Developmental Signals - Homeobox: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| (2 intermediate revisions by the same user not shown) | |||
| Line 34: | Line 34: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* '''The Hox Gene egl-5 Acts as a Terminal Selector for VD13 Development via Wnt Signaling'''{{#pmid:32138237|PMID32138237}} "Nervous systems are comprised of diverse cell types that differ functionally and morphologically. During development, extrinsic signals, e.g., growth factors, can activate intrinsic programs, usually orchestrated by networks of transcription factors. Within that network, transcription factors that drive the specification of features specific to a limited number of cells are often referred to as terminal selectors. While we still have an incomplete view of how individual neurons within organisms become specified, reporters limited to a subset of neurons in a nervous system can facilitate the discovery of cell specification programs. We have identified a fluorescent reporter that labels VD13, the most posterior of the 19 inhibitory GABA (γ-amino butyric acid)-ergic motorneurons, and two additional neurons, LUAL and LUAR. Loss of function in multiple Wnt signaling genes resulted in an incompletely penetrant loss of the marker, selectively in VD13, but not the LUAs, even though other aspects of GABAergic specification in VD13 were normal. The posterior Hox gene, egl-5, was necessary for expression of our marker in VD13, and ectopic expression of egl-5 in more anterior GABAergic neurons induced expression of the marker. These results suggest egl-5 is a terminal selector of VD13, subsequent to GABAergic specification." {{worm}} {{neural}} | |||
* '''The formation of the thumb requires direct modulation of Gli3 transcription by {{Hox}}a13'''{{#pmid:31896583|PMID31896583}} "In the tetrapod limb, the digits (fingers or toes) are the elements most subject to morphological diversification in response to functional adaptations. However, despite their functional importance, the mechanisms controlling digit morphology remain poorly understood. Here we have focused on understanding the special morphology of the thumb (digit 1), the acquisition of which was an important adaptation of the human hand. To this end, we have studied the limbs of the Hoxa13 mouse mutant that specifically fail to form digit 1. We show that, consistent with the role of Hoxa13 in Hoxd transcriptional regulation, the expression of Hoxd13 in Hoxa13 mutant limbs does not extend into the presumptive digit 1 territory, which is therefore devoid of distal Hox transcripts, a circumstance that can explain its agenesis. The loss of Hoxd13 expression, exclusively in digit 1 territory, correlates with increased Gli3 repressor activity, a Hoxd negative regulator, resulting from increased Gli3 transcription that, in turn, is due to the release from the negative modulation exerted by Hox13 paralogs on Gli3 regulatory sequences. Our results indicate that Hoxa13 acts hierarchically to initiate the formation of digit 1 by reducing Gli3 transcription and by enabling expansion of the 5'Hoxd second expression phase, thereby establishing anterior-posterior asymmetry in the handplate. Our work uncovers a mutual antagonism between Gli3 and Hox13 paralogs that has important implications for Hox and Gli3 gene regulation in the context of development and evolution." {{limb}} | |||
* '''Evolutionary selection and morphological integration in the vertebral column of modern humans'''{{#pmid:31675109|PMID31675109}} "The main objective is to quantify integration, modularity, and response to selection in the presacral vertebral column of modern humans. MATERIALS AND METHODS: Seventeen linear variables on each presacral vertebra were collected in 108 modern humans producing a total of ~39,000 measurements. Then, we studied patterns and magnitudes of integration at regional, vertebral, and intra-vertebral levels. Additionally, we calculated the ability of vertebrae to respond to selection by quantifying differences in evolvability, flexibility, and constraint throughout the spine. The results indicate that caudal vertebrae are more evolvable than those located more cranially in the presacral vertebral column, following an increasing pattern of evolvability from the cervical to the lumbar region. Additionally, the atlas and fifth lumbar vertebra show the lowest values of integration, while central thoracic vertebrae display the highest magnitudes of integration. These results could be related to three main factors: body plan organization expressed by the Hox genes, the strong developmental constraints that determine the number of mammalian vertebrae, and, finally, the functional requirements of an adaptation to bipedal locomotion in the human lineage." | * '''Evolutionary selection and morphological integration in the vertebral column of modern humans'''{{#pmid:31675109|PMID31675109}} "The main objective is to quantify integration, modularity, and response to selection in the presacral vertebral column of modern humans. MATERIALS AND METHODS: Seventeen linear variables on each presacral vertebra were collected in 108 modern humans producing a total of ~39,000 measurements. Then, we studied patterns and magnitudes of integration at regional, vertebral, and intra-vertebral levels. Additionally, we calculated the ability of vertebrae to respond to selection by quantifying differences in evolvability, flexibility, and constraint throughout the spine. The results indicate that caudal vertebrae are more evolvable than those located more cranially in the presacral vertebral column, following an increasing pattern of evolvability from the cervical to the lumbar region. Additionally, the atlas and fifth lumbar vertebra show the lowest values of integration, while central thoracic vertebrae display the highest magnitudes of integration. These results could be related to three main factors: body plan organization expressed by the Hox genes, the strong developmental constraints that determine the number of mammalian vertebrae, and, finally, the functional requirements of an adaptation to bipedal locomotion in the human lineage." | ||
| Line 40: | Line 44: | ||
* '''Analysis of a {{limb}}-specific regulatory element in the promoter of the link protein gene'''{{#pmid:31470976|PMID31470976}} "Link protein is encoded by the Hapln1 gene and is a prototypical protein found in the {{cartilage}} matrix. It acts as an important component of the endochondral skeleton during early development. To study its transcriptional regulation, promoter fragments derived from the link protein gene were coupled to the β-galactosidase reporter and used to study in vivo transgene expression in mice. In day 15.5 mouse embryos, a link promoter fragment spanning -1020 to +40 nucleotides demonstrated highly specific β-galactosidase staining of skeletal structures, including the appendicular and axial cartilaginous tissues. Two shorter promoter fragments, spanning -690 to +40 and -315 to +40 nucleotides, demonstrated limb- and genitalia-specific expression resembling that of homeodomain-regulated tissues. Bioinformatic analysis revealed a highly conserved, Hox-like binding site (HLBS) at approximately -220 bp of the promoter, shared by both constructs, which contained the Hox-core consensus sequence TAATTA. Electromobility shift assays demonstrated binding of Hox-B4 recombinant protein to the HLBS, which was eliminated with nucleotide substitutions within the core-binding element. Co-transfection analysis of the HLBS demonstrated a 22-fold transcriptional activation by HoxA9 expression, which was ablated with a substitution within the core HLBS element. Together these findings establish promoter regions within the link protein gene that are important for in vivo expression and identify the potential role of homeodomain-containing proteins in controlling {{cartilage}} and {{limb}} gene expression." | * '''Analysis of a {{limb}}-specific regulatory element in the promoter of the link protein gene'''{{#pmid:31470976|PMID31470976}} "Link protein is encoded by the Hapln1 gene and is a prototypical protein found in the {{cartilage}} matrix. It acts as an important component of the endochondral skeleton during early development. To study its transcriptional regulation, promoter fragments derived from the link protein gene were coupled to the β-galactosidase reporter and used to study in vivo transgene expression in mice. In day 15.5 mouse embryos, a link promoter fragment spanning -1020 to +40 nucleotides demonstrated highly specific β-galactosidase staining of skeletal structures, including the appendicular and axial cartilaginous tissues. Two shorter promoter fragments, spanning -690 to +40 and -315 to +40 nucleotides, demonstrated limb- and genitalia-specific expression resembling that of homeodomain-regulated tissues. Bioinformatic analysis revealed a highly conserved, Hox-like binding site (HLBS) at approximately -220 bp of the promoter, shared by both constructs, which contained the Hox-core consensus sequence TAATTA. Electromobility shift assays demonstrated binding of Hox-B4 recombinant protein to the HLBS, which was eliminated with nucleotide substitutions within the core-binding element. Co-transfection analysis of the HLBS demonstrated a 22-fold transcriptional activation by HoxA9 expression, which was ablated with a substitution within the core HLBS element. Together these findings establish promoter regions within the link protein gene that are important for in vivo expression and identify the potential role of homeodomain-containing proteins in controlling {{cartilage}} and {{limb}} gene expression." | ||
* Hox genes limit germ cell formation in the short germ insect Gryllus bimaculatus | * Hox genes limit germ cell formation in the short germ insect Gryllus bimaculatus{{#pmid:31346080|PMID31346080}} "Hox genes are conserved transcription factor-encoding genes that specify the identity of body regions in bilaterally symmetrical animals. In the cricket Gryllus bimaculatus, a member of the hemimetabolous insect group Orthoptera, the induction of a subset of mesodermal cells to form the {{primordial germ cell}}s (PGCs) is restricted to the second through the fourth abdominal segments (A2 to A4). In numerous insect species, the Hox genes Sex-combs reduced (Scr), Antennapedia (Antp), Ultrabithorax (Ubx), and abdominal-A (abd-A) jointly regulate the identities of middle and posterior body segments, suggesting that these genes may restrict PGC formation to specific abdominal segments in G. bimaculatus Here we show that reducing transcript levels of some or all of these Hox genes results in supernumerary and/or ectopic PGCs, either individually or in segment-specific combinations, suggesting that the role of these Hox genes is to limit PGC development with respect to their number, segmental location, or both. These data provide evidence of a role for this ancient group of genes in PGC development." | ||
* '''Developmental Dynamics''' special issue January 2014 [http://onlinelibrary.wiley.com/doi/10.1002/dvdy.24098/full Hox/Tale transcription factors in development and disease] | * '''Developmental Dynamics''' special issue January 2014 [http://onlinelibrary.wiley.com/doi/10.1002/dvdy.24098/full Hox/Tale transcription factors in development and disease] | ||
Latest revision as of 11:53, 30 April 2020
| Embryology - 4 May 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
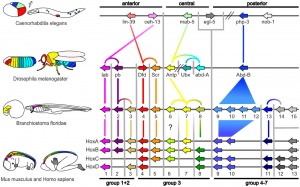
The family of homeobox (Hox) proteins has been a focus of research for over 30 years. In humans, the homeobox gene family contains 300 human homeobox loci, which we divide into 235 probable functional genes and 65 probable pseudogenes.[2]
| Human Hox Family | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
This family of genes were also the basis of the embryo patterning studies that led to the Nobel Prize in Medicine 1995. We now know that in addition to whole embryo axes patterning, this family of genes has many roles in establishing pattern throughout the embryo in different tissues and organs.
There has recently been a revival of an earlier theory[3] that Hox expression during vertebrate pattern formation is linked to the process of segmentation of paraxial mesoderm during somitogenesis.[4]
This signalling pathway has also been implicated in many developmental abnormalities and diseases.
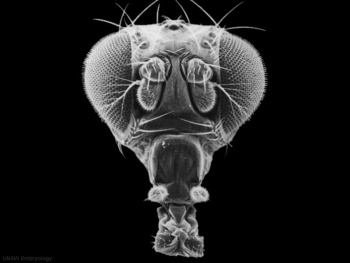
|

|
| Fly wild-type head[5] | Fly antennapedia head[5] |
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Some Recent Findings
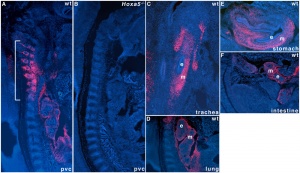
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Hox | antennapedia |
| Older papers |
|---|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
|
Classification
Commonly recognized HOX groupings include: ANTP, PRD, LIM, POU, HNF, SINE, TALE, CUT, PROS and ZF.
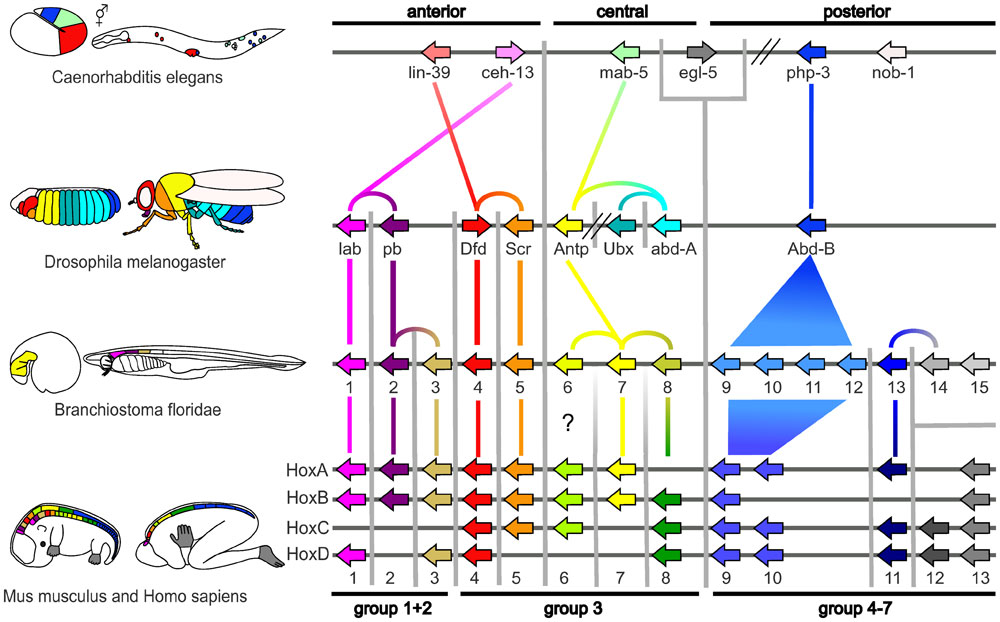
|
| Proposed Hox protein classification[1] |
- PAX2, PAX5 and PAX8 have a partial homeobox
- PAX1 and PAX9 lack a homeobox.
Human Hox
| Table - Human Homeobox Family | ||||
| Class | Subclass | Number of gene families | Number of genes | Number of pseudogenes |
|---|---|---|---|---|
| ANTP | HOXL | 14 | 52 | 0 |
| NKL | 23 | 48 | 19 | |
| PRD | PAX | 3 | 7 | 0 |
| PAXL | 28 | 43 | 24 | |
| LIM | 6 | 12 | 0 | |
| POU | 7 | 16 | 8 | |
| HNF | 2 | 3 | 0 | |
| SINE | 3 | 6 | 0 | |
| TALE | 6 | 20 | 10 | |
| CUT | 3 | 7 | 3 | |
| PROS | 1 | 2 | 0 | |
| ZF | 5 | 14 | 1h | |
| CERS | 1 | 5 | 10 | |
| Totals | 102 | 235 | 65 | |
| Classification data from[2] does not include table supplementary information.
Links: Hox | Hox family | Bmp Family | Fgf Family | Hedgehog Family | Sox Family | Tbx Family | ||||
Chromosomal Distribution of Human Homeobox Genes[2]
Functions
Developmental patterning signal.
Neural
Segmentation
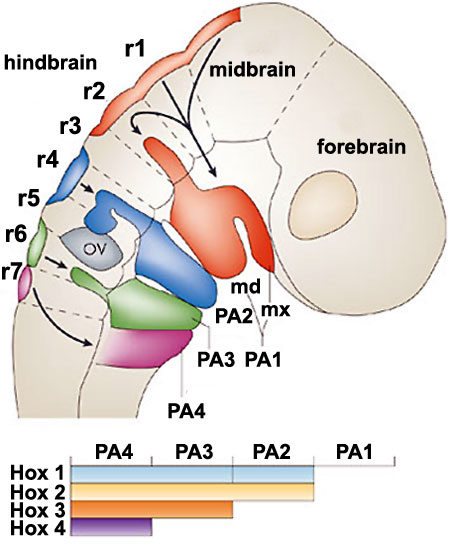
|
Hindbrain neural crest migration and Hox expression pattern[22]
Legend
Adapted by permission from Macmillan Publishers Ltd: Nature Reviews Neuroscience[22], copyright (2007) |
Phrenic Motor Neurons
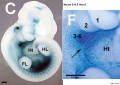
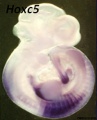
Hox5 (Hoxa5 and Hoxc5) required for phrenic motor column (PMC) development that form the respiratory motor neurons driving the diaphragm for respiration.[23]
Axial Skeleton
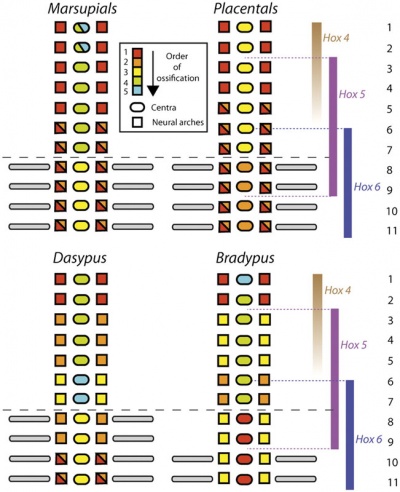
Vertebral element ossification between species.[24] |
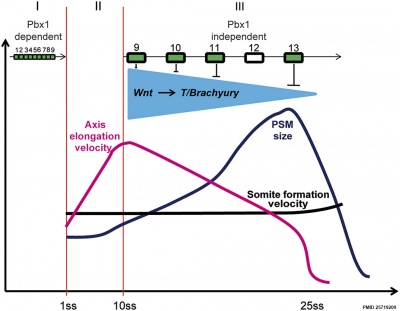
Chicken model of Hox in paraxial mesoderm precursors in the epiblast/tail-bud during axis elongation[25] |
- Links: Somitogenesis | Axial Skeleton Development
Limb
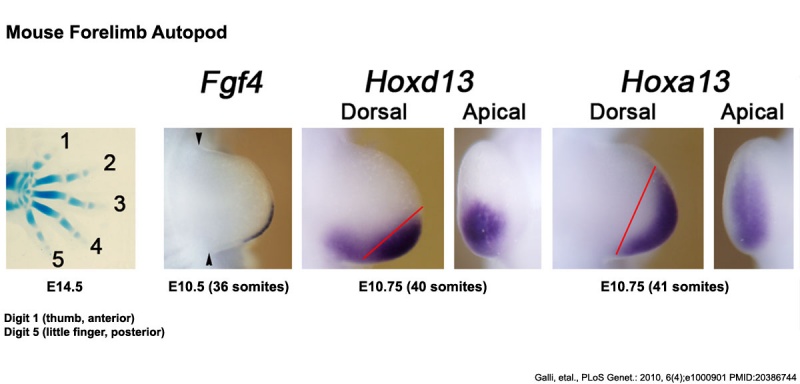
Mouse Limb Patterning Fgf and Hox Expression[26]
Fgf and Hox expression in E10.5 to 10.75 wild-type embryonic forelimb autopod, compared to future E14.5 digit arrangement.
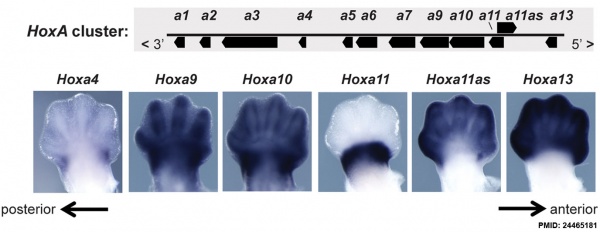
Expression of Hoxa4, Hoxa9, Hoxa10, Hoxa11, Hoxa11 antisense (Hoxa11as), and Hoxa13 in E12.5 limb buds.[27]
- Links: Limb Development
Other
Signaling Pathway
Otx
Otx is a txanscription factor essential for the normal development of the brain, cerebellum, pineal gland, and eye.
OTX is a homeobox family gene related to a gene expressed in the developing Drosophila head termed 'orthodenticle.' OTX transcription factors bind with high affinity to TAATCC/T elements on DNA.
- Links: Pineal | Vision | PubMed - Otx
Additional Images
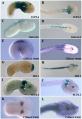
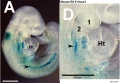
Hoxa3 Mouse E8.5
Hoxa3 Mouse E9.5
Hoxa3 Mouse E10.5
References
- ↑ 1.0 1.1 1.2 Hueber SD, Weiller GF, Djordjevic MA & Frickey T. (2010). Improving Hox protein classification across the major model organisms. PLoS ONE , 5, e10820. PMID: 20520839 DOI.
- ↑ 2.0 2.1 2.2 2.3 Holland PW, Booth HA & Bruford EA. (2007). Classification and nomenclature of all human homeobox genes. BMC Biol. , 5, 47. PMID: 17963489 DOI.
- ↑ Meinhardt H. Models For Biological Pattern Formation. Academic Press; 1982.
- ↑ Gu S, Gu W, Shou J, Xiong J, Liu X, Sun B, Yang D & Xie R. (2017). The Molecular Feature of HOX Gene Family in the Intramedullary Spinal Tumors. Spine , 42, 291-297. PMID: 25785959 DOI.
- ↑ 5.0 5.1 Turner FR & Mahowald AP. (1979). Scanning electron microscopy of Drosophila melanogaster embryogenesis. III. Formation of the head and caudal segments. Dev. Biol. , 68, 96-109. PMID: 108157
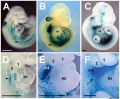
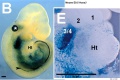
- ↑ 6.0 6.1 Coulombe Y, Lemieux M, Moreau J, Aubin J, Joksimovic M, Bérubé-Simard FA, Tabariès S, Boucherat O, Guillou F, Larochelle C, Tuggle CK & Jeannotte L. (2010). Multiple promoters and alternative splicing: Hoxa5 transcriptional complexity in the mouse embryo. PLoS ONE , 5, e10600. PMID: 20485555 DOI.
- ↑ Kurland M, O'Meara B, Tucker DK & Ackley BD. (2020). The Hox Gene egl-5 Acts as a Terminal Selector for VD13 Development via Wnt Signaling. J Dev Biol , 8, . PMID: 32138237 DOI.
- ↑ Bastida MF, Pérez-Gómez R, Trofka A, Zhu J, Rada-Iglesias A, Sheth R, Stadler HS, Mackem S & Ros MA. (2020). The formation of the thumb requires direct modulation of Gli3 transcription by Hoxa13. Proc. Natl. Acad. Sci. U.S.A. , 117, 1090-1096. PMID: 31896583 DOI.
- ↑ Arlegi M, Veschambre-Couture C & Gómez-Olivencia A. (2019). Evolutionary selection and morphological integration in the vertebral column of modern humans. Am. J. Phys. Anthropol. , , . PMID: 31675109 DOI.
- ↑ Tajima Y, Hozumi A, Yoshida K, Treen N, Sakuma T, Yamamoto T & Sasakura Y. (2019). Hox13 is essential for formation of a sensory organ at the terminal end of the sperm duct in Ciona. Dev. Biol. , , . PMID: 31682808 DOI.
- ↑ Rhodes CS, Matsunobu T & Yamada Y. (2019). Analysis of a limb-specific regulatory element in the promoter of the link protein gene. Biochem. Biophys. Res. Commun. , 518, 672-677. PMID: 31470976 DOI.
- ↑ Barnett AA, Nakamura T & Extavour CG. (2019). Hox genes limit germ cell formation in the short germ insect Gryllus bimaculatus. Proc. Natl. Acad. Sci. U.S.A. , 116, 16430-16435. PMID: 31346080 DOI.
- ↑ Gordon J. (2018). Hox genes in the pharyngeal region: how Hoxa3 controls early embryonic development of the pharyngeal organs. Int. J. Dev. Biol. , 62, 775-783. PMID: 30604847 DOI.
- ↑ Hockman D, Chong-Morrison V, Green SA, Gavriouchkina D, Candido-Ferreira I, Ling ITC, Williams RM, Amemiya CT, Smith JJ, Bronner ME & Sauka-Spengler T. (2019). A genome-wide assessment of the ancestral neural crest gene regulatory network. Nat Commun , 10, 4689. PMID: 31619682 DOI.
- ↑ Acemel RD, Tena JJ, Irastorza-Azcarate I, Marlétaz F, Gómez-Marín C, de la Calle-Mustienes E, Bertrand S, Diaz SG, Aldea D, Aury JM, Mangenot S, Holland PW, Devos DP, Maeso I, Escrivá H & Gómez-Skarmeta JL. (2016). A single three-dimensional chromatin compartment in amphioxus indicates a stepwise evolution of vertebrate Hox bimodal regulation. Nat. Genet. , 48, 336-41. PMID: 26829752 DOI.
- ↑ Hench J, Henriksson J, Abou-Zied AM, Lüppert M, Dethlefsen J, Mukherjee K, Tong YG, Tang L, Gangishetti U, Baillie DL & Bürglin TR. (2015). The Homeobox Genes of Caenorhabditis elegans and Insights into Their Spatio-Temporal Expression Dynamics during Embryogenesis. PLoS ONE , 10, e0126947. PMID: 26024448 DOI.
- ↑ Parker HJ, Bronner ME & Krumlauf R. (2014). A Hox regulatory network of hindbrain segmentation is conserved to the base of vertebrates. Nature , 514, 490-3. PMID: 25219855 DOI.
- ↑ Zigman M, Laumann-Lipp N, Titus T, Postlethwait J & Moens CB. (2014). Hoxb1b controls oriented cell division, cell shape and microtubule dynamics in neural tube morphogenesis. Development , 141, 639-49. PMID: 24449840 DOI.
- ↑ Natale A, Sims C, Chiusano ML, Amoroso A, D'Aniello E, Fucci L, Krumlauf R, Branno M & Locascio A. (2011). Evolution of anterior Hox regulatory elements among chordates. BMC Evol. Biol. , 11, 330. PMID: 22085760 DOI.
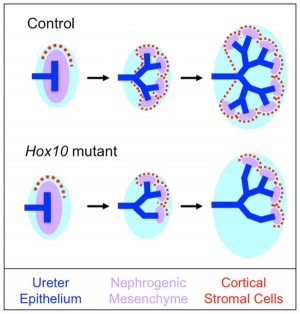
- ↑ Yallowitz AR, Hrycaj SM, Short KM, Smyth IM & Wellik DM. (2011). Hox10 genes function in kidney development in the differentiation and integration of the cortical stroma. PLoS ONE , 6, e23410. PMID: 21858105 DOI.
- ↑ Zhang Y, Huang Q, Cheng JC, Nishi Y, Yanase T, Huang HF & Leung PC. (2010). Homeobox A7 increases cell proliferation by up-regulation of epidermal growth factor receptor expression in human granulosa cells. Reprod. Biol. Endocrinol. , 8, 61. PMID: 20540809 DOI.
- ↑ 22.0 22.1 Guthrie S. (2007). Patterning and axon guidance of cranial motor neurons. Nat. Rev. Neurosci. , 8, 859-71. PMID: 17948031 DOI.
- ↑ Philippidou P, Walsh CM, Aubin J, Jeannotte L & Dasen JS. (2012). Sustained Hox5 gene activity is required for respiratory motor neuron development. Nat. Neurosci. , 15, 1636-44. PMID: 23103965 DOI.
- ↑ Hautier L, Weisbecker V, Sánchez-Villagra MR, Goswami A & Asher RJ. (2010). Skeletal development in sloths and the evolution of mammalian vertebral patterning. Proc. Natl. Acad. Sci. U.S.A. , 107, 18903-8. PMID: 20956304 DOI.
- ↑ Denans N, Iimura T & Pourquié O. (2015). Hox genes control vertebrate body elongation by collinear Wnt repression. Elife , 4, . PMID: 25719209 DOI.
- ↑ Galli A, Robay D, Osterwalder M, Bao X, Bénazet JD, Tariq M, Paro R, Mackem S & Zeller R. (2010). Distinct roles of Hand2 in initiating polarity and posterior Shh expression during the onset of mouse limb bud development. PLoS Genet. , 6, e1000901. PMID: 20386744 DOI.
- ↑ Woltering JM, Noordermeer D, Leleu M & Duboule D. (2014). Conservation and divergence of regulatory strategies at Hox Loci and the origin of tetrapod digits. PLoS Biol. , 12, e1001773. PMID: 24465181 DOI.
Reviews
Mallo M, Wellik DM & Deschamps J. (2010). Hox genes and regional patterning of the vertebrate body plan. Dev. Biol. , 344, 7-15. PMID: 20435029 DOI.
Narita Y & Rijli FM. (2009). Hox genes in neural patterning and circuit formation in the mouse hindbrain. Curr. Top. Dev. Biol. , 88, 139-67. PMID: 19651304 DOI.
Deschamps J & van Nes J. (2005). Developmental regulation of the Hox genes during axial morphogenesis in the mouse. Development , 132, 2931-42. PMID: 15944185 DOI.
Schilling TF & Knight RD. (2001). Origins of anteroposterior patterning and Hox gene regulation during chordate evolution. Philos. Trans. R. Soc. Lond., B, Biol. Sci. , 356, 1599-613. PMID: 11604126 DOI.
Articles
Brauchle M, Bilican A, Eyer C, Bailly X, Martínez P, Ladurner P, Bruggmann R & Sprecher SG. (2018). Xenacoelomorpha Survey Reveals That All 11 Animal Homeobox Gene Classes Were Present in the First Bilaterians. Genome Biol Evol , 10, 2205-2217. PMID: 30102357 DOI.
Rodrigues MF, Esteves CM, Xavier FC & Nunes FD. (2016). Methylation status of homeobox genes in common human cancers. Genomics , 108, 185-193. PMID: 27826049 DOI.
Dunwell TL & Holland PW. (2016). Diversity of human and mouse homeobox gene expression in development and adult tissues. BMC Dev. Biol. , 16, 40. PMID: 27809766 DOI.
Search Pubmed
Search Bookshelf hox
Search Pubmed Now: Hox Homeobox
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- Nobel Prize - Medicine 1995
- OMIM - SONIC HEDGEHOG
- Genetics Home Reference
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, May 4) Embryology Developmental Signals - Homeobox. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Homeobox
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G