Developmental Signals - Fibroblast Growth Factor: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 34: | Line 34: | ||
* '''{{BMP}} and {{FGF}} signaling interact to pattern {{mesoderm}} by controlling basic helix-loop-helix transcription factor activity'''{{#pmid:29877796|PMID29877796}} "The {{mesoderm}}al germ layer is patterned into mediolateral subtypes by signaling factors including {{BMP}} and {{FGF}}. How these pathways are integrated to induce specific mediolateral cell fates is not well understood. We used mesoderm derived from post-gastrulation neuromesodermal progenitors (NMPs), which undergo a binary mediolateral patterning decision, as a simplified model to understand how FGF acts together with BMP to impart mediolateral fate. Using zebrafish and mouse NMPs, we identify an evolutionarily conserved mechanism of BMP and FGF mediated mediolateral mesodermal patterning that occurs through modulation of basic helix-loop-helix (bHLH) transcription factor activity. BMP imparts lateral fate through induction of Id helix loop helix (HLH) proteins, which antagonize bHLH transcription factors, induced by FGF signaling, that specify medial fate. We extend our analysis of zebrafish development to show that bHLH activity is responsible for the mediolateral patterning of the entire mesodermal germ layer." | * '''{{BMP}} and {{FGF}} signaling interact to pattern {{mesoderm}} by controlling basic helix-loop-helix transcription factor activity'''{{#pmid:29877796|PMID29877796}} "The {{mesoderm}}al germ layer is patterned into mediolateral subtypes by signaling factors including {{BMP}} and {{FGF}}. How these pathways are integrated to induce specific mediolateral cell fates is not well understood. We used mesoderm derived from post-gastrulation neuromesodermal progenitors (NMPs), which undergo a binary mediolateral patterning decision, as a simplified model to understand how FGF acts together with BMP to impart mediolateral fate. Using zebrafish and mouse NMPs, we identify an evolutionarily conserved mechanism of BMP and FGF mediated mediolateral mesodermal patterning that occurs through modulation of basic helix-loop-helix (bHLH) transcription factor activity. BMP imparts lateral fate through induction of Id helix loop helix (HLH) proteins, which antagonize bHLH transcription factors, induced by FGF signaling, that specify medial fate. We extend our analysis of zebrafish development to show that bHLH activity is responsible for the mediolateral patterning of the entire mesodermal germ layer." | ||
* '''A review of FGF signaling in {{palate}} development'''{{#pmid:29655165|PMID29655165}} "The fibroblast growth factors ({{FGF}}s) play a critical role during palatogenesis by mediating a variety of cellular responses. Extensive epidemiological and genetic studies over several decades in humans have revealed members of the FGF family function as candidate genes for syndromic and nonsyndromic cleft lip and cleft palate. The findings that FGFs signaling work delicately in the development of palate have been confirmed in mice carrying targeted mutations. Here we try to review recent progress toward a detailed understanding of FGF signaling including FGF7, FGF8, FGF9, FGF10, FGF18 and their receptors FGFR1, FGFR2 in palate development studies and discuss how they interact with other factors on the basis of animal studies regarding cleft palate." | * '''A review of FGF signaling in {{palate}} development'''{{#pmid:29655165|PMID29655165}} "The fibroblast growth factors ({{FGF}}s) play a critical role during palatogenesis by mediating a variety of cellular responses. Extensive epidemiological and genetic studies over several decades in humans have revealed members of the FGF family function as candidate genes for syndromic and nonsyndromic cleft lip and cleft palate. The findings that FGFs signaling work delicately in the development of palate have been confirmed in mice carrying targeted mutations. Here we try to review recent progress toward a detailed understanding of FGF signaling including FGF7, FGF8, FGF9, FGF10, FGF18 and their receptors FGFR1, FGFR2 in palate development studies and discuss how they interact with other factors on the basis of animal studies regarding {{cleft palate}}." {{palate}} | ||
* '''Review - The Multiple Roles of FGF Signaling in the Developing Spinal Cord'''{{#pmid:28626748|PMID28626748}} "During vertebrate embryonic development, the {{spinal cord}} is formed by the neural derivatives of a neuromesodermal population that is specified at early stages of development and which develops in concert with the caudal regression of the primitive streak. Several processes related to {{spinal cord}} specification and maturation are coupled to this caudal extension including neurogenesis, ventral patterning and {{neural crest}} specification and all of them seem to be crucially regulated by Fibroblast Growth Factor ({{FGF}}) signaling, which is prominently active in the neuromesodermal region and transiently in its derivatives. Here we review the role of FGF signaling in those processes, trying to separate its different functions and highlighting the interactions with other signaling pathways." {{spinal cord}} | * '''Review - The Multiple Roles of FGF Signaling in the Developing Spinal Cord'''{{#pmid:28626748|PMID28626748}} "During vertebrate embryonic development, the {{spinal cord}} is formed by the neural derivatives of a neuromesodermal population that is specified at early stages of development and which develops in concert with the caudal regression of the primitive streak. Several processes related to {{spinal cord}} specification and maturation are coupled to this caudal extension including neurogenesis, ventral patterning and {{neural crest}} specification and all of them seem to be crucially regulated by Fibroblast Growth Factor ({{FGF}}) signaling, which is prominently active in the neuromesodermal region and transiently in its derivatives. Here we review the role of FGF signaling in those processes, trying to separate its different functions and highlighting the interactions with other signaling pathways." {{spinal cord}} | ||
| Line 93: | Line 93: | ||
:'''Links:''' | :'''Links:''' {{Somitogenesis}} | [[Musculoskeletal System - Axial Skeleton Development|Axial Skeleton Development]] | [[Developmental Signals - Notch|Notch]] | [[Developmental_Signals_-_Fibroblast_Growth_Factor|FGF]] | ||
| Line 102: | Line 102: | ||
==Respiration== | ==Respiration== | ||
[[File:Mouse- respiratory development 04.jpg|thumb|Mouse Respiratory Development Fibroblast Growth Factor Signaling | [[File:Mouse- respiratory development 04.jpg|thumb|Mouse Respiratory Development Fibroblast Growth Factor Signaling{{#pmid:20614584|PMID20614584}}]] | ||
===Lung Buds=== | ===Lung Buds=== | ||
Fibroblast growth factor 10 (FGF10) expression in mesoderm required for initial lung buds, through FGFR2IIIb transmembrane tyrosine kinase receptor protein. | Fibroblast growth factor 10 (FGF10) expression in mesoderm required for initial lung buds, through FGFR2IIIb transmembrane tyrosine kinase receptor protein. | ||
===Branching=== | ===Branching=== | ||
Fibroblast growth factor 10 (FGF10) and | Fibroblast growth factor 10 (FGF10) and {{sonic hedgehog}} ({{SHH}}) form a feedback loop for branching | ||
* mesenchyme produced FGF10 signals to the distal epithelium to upregulate SHH expression. | * mesenchyme produced FGF10 signals to the distal epithelium to upregulate SHH expression. | ||
* SHH then feeds back to inhibit Fgf10 expression in the adjacent mesenchyme, dividing in two the Fgf10 expression domain. | * SHH then feeds back to inhibit Fgf10 expression in the adjacent mesenchyme, dividing in two the Fgf10 expression domain. | ||
| Line 114: | Line 114: | ||
Loop process mediated through FGF-activated transcription factor genes Etv4 and Etv5.{{#pmid:26555052|PMID26555052}} | Loop process mediated through FGF-activated transcription factor genes Etv4 and Etv5.{{#pmid:26555052|PMID26555052}} | ||
:'''Links:''' | :'''Links:''' {{Respiratory}} | ||
==Limb== | ==Limb== | ||
| Line 120: | Line 120: | ||
FGF soaked beads (FGF-1, FGF-2 and FGF-4) are capable of inducing additional limbs in chicken embryos.{{#pmid:7889567|PMID7889567}} A later study{{#pmid:8674413|PMID8674413}} identified the endogenous signal as Fgf-8 from initially the intermediate mesoderm and then the prelimb field ectoderm for limb initiation and outgrowth, respectively. | FGF soaked beads (FGF-1, FGF-2 and FGF-4) are capable of inducing additional limbs in chicken embryos.{{#pmid:7889567|PMID7889567}} A later study{{#pmid:8674413|PMID8674413}} identified the endogenous signal as Fgf-8 from initially the intermediate mesoderm and then the prelimb field ectoderm for limb initiation and outgrowth, respectively. | ||
:'''Links:''' | :'''Links:''' {{Limb}}] | {{Chicken}} | ||
==Hearing== | ==Hearing== | ||
| Line 127: | Line 127: | ||
* '''fibroblast growth factor receptor 3''' - (Fgfr-3) a tyrosine kinase receptor with a role in the commitment, differentiation and position of pillar cells in the organ of corti | * '''fibroblast growth factor receptor 3''' - (Fgfr-3) a tyrosine kinase receptor with a role in the commitment, differentiation and position of pillar cells in the organ of corti | ||
:'''Links:''' | :'''Links:''' {{Hearing}} | ||
==Palate== | |||
Fibroblast growth factors are required during palatogenesis and are candidate genes for syndromic and nonsyndromic cleft lip and cleft palate, see the recent review.{{#pmid:29655165|PMID29655165}} | |||
FGF7, FGF8, FGF9, FGF10, FGF18 and their receptors FGFR1, FGFR2 | |||
:'''Links:''' {{palate}} | {{cleft palate}} | |||
==Abnormalities== | ==Abnormalities== | ||
Revision as of 06:59, 24 November 2018
| Embryology - 14 May 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
Fibroblast Growth Factors (FGF) were originally identified by their ability to stimulate fibroblast cell proliferation but have a role in a growing number of different tissues development and differentiation and continue to have a role in the adult.
The first two identified factors were originally given the nomenclature of acidic or basic. We now know there to be at least 22 different human FGFs (Fgf1–Fgf23). These protein growth factors are bound by 4 different cell membrane receptors (FGFR1-4). FGFRs belong to the tyrosine kinase receptor family.
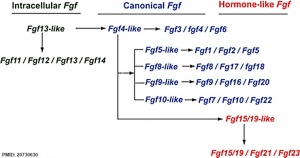
The mammalian Fgf family can be divided into the intracellular Fgf11/12/13/14 subfamily (iFGFs), the endocrine hormone-like Fgf15/21/23 subfamily (hFGFs), and the paracrine canonical Fgf subfamilies, including Fgf1/2/5, Fgf3/4/6, Fgf7/10/22, Fgf8/17/18, and Fgf9/16/20.
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Human FGF Family
| Table - Human Fibroblast Growth Factor Family FGF | ||||
| Approved Symbol |
Approved Name | Previous Symbols |
Synonyms | Chromosome |
|---|---|---|---|---|
| FGF1 | fibroblast growth factor 1 | FGFA | "AFGF, ECGF, ECGFA, ECGFB, HBGF1, ECGF-beta, FGF-alpha, GLIO703" | 5q31.3 |
| FGF2 | fibroblast growth factor 2 | FGFB | 4q28.1 | |
| FGF4 | fibroblast growth factor 4 | HSTF1 | "K-FGF, HBGF-4, HST, HST-1, KFGF" | 11q13.3 |
| FGF5 | fibroblast growth factor 5 | 4q21.21 | ||
| FGF6 | fibroblast growth factor 6 | 12p13.32 | ||
| FGF7 | fibroblast growth factor 7 | KGF | 15q21.2 | |
| FGF8 | fibroblast growth factor 8 | AIGF | 10q24.32 | |
| FGF9 | fibroblast growth factor 9 | 13q12.11 | ||
| Links: Developmental Signals - Fibroblast Growth Factor | OMIM Fgf1 | HGNC | Bmp Family | Fgf Family | Sox Family | Tbx Family | ||||
| Human FGF Family | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
Protein Properties
Human FGF
- ~150–300 amino acids
- have a conserved ~120-residue core with ~30–60% identity
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Fibroblast Growth Factor <pubmed limit=5>Fibroblast Growth Factor</pubmed> |
| Older papers |
|---|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
|
Endoderm
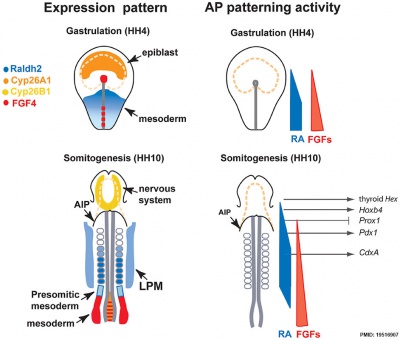
Chicken antero-posterior endoderm patterning[11]
- Links: Endoderm | Chicken Development
Mesoderm
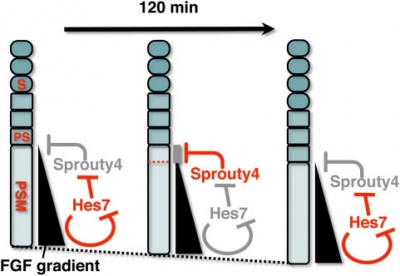
A Putative Model for the role of Sprouty4 as a mediator that links the mouse segmentation clock to the gradient of FGF signaling.[12]
The FGF signaling may be periodically inhibited by Sprouty4, by which temporal periodicity of Notch segmentation clock may be translated to spatial periodicity of the array of somites.
- In the PSM - FGF signaling establishes a posterior-to-anterior gradient, which is involved in the positioning of presumptive somite boundaries.
- Cyclic Sprouty4
- which is controlled by the Notch segmentation clock, the mechanism of which includes negative feedback loop of Hes7,
- may inhibit the FGF signaling possibly around the anterior border of the FGF signaling positive area
- where the FGF signaling is close to its threshold.
- S - somite
- PS - presumptive somite.
- Links: somitogenesis | Axial Skeleton Development | Notch | FGF
Bone
FGF9 has been shown to induce endochondral ossification in cranial mesenchyme in the mouse model.[13]
Respiration
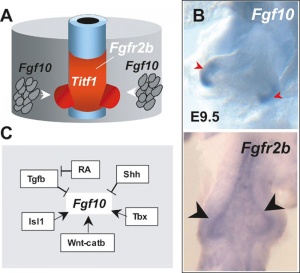
Lung Buds
Fibroblast growth factor 10 (FGF10) expression in mesoderm required for initial lung buds, through FGFR2IIIb transmembrane tyrosine kinase receptor protein.
Branching
Fibroblast growth factor 10 (FGF10) and sonic hedgehog (SHH) form a feedback loop for branching
- mesenchyme produced FGF10 signals to the distal epithelium to upregulate SHH expression.
- SHH then feeds back to inhibit Fgf10 expression in the adjacent mesenchyme, dividing in two the Fgf10 expression domain.
- new FGF10 signaling domains serve as two chemoattractant sources, leading to bifurcation of the epithelial tip.
Loop process mediated through FGF-activated transcription factor genes Etv4 and Etv5.[15]
- Links: respiratory
Limb
FGF soaked beads (FGF-1, FGF-2 and FGF-4) are capable of inducing additional limbs in chicken embryos.[16] A later study[17] identified the endogenous signal as Fgf-8 from initially the intermediate mesoderm and then the prelimb field ectoderm for limb initiation and outgrowth, respectively.
Hearing
- fibroblast growth factor 1 - (Fgf-1) a growth factor released from cochlea sensory epithelium which stimulates spiral ganglion neurite branching.
- fibroblast growth factor 8 - (Fgf-8) a growth factor released by inner hair cells which regulates pillar cell number, position and rate of development.
- fibroblast growth factor receptor 3 - (Fgfr-3) a tyrosine kinase receptor with a role in the commitment, differentiation and position of pillar cells in the organ of corti
- Links: hearing
Palate
Fibroblast growth factors are required during palatogenesis and are candidate genes for syndromic and nonsyndromic cleft lip and cleft palate, see the recent review.[4]
FGF7, FGF8, FGF9, FGF10, FGF18 and their receptors FGFR1, FGFR2
- Links: palate | cleft palate
Abnormalities
- FGFR1 mutation has been associated with the relatively milder form of Pfeiffer syndrome type 1.
- FGFR2 and FGFR3 have been associated with the Apert, Crouzon and Pfeiffer syndromes.
References
- ↑ Itoh N. (2010). Hormone-like (endocrine) Fgfs: their evolutionary history and roles in development, metabolism, and disease. Cell Tissue Res. , 342, 1-11. PMID: 20730630 DOI.
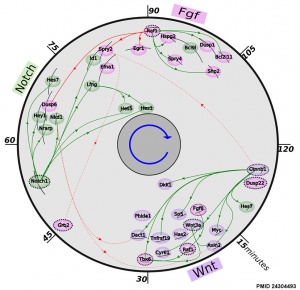
- ↑ 2.0 2.1 Fongang B & Kudlicki A. (2013). The precise timeline of transcriptional regulation reveals causation in mouse somitogenesis network. BMC Dev. Biol. , 13, 42. PMID: 24304493 DOI.
- ↑ Row RH, Pegg A, Kinney B, Farr GH, Maves L, Lowell S, Wilson V & Martin BL. (2018). BMP and FGF signaling interact to pattern mesoderm by controlling basic helix-loop-helix transcription factor activity. Elife , 7, . PMID: 29877796 DOI.
- ↑ 4.0 4.1 Weng M, Chen Z, Xiao Q, Li R & Chen Z. (2018). A review of FGF signaling in palate development. Biomed. Pharmacother. , 103, 240-247. PMID: 29655165 DOI.
- ↑ Diez Del Corral R & Morales AV. (2017). The Multiple Roles of FGF Signaling in the Developing Spinal Cord. Front Cell Dev Biol , 5, 58. PMID: 28626748 DOI.
- ↑ Atsuta Y & Takahashi Y. (2015). FGF8 coordinates tissue elongation and cell epithelialization during early kidney tubulogenesis. Development , 142, 2329-37. PMID: 26130757 DOI.
- ↑ Gredler ML, Seifert AW & Cohn MJ. (2015). Tissue-specific roles of Fgfr2 in development of the external genitalia. Development , 142, 2203-12. PMID: 26081573 DOI.
- ↑ Shifley ET, Kenny AP, Rankin SA & Zorn AM. (2012). Prolonged FGF signaling is necessary for lung and liver induction in Xenopus. BMC Dev. Biol. , 12, 27. PMID: 22988910 DOI.
- ↑ Lahti L, Saarimäki-Vire J, Rita H & Partanen J. (2011). FGF signaling gradient maintains symmetrical proliferative divisions of midbrain neuronal progenitors. Dev. Biol. , 349, 270-82. PMID: 21074523 DOI.
- ↑ Yu SR, Burkhardt M, Nowak M, Ries J, Petrásek Z, Scholpp S, Schwille P & Brand M. (2009). Fgf8 morphogen gradient forms by a source-sink mechanism with freely diffusing molecules. Nature , 461, 533-6. PMID: 19741606 DOI.
- ↑ Bayha E, Jørgensen MC, Serup P & Grapin-Botton A. (2009). Retinoic acid signaling organizes endodermal organ specification along the entire antero-posterior axis. PLoS ONE , 4, e5845. PMID: 19516907 DOI.
- ↑ Hayashi S, Shimoda T, Nakajima M, Tsukada Y, Sakumura Y, Dale JK, Maroto M, Kohno K, Matsui T & Bessho Y. (2009). Sprouty4, an FGF inhibitor, displays cyclic gene expression under the control of the notch segmentation clock in the mouse PSM. PLoS ONE , 4, e5603. PMID: 19440349 DOI.
- ↑ Govindarajan V & Overbeek PA. (2006). FGF9 can induce endochondral ossification in cranial mesenchyme. BMC Dev. Biol. , 6, 7. PMID: 16504022 DOI.
- ↑ Cardoso WV & Kotton DN. (2008). Specification and patterning of the respiratory system. , , . PMID: 20614584 DOI.
- ↑ Herriges JC, Verheyden JM, Zhang Z, Sui P, Zhang Y, Anderson MJ, Swing DA, Zhang Y, Lewandoski M & Sun X. (2015). FGF-Regulated ETV Transcription Factors Control FGF-SHH Feedback Loop in Lung Branching. Dev. Cell , 35, 322-32. PMID: 26555052 DOI.
- ↑ Cohn MJ, Izpisúa-Belmonte JC, Abud H, Heath JK & Tickle C. (1995). Fibroblast growth factors induce additional limb development from the flank of chick embryos. Cell , 80, 739-46. PMID: 7889567
- ↑ Vogel A, Rodriguez C & Izpisúa-Belmonte JC. (1996). Involvement of FGF-8 in initiation, outgrowth and patterning of the vertebrate limb. Development , 122, 1737-50. PMID: 8674413
Reviews
Articles
Su N, Xu X, Li C, He Q, Zhao L, Li C, Chen S, Luo F, Yi L, Du X, Huang H, Deng C & Chen L. (2010). Generation of Fgfr3 conditional knockout mice. Int. J. Biol. Sci. , 6, 327-32. PMID: 20582225
Search Pubmed
Search Bookshelf Fibroblast Growth Factor
Search Pubmed Now: Fibroblast Growth Factor
Cite this page: Hill, M.A. (2024, May 14) Embryology Developmental Signals - Fibroblast Growth Factor. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Fibroblast_Growth_Factor
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G