated, and previously unidentified activation dynamics and ERK domains were identified." | * '''ERK Activity Dynamics during {{Zebrafish}} Embryonic Development'''{{#pmid:30597912|PMID30597912}} "During vertebrate development, extracellular signal-regulated kinase (ERK) is activated by growth factors such as fibroblast growth factor (FGF), and it regulates the formation of tissues/organs including eyes, brains, somites, limbs, and inner ears. However, an experimental system to monitor ERK activity dynamics in the entire body of the vertebrate embryo is lacking. We recently studied ERK activity dynamics in the pre-somitic mesoderm of living zebrafish embryos injected with mRNAs encoding a Förster resonance energy transfer (FRET)-based ERK biosensor. ... A spatiotemporal map of ERK activity in the entire body during zebrafish embryogenesis was generated, and previously unidentified activation dynamics and ERK domains were identified." | ||
| Line 37: | Line 40: | ||
* '''A review of FGF signaling in {{palate}} development'''{{#pmid:29655165|PMID29655165}} "The fibroblast growth factors ({{FGF}}s) play a critical role during palatogenesis by mediating a variety of cellular responses. Extensive epidemiological and genetic studies over several decades in humans have revealed members of the FGF family function as candidate genes for syndromic and nonsyndromic cleft lip and cleft palate. The findings that FGFs signaling work delicately in the development of palate have been confirmed in mice carrying targeted mutations. Here we try to review recent progress toward a detailed understanding of FGF signaling including FGF7, FGF8, FGF9, FGF10, FGF18 and their receptors FGFR1, FGFR2 in palate development studies and discuss how they interact with other factors on the basis of animal studies regarding {{cleft palate}}." {{palate}} | * '''A review of FGF signaling in {{palate}} development'''{{#pmid:29655165|PMID29655165}} "The fibroblast growth factors ({{FGF}}s) play a critical role during palatogenesis by mediating a variety of cellular responses. Extensive epidemiological and genetic studies over several decades in humans have revealed members of the FGF family function as candidate genes for syndromic and nonsyndromic cleft lip and cleft palate. The findings that FGFs signaling work delicately in the development of palate have been confirmed in mice carrying targeted mutations. Here we try to review recent progress toward a detailed understanding of FGF signaling including FGF7, FGF8, FGF9, FGF10, FGF18 and their receptors FGFR1, FGFR2 in palate development studies and discuss how they interact with other factors on the basis of animal studies regarding {{cleft palate}}." {{palate}} | ||
|} | |} | ||
{| class="wikitable mw-collapsible mw-collapsed" | {| class="wikitable mw-collapsible mw-collapsed" | ||
| Line 46: | Line 46: | ||
| [[File:Mark_Hill.jpg|90px|left]] {{Most_Recent_Refs}} | | [[File:Mark_Hill.jpg|90px|left]] {{Most_Recent_Refs}} | ||
Search term: [http://www.ncbi.nlm.nih.gov/pubmed/?term=Fibroblast+Growth+Factor ''Fibroblast Growth Factor''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=FGF ''FGF''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=ERK ''ERK''] | Search term: [http://www.ncbi.nlm.nih.gov/pubmed/?term=Fibroblast+Growth+Factor ''Fibroblast Growth Factor''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=FGF ''FGF''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=ERK ''ERK''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Fibroblast+Growth+Factor+Receptor ''Fibroblast Growth Factor Receptor''] | ||
|} | |} | ||
| Line 53: | Line 53: | ||
|- | |- | ||
| {{Older papers}} | | {{Older papers}} | ||
* '''FGF8 morphogen gradients are differentially regulated by heparan sulphotransferases Hs2st and Hs6st1 in the developing brain'''{{#pmid:29158323|PMID29158323}} "Fibroblast growth factor (FGF) morphogen signalling through the evolutionarily ancient extracellular signalling-regulated kinase/mitogen activated protein kinase (ERK/MAPK) pathway recurs in many neural and non-neural developmental contexts, and understanding the mechanisms that regulate FGF/ERK function are correspondingly important. The glycosaminoglycan heparan sulphate (HS) binds to FGFs and exists in an enormous number of differentially sulphated forms produced by the action of HS modifying enzymes, and so has the potential to present an extremely large amount of information in FGF/ERK signalling. ...We discover that two different HS modifying enzymes, Hs2st and Hs6st1, indeed differentially modulate the properties of emerging Fgf8 protein concentration gradients and the Erk signalling output in response to Fgf8 in living tissue in ex vivo cultures. Both Hs2st and Hs6st1 are required for stable Fgf8 gradients to form as rapidly as they do in wild-type tissue while only Hs6st1 has a significant effect on suppressing the levels of Fgf8 protein in the gradient compared to wild type. Next we show that Hs2st and Hs6st1 act to antagonise and agonise the Erk signalling in response to Fgf8 protein, respectively, in ex vivo cultures of living tissue. Examination of endogenous Fgf8 protein and Erk signalling outputs in Hs2st-/- and Hs6st1-/- embryos suggests that our ex vivo findings have physiological relevance in vivo Our discovery identifies a new class of mechanism to tune Fgf8 function by regulated expression of Hs2st and Hs6st1 that is likely to have broader application to the >200 other signalling proteins that interact with HS and their function in neural development and disease." | |||
* '''Review - The Multiple Roles of FGF Signaling in the Developing Spinal Cord'''{{#pmid:28626748|PMID28626748}} "During vertebrate embryonic development, the {{spinal cord}} is formed by the neural derivatives of a neuromesodermal population that is specified at early stages of development and which develops in concert with the caudal regression of the primitive streak. Several processes related to {{spinal cord}} specification and maturation are coupled to this caudal extension including neurogenesis, ventral patterning and {{neural crest}} specification and all of them seem to be crucially regulated by Fibroblast Growth Factor ({{FGF}}) signaling, which is prominently active in the neuromesodermal region and transiently in its derivatives. Here we review the role of FGF signaling in those processes, trying to separate its different functions and highlighting the interactions with other signaling pathways." {{spinal cord}} | * '''Review - The Multiple Roles of FGF Signaling in the Developing Spinal Cord'''{{#pmid:28626748|PMID28626748}} "During vertebrate embryonic development, the {{spinal cord}} is formed by the neural derivatives of a neuromesodermal population that is specified at early stages of development and which develops in concert with the caudal regression of the primitive streak. Several processes related to {{spinal cord}} specification and maturation are coupled to this caudal extension including neurogenesis, ventral patterning and {{neural crest}} specification and all of them seem to be crucially regulated by Fibroblast Growth Factor ({{FGF}}) signaling, which is prominently active in the neuromesodermal region and transiently in its derivatives. Here we review the role of FGF signaling in those processes, trying to separate its different functions and highlighting the interactions with other signaling pathways." {{spinal cord}} | ||
| Line 72: | Line 74: | ||
{{Neural crest}} cell migration during early {{chicken}} embryo development{{#pmid:30520400|PMID30520400}} | {{Neural crest}} cell migration during early {{chicken}} embryo development{{#pmid:30520400|PMID30520400}} | ||
:"Fibroblast growth factor (FGF) signalling acts as one of modulators that control neural crest cell (NCC) migration, but how this is achieved is still unclear. In this study, we investigated the effects of FGF signalling on NCC migration by blocking this process. Constructs that were capable of inducing Sprouty2 (Spry2) or dominant-negative FGFR1 (Dn-FGFR1) expression were transfected into the cells making up the neural tubes. Our results revealed that blocking FGF signalling at stage HH10 (neurulation stage) could enhance NCC migration at both the cranial and trunk levels in the developing embryos. It was established that FGF-mediated NCC migration was not due to altering the expression of N-cadherin in the neural tube. Instead, we determined that cyclin D1 was overexpressed in the cranial and trunk levels when Sprouty2 was upregulated in the dorsal neural tube. These results imply that the cell cycle was a target of FGF signalling through which it regulates NCC migration at the neurulation stage." | :"Fibroblast growth factor (FGF) signalling acts as one of modulators that control neural crest cell (NCC) migration, but how this is achieved is still unclear. In this study, we investigated the effects of FGF signalling on NCC migration by blocking this process. Constructs that were capable of inducing Sprouty2 (Spry2) or dominant-negative FGFR1 (Dn-FGFR1) expression were transfected into the cells making up the neural tubes. Our results revealed that blocking FGF signalling at stage HH10 (neurulation stage) could enhance NCC migration at both the cranial and trunk levels in the developing embryos. It was established that FGF-mediated NCC migration was not due to altering the expression of N-cadherin in the neural tube. Instead, we determined that cyclin D1 was overexpressed in the cranial and trunk levels when Sprouty2 was upregulated in the dorsal neural tube. These results imply that the cell cycle was a target of FGF signalling through which it regulates NCC migration at the neurulation stage." | ||
===Spinal Cord=== | |||
'''Differentiation and localization of interneurons in the developing spinal cord depends on DOT1L expression'''{{#pmid:32471461|PMID32471461}} | |||
"Genetic and epigenetic factors contribute to the development of the spinal cord. Failure in correct exertion of the developmental programs, including neurulation, neural tube closure and neurogenesis of the diverse spinal cord neuronal subtypes results in defects of variable severity. We here report on the histone methyltransferase Disruptor of Telomeric 1 Like (DOT1L), which mediates histone H3 lysine 79 (H3K79) methylation. Conditional inactivation of DOT1L using Wnt1-cre as driver (Dot1l-cKO) showed that DOT1L expression is essential for spinal cord neurogenesis and localization of diverse neuronal subtypes, similar to its function in the development of the cerebral cortex and cerebellum. Transcriptome analysis revealed that DOT1L deficiency favored differentiation over progenitor proliferation. Dot1l-cKO mainly decreased the numbers of dI1 interneurons expressing Lhx2. In contrast, Lhx9 expressing dI1 interneurons did not change in numbers but localized differently upon Dot1l-cKO. Similarly, loss of DOT1L affected localization but not generation of dI2, dI3, dI5, V0 and V1 interneurons. The resulting derailed interneuron patterns might be responsible for increased cell death, occurrence of which was restricted to the late developmental stage {{ME18.5}}. Together our data indicate that DOT1L is essential for subtype-specific neurogenesis, migration and localization of dorsal and ventral interneurons in the developing spinal cord, in part by regulating transcriptional activation of Lhx2." | |||
==Endoderm== | ==Endoderm== | ||
| Line 80: | Line 88: | ||
:'''Links:''' | :'''Links:''' {{Endoderm}} | {{Chicken}} | ||
==Mesoderm== | ==Mesoderm== | ||
[[File:Model for Sprouty4 and FGF in mesoderm segmentation.jpg|400px]] | [[File:Model for Sprouty4 and FGF in mesoderm segmentation.jpg|400px]] | ||
| Line 150: | Line 157: | ||
* FGFR2 and FGFR3 have been associated with the Apert, Crouzon and Pfeiffer syndromes. | * FGFR2 and FGFR3 have been associated with the Apert, Crouzon and Pfeiffer syndromes. | ||
===FGF10=== | |||
A recent mouse knock-out study has shown a single-gene deletion of {{Fgf}}10 can generate {{Duodenal atresia}} in this animal model.{{#pmid:30473704|PMID30473704}} | |||
{| | |||
|+ Duodenal Atresia (Mouse Model){{#pmid:30473704|PMID30473704}} | |||
|- | |||
| [[File:Duodenal atresia 03.jpg|300px]] | |||
| valign=middle| | |||
('''A''') Normal gastric, pyloric, and duodenal morphology was demonstrated by wild type embryos. | |||
Null Fgf10 embryos universally demonstrated microgastria, but duodenal morphology varied according to presence and type of DA. Null embryos provided examples of: normal continuity and morphology of the duodenum for tm1 (D) and tm2 (E); type 1 DA, tm1 (F) and tm2 (G); type 2 DA, tm1 (H) and tm2 (I); and type 3 DA, tm1 (J) and tm2 (K). | |||
('''L''') Type 2 DA demonstrating a “double bubble” with significantly dilated proximal duodenum; tm2. | |||
('''M''') Type 3 DA demonstrating an intact esophagus, in the presence of tracheal atresia; tm1. Annotations denote genotype, scale bars, and location of pylorus, P. Arrows indicate location atresia in type 1 DA and type 2 DA examples. | |||
|} | |||
==References== | ==References== | ||
Latest revision as of 13:00, 25 July 2020
| Embryology - 28 Apr 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
Fibroblast Growth Factors (FGF) were originally identified by their ability to stimulate fibroblast cell proliferation[3][4] but have since been identified as having a role in a growth of a number of different tissues development and differentiation and continue to have a role in the adult.
The first two identified factors were originally given the simple nomenclature of "acidic" or "basic". We now know there to be at least 22 different human FGFs (Fgf1–Fgf23). These protein growth factors are bound by 4 different cell membrane receptors (FGFR1-4). FGFRs belong to the tyrosine kinase receptor family.
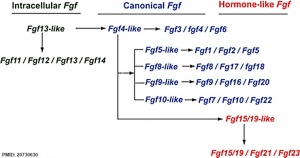
The mammalian Fgf family can be divided into the intracellular Fgf11/12/13/14 subfamily (iFGFs), the endocrine hormone-like Fgf15/21/23 subfamily (hFGFs), and the paracrine canonical Fgf subfamilies, including Fgf1/2/5, Fgf3/4/6, Fgf7/10/22, Fgf8/17/18, and Fgf9/16/20.
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Human FGF Family
| Table - Human Fibroblast Growth Factor Family FGF | ||||
| Approved Symbol |
Approved Name | Previous Symbols |
Synonyms | Chromosome |
|---|---|---|---|---|
| FGF1 | fibroblast growth factor 1 | FGFA | "AFGF, ECGF, ECGFA, ECGFB, HBGF1, ECGF-beta, FGF-alpha, GLIO703" | 5q31.3 |
| FGF2 | fibroblast growth factor 2 | FGFB | 4q28.1 | |
| FGF4 | fibroblast growth factor 4 | HSTF1 | "K-FGF, HBGF-4, HST, HST-1, KFGF" | 11q13.3 |
| FGF5 | fibroblast growth factor 5 | 4q21.21 | ||
| FGF6 | fibroblast growth factor 6 | 12p13.32 | ||
| FGF7 | fibroblast growth factor 7 | KGF | 15q21.2 | |
| FGF8 | fibroblast growth factor 8 | AIGF | 10q24.32 | |
| FGF9 | fibroblast growth factor 9 | 13q12.11 | ||
| Links: Developmental Signals - Fibroblast Growth Factor | OMIM Fgf1 | HGNC | Bmp Family | Fgf Family | Sox Family | Tbx Family | ||||
| Human FGF Family | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
Protein Properties
Human FGF
- ~150–300 amino acids
- have a conserved ~120-residue core with ~30–60% identity
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Fibroblast Growth Factor | FGF | ERK | Fibroblast Growth Factor Receptor |
| Older papers |
|---|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
|
Ectoderm
neural crest cell migration during early chicken embryo development[17]
- "Fibroblast growth factor (FGF) signalling acts as one of modulators that control neural crest cell (NCC) migration, but how this is achieved is still unclear. In this study, we investigated the effects of FGF signalling on NCC migration by blocking this process. Constructs that were capable of inducing Sprouty2 (Spry2) or dominant-negative FGFR1 (Dn-FGFR1) expression were transfected into the cells making up the neural tubes. Our results revealed that blocking FGF signalling at stage HH10 (neurulation stage) could enhance NCC migration at both the cranial and trunk levels in the developing embryos. It was established that FGF-mediated NCC migration was not due to altering the expression of N-cadherin in the neural tube. Instead, we determined that cyclin D1 was overexpressed in the cranial and trunk levels when Sprouty2 was upregulated in the dorsal neural tube. These results imply that the cell cycle was a target of FGF signalling through which it regulates NCC migration at the neurulation stage."
Spinal Cord
Differentiation and localization of interneurons in the developing spinal cord depends on DOT1L expression[18]
"Genetic and epigenetic factors contribute to the development of the spinal cord. Failure in correct exertion of the developmental programs, including neurulation, neural tube closure and neurogenesis of the diverse spinal cord neuronal subtypes results in defects of variable severity. We here report on the histone methyltransferase Disruptor of Telomeric 1 Like (DOT1L), which mediates histone H3 lysine 79 (H3K79) methylation. Conditional inactivation of DOT1L using Wnt1-cre as driver (Dot1l-cKO) showed that DOT1L expression is essential for spinal cord neurogenesis and localization of diverse neuronal subtypes, similar to its function in the development of the cerebral cortex and cerebellum. Transcriptome analysis revealed that DOT1L deficiency favored differentiation over progenitor proliferation. Dot1l-cKO mainly decreased the numbers of dI1 interneurons expressing Lhx2. In contrast, Lhx9 expressing dI1 interneurons did not change in numbers but localized differently upon Dot1l-cKO. Similarly, loss of DOT1L affected localization but not generation of dI2, dI3, dI5, V0 and V1 interneurons. The resulting derailed interneuron patterns might be responsible for increased cell death, occurrence of which was restricted to the late developmental stage E18.5. Together our data indicate that DOT1L is essential for subtype-specific neurogenesis, migration and localization of dorsal and ventral interneurons in the developing spinal cord, in part by regulating transcriptional activation of Lhx2."
Endoderm
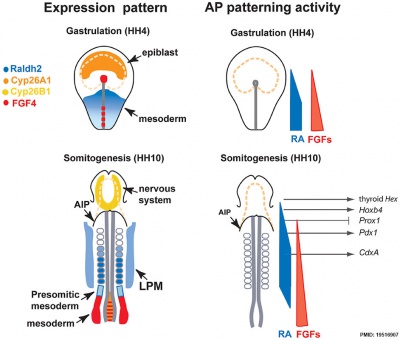
Chicken antero-posterior endoderm patterning[19]
Mesoderm
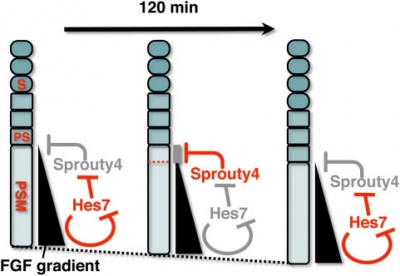
A Putative Model for the role of Sprouty4 as a mediator that links the mouse segmentation clock to the gradient of FGF signaling.[20]
The FGF signaling may be periodically inhibited by Sprouty4, by which temporal periodicity of Notch segmentation clock may be translated to spatial periodicity of the array of somites.
- In the PSM - FGF signaling establishes a posterior-to-anterior gradient, which is involved in the positioning of presumptive somite boundaries.
- Cyclic Sprouty4
- which is controlled by the Notch segmentation clock, the mechanism of which includes negative feedback loop of Hes7,
- may inhibit the FGF signaling possibly around the anterior border of the FGF signaling positive area
- where the FGF signaling is close to its threshold.
- S - somite
- PS - presumptive somite.
- Links: somitogenesis | Axial Skeleton Development | Notch | FGF
Bone
FGF9 has been shown to induce endochondral ossification in cranial mesenchyme in the mouse model.[21]
Respiration
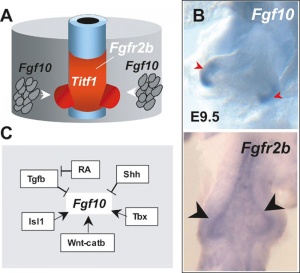
Lung Buds
Fibroblast growth factor 10 (FGF10) expression in mesoderm required for initial lung buds, through FGFR2IIIb transmembrane tyrosine kinase receptor protein.
Branching
Fibroblast growth factor 10 (FGF10) and sonic hedgehog (SHH) form a feedback loop for branching
- mesenchyme produced FGF10 signals to the distal epithelium to upregulate SHH expression.
- SHH then feeds back to inhibit Fgf10 expression in the adjacent mesenchyme, dividing in two the Fgf10 expression domain.
- new FGF10 signaling domains serve as two chemoattractant sources, leading to bifurcation of the epithelial tip.
Loop process mediated through FGF-activated transcription factor genes Etv4 and Etv5.[23]
- Links: respiratory
Limb
FGF soaked beads (FGF-1, FGF-2 and FGF-4) are capable of inducing additional limbs in chicken embryos.[24] A later study[25] identified the endogenous signal as Fgf-8 from initially the intermediate mesoderm and then the prelimb field ectoderm for limb initiation and outgrowth, respectively.
Hearing
- fibroblast growth factor 1 - (Fgf-1) a growth factor released from cochlea sensory epithelium which stimulates spiral ganglion neurite branching.
- fibroblast growth factor 8 - (Fgf-8) a growth factor released by inner hair cells which regulates pillar cell number, position and rate of development.
- fibroblast growth factor receptor 3 - (Fgfr-3) a tyrosine kinase receptor with a role in the commitment, differentiation and position of pillar cells in the organ of corti
- Links: hearing
Palate
Fibroblast growth factors are required during palatogenesis and are candidate genes for syndromic and nonsyndromic cleft lip and cleft palate, see the recent review.[9]
FGF7, FGF8, FGF9, FGF10, FGF18 and their receptors FGFR1, FGFR2
- Links: palate | cleft palate
Abnormalities
- FGFR1 mutation has been associated with the relatively milder form of Pfeiffer syndrome type 1.
- FGFR2 and FGFR3 have been associated with the Apert, Crouzon and Pfeiffer syndromes.
FGF10
A recent mouse knock-out study has shown a single-gene deletion of Fgf10 can generate duodenal atresia in this animal model.[26]
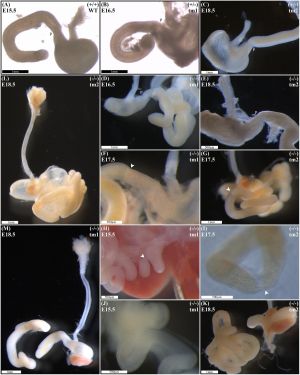
|
(A) Normal gastric, pyloric, and duodenal morphology was demonstrated by wild type embryos. Null Fgf10 embryos universally demonstrated microgastria, but duodenal morphology varied according to presence and type of DA. Null embryos provided examples of: normal continuity and morphology of the duodenum for tm1 (D) and tm2 (E); type 1 DA, tm1 (F) and tm2 (G); type 2 DA, tm1 (H) and tm2 (I); and type 3 DA, tm1 (J) and tm2 (K).
|
References
- ↑ Itoh N. (2010). Hormone-like (endocrine) Fgfs: their evolutionary history and roles in development, metabolism, and disease. Cell Tissue Res. , 342, 1-11. PMID: 20730630 DOI.
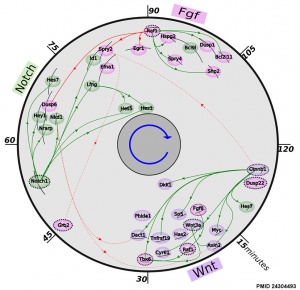
- ↑ 2.0 2.1 Fongang B & Kudlicki A. (2013). The precise timeline of transcriptional regulation reveals causation in mouse somitogenesis network. BMC Dev. Biol. , 13, 42. PMID: 24304493 DOI.
- ↑ Gospodarowicz D. (1974). Localisation of a fibroblast growth factor and its effect alone and with hydrocortisone on 3T3 cell growth. Nature , 249, 123-7. PMID: 4364816 DOI.
- ↑ Gospodarowicz D & Moran JS. (1975). Mitogenic effect of fibroblast growth factor on early passage cultures of human and murine fibroblasts. J. Cell Biol. , 66, 451-7. PMID: 1170180 DOI.
- ↑ Bird AD, Croft BM, Harada M, Tang L, Zhao L, Ming Z, Bagheri-Fam S, Koopman P, Wang Z, Akita K & Harley VR. (2020). Ovotesticular disorders of sex development (DSD) in FGF9 mouse models of human synostosis syndromes. Hum. Mol. Genet. , , . PMID: 32452519 DOI.
- ↑ Kantarci H, Gou Y & Riley BB. (2020). The Warburg Effect and lactate signaling augment Fgf-MAPK to promote sensory-neural development in the otic vesicle. Elife , 9, . PMID: 32338604 DOI.
- ↑ Wong KL, Akiyama R, Bessho Y & Matsui T. (2018). ERK Activity Dynamics during Zebrafish Embryonic Development. Int J Mol Sci , 20, . PMID: 30597912 DOI.
- ↑ Row RH, Pegg A, Kinney B, Farr GH, Maves L, Lowell S, Wilson V & Martin BL. (2018). BMP and FGF signaling interact to pattern mesoderm by controlling basic helix-loop-helix transcription factor activity. Elife , 7, . PMID: 29877796 DOI.
- ↑ 9.0 9.1 Weng M, Chen Z, Xiao Q, Li R & Chen Z. (2018). A review of FGF signaling in palate development. Biomed. Pharmacother. , 103, 240-247. PMID: 29655165 DOI.
- ↑ Chan WK, Price DJ & Pratt T. (2017). FGF8 morphogen gradients are differentially regulated by heparan sulphotransferases Hs2st and Hs6st1 in the developing brain. Biol Open , 6, 1933-1942. PMID: 29158323 DOI.
- ↑ Diez Del Corral R & Morales AV. (2017). The Multiple Roles of FGF Signaling in the Developing Spinal Cord. Front Cell Dev Biol , 5, 58. PMID: 28626748 DOI.
- ↑ Atsuta Y & Takahashi Y. (2015). FGF8 coordinates tissue elongation and cell epithelialization during early kidney tubulogenesis. Development , 142, 2329-37. PMID: 26130757 DOI.
- ↑ Gredler ML, Seifert AW & Cohn MJ. (2015). Tissue-specific roles of Fgfr2 in development of the external genitalia. Development , 142, 2203-12. PMID: 26081573 DOI.
- ↑ Shifley ET, Kenny AP, Rankin SA & Zorn AM. (2012). Prolonged FGF signaling is necessary for lung and liver induction in Xenopus. BMC Dev. Biol. , 12, 27. PMID: 22988910 DOI.
- ↑ Lahti L, Saarimäki-Vire J, Rita H & Partanen J. (2011). FGF signaling gradient maintains symmetrical proliferative divisions of midbrain neuronal progenitors. Dev. Biol. , 349, 270-82. PMID: 21074523 DOI.
- ↑ Yu SR, Burkhardt M, Nowak M, Ries J, Petrásek Z, Scholpp S, Schwille P & Brand M. (2009). Fgf8 morphogen gradient forms by a source-sink mechanism with freely diffusing molecules. Nature , 461, 533-6. PMID: 19741606 DOI.
- ↑ Zhang XT, Wang G, Li Y, Chuai M, Lee KKH & Yang X. (2018). Role of FGF signalling in neural crest cell migration during early chick embryo development. Zygote , , 1-8. PMID: 30520400 DOI.
- ↑ Gray de Cristoforis A, Ferrari F, Clotman F & Vogel T. (2020). Differentiation and localization of interneurons in the developing spinal cord depends on DOT1L expression. Mol Brain , 13, 85. PMID: 32471461 DOI.
- ↑ Bayha E, Jørgensen MC, Serup P & Grapin-Botton A. (2009). Retinoic acid signaling organizes endodermal organ specification along the entire antero-posterior axis. PLoS ONE , 4, e5845. PMID: 19516907 DOI.
- ↑ Hayashi S, Shimoda T, Nakajima M, Tsukada Y, Sakumura Y, Dale JK, Maroto M, Kohno K, Matsui T & Bessho Y. (2009). Sprouty4, an FGF inhibitor, displays cyclic gene expression under the control of the notch segmentation clock in the mouse PSM. PLoS ONE , 4, e5603. PMID: 19440349 DOI.
- ↑ Govindarajan V & Overbeek PA. (2006). FGF9 can induce endochondral ossification in cranial mesenchyme. BMC Dev. Biol. , 6, 7. PMID: 16504022 DOI.
- ↑ Cardoso WV & Kotton DN. (2008). Specification and patterning of the respiratory system. , , . PMID: 20614584 DOI.
- ↑ Herriges JC, Verheyden JM, Zhang Z, Sui P, Zhang Y, Anderson MJ, Swing DA, Zhang Y, Lewandoski M & Sun X. (2015). FGF-Regulated ETV Transcription Factors Control FGF-SHH Feedback Loop in Lung Branching. Dev. Cell , 35, 322-32. PMID: 26555052 DOI.
- ↑ Cohn MJ, Izpisúa-Belmonte JC, Abud H, Heath JK & Tickle C. (1995). Fibroblast growth factors induce additional limb development from the flank of chick embryos. Cell , 80, 739-46. PMID: 7889567
- ↑ Vogel A, Rodriguez C & Izpisúa-Belmonte JC. (1996). Involvement of FGF-8 in initiation, outgrowth and patterning of the vertebrate limb. Development , 122, 1737-50. PMID: 8674413
- ↑ 26.0 26.1 Teague WJ, Jones MLM, Hawkey L, Smyth IM, Catubig A, King SK, Sarila G, Li R & Hutson JM. (2018). FGF10 and the Mystery of Duodenal Atresia in Humans. Front Genet , 9, 530. PMID: 30473704 DOI.
Reviews
Articles
Su N, Xu X, Li C, He Q, Zhao L, Li C, Chen S, Luo F, Yi L, Du X, Huang H, Deng C & Chen L. (2010). Generation of Fgfr3 conditional knockout mice. Int. J. Biol. Sci. , 6, 327-32. PMID: 20582225
Search Pubmed
Search Bookshelf Fibroblast Growth Factor
Search Pubmed Now: Fibroblast Growth Factor
Cite this page: Hill, M.A. (2024, April 28) Embryology Developmental Signals - Fibroblast Growth Factor. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Fibroblast_Growth_Factor
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G