Developmental Signals - basic Helix-Loop-Helix: Difference between revisions
(Created page with "{{Header}} ==Introduction== The family of basic Helix-Loop-Helix ({{bHLH}}) transcription factors is encoded in humans between 110 to 125 genes, compared with only 59 in the...") |
mNo edit summary |
||
| (3 intermediate revisions by the same user not shown) | |||
| Line 51: | Line 51: | ||
===Neural=== | ===Neural=== | ||
{{neural}} | |||
===Hearing=== | |||
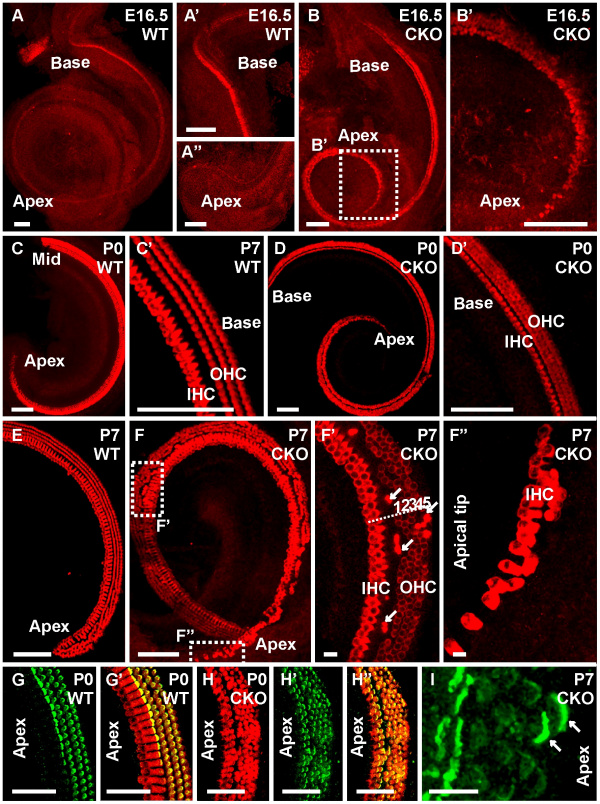
Mouse organ of Corti development requires Neurod1{{#pmid:20661473|PMID20661473}} | |||
[[File:Mouse organ of corti NeuroD1.jpg|600px]] | |||
:Links: {{inner ear}} | {{hearing}} | |||
===Axial Skeleton=== | ===Axial Skeleton=== | ||
===Limb=== | ===Limb=== | ||
{{limb}} | |||
===Other=== | ===Other=== | ||
| Line 81: | Line 93: | ||
{{Footer}} | {{Footer}} | ||
[[Category:bHLH]] | |||
[[Category:Molecular]] [[Category:Pattern]] | [[Category:Molecular]][[Category:Pattern]] | ||
[[Category:Neural]] | [[Category:Neural]] | ||
[[Category:Transcription Factor]] | [[Category:Transcription Factor]] | ||
Latest revision as of 18:33, 30 July 2018
| Embryology - 21 Jun 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
The family of basic Helix-Loop-Helix (bHLH) transcription factors is encoded in humans between 110 to 125 genes, compared with only 59 in the fly (Drosophila).[1] The structural motif describing this transcription factor family refers to the two α-helices connected by a loop. These transcription factors are generally domain are dimeric, with each helix containing basic amino acid residues that aid DNA binding.
This signalling pathway has also been implicated in many developmental abnormalities and diseases.
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Some Recent Findings
|
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Hox <pubmed limit=5>Hox</pubmed> |
| Older papers |
|---|
Classification
Human Genes
| Table - basic Helix Loop Helix Factor Family bHLH | |||||
| Approved Symbol |
Approved Name | Previous Symbols |
Synonyms | Chromosome | |
|---|---|---|---|---|---|
| NEUROD2 | neuronal differentiation 2 | bHLHa1, NDRF | 17q12 | ||
| NEUROD6 | neuronal differentiation 6 | Nex1, Math-2, bHLHa2, NEX1M, Atoh2 | 7p14.3 | ||
| NEUROD1 | neuronal differentiation 1 | NEUROD | MODY6, bHLHa3, NeuroD, BHF-1, BETA2 | 2q31.3 | |
| NEUROD4 | neuronal differentiation 4 | Atoh3, MATH-3, ATH-3, bHLHa4 | 12q13.13 | ||
| NEUROG1 | neurogenin 1 | NEUROD3 | bHLHa6, Math4C, ngn1, AKA | 5q31.1 | |
| NEUROG3 | neurogenin 3 | Math4B, Atoh5, ngn3, bHLHa7 | 10q22.1 | ||
| NEUROG2 | neurogenin 2 | Atoh4, bHLHa8, ngn-2, NGN2, Math4A | 4q25 | ||
| BHLHA9 | basic helix-loop-helix family member a9 | bHLHa9, BHLHF42 | 17p13.3 | ||
| ATOH7 | atonal bHLH transcription factor 7 | Math5, bHLHa13 | 10q21.3 | ||
| ATOH1 | atonal bHLH transcription factor 1 | MATH-1, bHLHa14, Math1, HATH1 | 4q22.2 | ||
| BHLHA15 | basic helix-loop-helix family member a15 | BHLHB8 | MIST1, bHLHa15 | 7q21.3 | |
| TAL1 | TAL bHLH transcription factor 1, erythroid differentiation factor | TCL5 | SCL, bHLHa17 | 1p33 | |
| LYL1 | LYL1, basic helix-loop-helix family member | bHLHa18 | 19p13.13 | ||
| TAL2 | TAL bHLH transcription factor 2 | bHLHa19 | 9q31.2 | ||
| ATOH8 | atonal bHLH transcription factor 8 | bHLHa21, HATH6, FLJ14708 | 2p11.2 | ||
| MSC | musculin | bHLHa22, ABF-1 | 8q13.3 | ||
| TCF21 | transcription factor 21 | bHLHa23, POD1 | 6q23.2 | ||
| TCF23 | transcription factor 23 | bHLHa24, OUT | 2p23.3 | ||
| TCF24 | transcription factor 24 | bHLHa25 | 8q13.1 | ||
| HAND2 | heart and neural crest derivatives expressed 2 | bHLHa26, Thing2, Hed, dHand | 4q34.1 | ||
| HAND1 | heart and neural crest derivatives expressed 1 | Hxt, Thing1, eHand, bHLHa27 | 5q33.2 | ||
| PTF1A | pancreas associated transcription factor 1a | p48, bHLHa29, PTF1-p48 | 10p12.2 | ||
| PERD3L | Fer3 like bHLH transcription factor | N-TWIST, bHLHa31, NATO3 | 7p21.1 | ||
| NHLH2 | nescient helix-loop-helix 2 | HEN2 | bHLHa34, NSCL2 | 1p13.1 | |
| NHLH1 | nescient helix-loop-helix 1 | HEN1 | bHLHa35, NSCL, NSCL1 | 1q23.2 | |
| TWIST1 | twist family bHLH transcription factor 1 | ACS3, BPES3, TWIST, CRS | bHLHa38, H-twist, SCS, CRS1, BPES2 | 7p21.1 | |
| TWIST2 | twist family bHLH transcription factor 2 | bHLHa39, Dermo-1, DERMO1 | 2q37.3 | ||
| TCF15 | transcription factor 15 | bHLHa40, PARAXIS, EC2 | 20p13 | ||
| ASCL3 | achaete-scute family bHLH transcription factor 3 | HASH3, bHLHa42, Sgn1 | 11p15.4 | ||
| ASCL4 | achaete-scute family bHLH transcription factor 4 | bHLHa44, HASH4 | 12q23.3 | ||
| ASCL2 | achaete-scute family bHLH transcription factor 2 | ASH2, bHLHa45, HASH2 | 11p15.5 | ||
| ASCL1 | achaete-scute family bHLH transcription factor 1 | bHLHa46, HASH1, ASH1 | 12q23.2 | ||
| ASCL5 | achaete-scute family bHLH transcription factor 5 | bHLHa47 | 1q32.1 | ||
| SCX | scleraxis bHLH transcription factor | SCXA, SCXB | bHLHa41, bHLHa48 | 8q24.3 | |
| BHLHB9 | basic helix-loop-helix family member b9 | p60TRP, KIAA1701, GASP3 | Xq22.1 | ||
| USF1 | upstream transcription factor 1 | bHLHb11, UEF, MLTFI | 1q23.3 | ||
| USF2 | upstream transcription factor 2, c-fos interacting | FIP, bHLHb12 | 19q13 | ||
| TCF4 | transcription factor 4 | SEF2-1B, ITF2, bHLHb19, E2-2 | 18q21.2 | ||
| TCF12 | transcription factor 12 | HsT17266, p64, HTF4, bHLHb20, HEB | 15q21.3 | ||
| TCF3 | transcription factor 3 | MGC129647, bHLHb21, p75, E2A, VDIR, E47, MGC129648, ITF1 | 19p13.3 | ||
| ID1 | inhibitor of DNA binding 1, HLH protein | bHLHb24, dJ857M17.1.2 | 20q11.21 | ||
| ID3 | inhibitor of DNA binding 3, HLH protein | bHLHb25, HEIR-1 | 1p36.12 | ||
| ID2 | inhibitor of DNA binding 2 | GIG8, bHLHb26 | 2p25.1 | ||
| ID4 | inhibitor of DNA binding 4, HLH protein | bHLHb27 | 6p22.3 | ||
| HEY1 | hes related family bHLH transcription factor with YRPW motif 1 | bHLHb31, CHF-2, HRT-1, HESR1, CHF2, HERP2, HESR-1 | 8q21.13 | ||
| HEY2 | hes related family bHLH transcription factor with YRPW motif 2 | bHLHb32, HESR2, HERP1 | 6q22.31 | ||
| HEYL | hes related family bHLH transcription factor with YRPW motif-like | HEY3, HESR3, bHLHb33 | 1p34.2 | ||
| HES7 | hes family bHLH transcription factor 7 | bHLHb37 | 17p13.1 | ||
| HES5 | hes family bHLH transcription factor 5 | bHLHb38 | 1p36.32 | ||
| HES1 | hes family bHLH transcription factor 1 | HRY | HES-1, Hes1, bHLHb39, FLJ20408 | 3q29 | |
| HES2 | hes family bHLH transcription factor 2 | bHLHb40 | 1p36.31 | ||
| HES6 | hes family bHLH transcription factor 6 | bHLHb41 | 2q37.3 | ||
| HES4 | hes family bHLH transcription factor 4 | bHLHb42 | 1p36.33 | ||
| HES3 | hes family bHLH transcription factor 3 | bHLHb43 | 1p36.31 | ||
| HELT | helt bHLH transcription factor | bHLHb44, HESL, HCM1228, MEGANE, Mgn | 4q35.1 | ||
| MYOD1 | myogenic differentiation 1 | MYF3 | PUM, bHLHc1, MYOD | 11p15.1 | |
| MYF5 | myogenic factor 5 | bHLHc2 | 12q21.31 | ||
| MYOG | myogenin | MYF4 | bHLHc3 | 1q32.1 | |
| MYF6 | myogenic factor 6 | MRF4, bHLHc4 | 12q21.31 | ||
| MESP1 | mesoderm posterior bHLH transcription factor 1 | bHLHc5, MGC10676 | 15q26.1 | ||
| MESP2 | mesoderm posterior bHLH transcription factor 2 | SCDO2, bHLHc6 | 15q26.1 | ||
| FIGLA | folliculogenesis specific bHLH transcription factor | bHLHc8 | 2p13.3 | ||
| MXI1 | MAX interactor 1, dimerization protein | MAD2, MXI, MXD2, bHLHc11 | 10q25.2 | ||
| MXD4 | MAX dimerization protein 4 | bHLHc12, MST149, MSTP149, MAD4 | 4p16.3 | ||
| MXD3 | MAX dimerization protein 3 | bHLHc13, MAD3 | 5q35.3 | ||
| TFAP4 | transcription factor AP-4 | bHLHc41, AP-4 | 16p13.3 | ||
| MXD1 | MAX dimerization protein 1 | MAD | bHLHc58, MAD1 | 2p13.3 | |
| SREBF1 | sterol regulatory element binding transcription factor 1 | SREBP1a, SREBP1, SREBP-1c, bHLHd1 | 17p11.2 | ||
| SREBF2 | sterol regulatory element binding transcription factor 2 | bHLHd2, SREBP2 | 22q13.2 | ||
| MNT | MAX network transcriptional repressor | ROX, MXD6, MAD6, bHLHd3 | 17p13.3 | ||
| MAX | MYC associated factor X | bHLHd7, bHLHd5, bHLHd8, bHLHd6, bHLHd4 | 14q23.3 | ||
| MLX | MLX, MAX dimerization protein | TCFL4 | bHLHd13, MXD7, MAD7 | 17q21.1 | |
| MLXIPL | MLX interacting protein like | WBSCR14 | bHLHd14, MONDOB, MIO, WS-bHLH, CHREBP | 7q11.23 | |
| ARNT2 | aryl hydrocarbon receptor nuclear translocator 2 | KIAA0307, bHLHe1 | 15q25.1 | ||
| ARNT | aryl hydrocarbon receptor nuclear translocator | bHLHe2, HIF-1beta | 1q21.3 | ||
| ARNTL | aryl hydrocarbon receptor nuclear translocator like | PASD3, BMAL1, MOP3, JAP3, bHLHe5 | 11p15.3 | ||
| ARNTL2 | aryl hydrocarbon receptor nuclear translocator like 2 | MOP9, bHLHe6, PASD9, CLIF, BMAL2 | 12p11.23 | ||
| CLOCK | clock circadian regulator | KIAA0334, bHLHe8, KAT13D | 4q12 | ||
| NPAS2 | neuronal PAS domain protein 2 | PASD4, MOP4, bHLHe9 | 2q11.2 | ||
| NPAS1 | neuronal PAS domain protein 1 | bHLHe11, MOP5, PASD5 | 19q13.32 | ||
| NPAS3 | neuronal PAS domain protein 3 | MOP6, bHLHe12, PASD6 | 14q13.1 | ||
| SIM1 | SIM bHLH transcription factor 1 | bHLHe14 | 6q16.3 | ||
| SIM2 | SIM bHLH transcription factor 2 | SIM | MGC119447, bHLHe15 | 21q22.13 | |
| HIF3A | hypoxia inducible factor 3 subunit alpha | MOP7, IPAS, bHLHe17, PASD7 | 19q13.32 | ||
| OLIG2 | oligodendrocyte transcription factor 2 | PRKCBP2, BHLHB1 | OLIGO2, RACK17, bHLHe19 | 21q22.11 | |
| OLIG3 | oligodendrocyte transcription factor 3 | bHLHe20, Bhlhb7 | 6q23.3 | ||
| OLIG1 | oligodendrocyte transcription factor 1 | BHLHB6, bHLHe21 | 21q22.11 | ||
| BHLHE22 | basic helix-loop-helix family member e22 | TNRC20, BHLHB5 | CAGL85, Beta3, bHLHe22 | 8q12.3 | |
| BHLHE23 | basic helix-loop-helix family member e23 | BHLHB4 | bA305P22.3, Beta4, bHLHe23 | 20q13.33 | |
| MITF | melanogenesis associated transcription factor | WS2A, WS2 | bHLHe32, MI | 3p13 | |
| TFE3 | transcription factor binding to IGHM enhancer 3 | TFEA, bHLHe33 | Xp11.23 | ||
| TFEC | transcription factor EC | bHLHe34, TCFEC, TFECL | 7q31.2 | ||
| TFEB | transcription factor EB | bHLHe35, TCFEB | 6p21.1 | ||
| MLXIP | MLX interacting protein | MIR, KIAA0867, bHLHe36, MONDOA | 12q21.31 | ||
| MYCN | MYCN proto-oncogene, bHLH transcription factor | NMYC | bHLHe37, N-myc, MYCNOT | 2p24.3 | |
| MYCL | MYCL proto-oncogene, bHLH transcription factor | MYCL1 | bHLHe38, LMYC | 1p34.2 | |
| MYC | MYC proto-oncogene, bHLH transcription factor | MYCC, bHLHe39, c-Myc | 8q24.21 | ||
| BHLHE40 | basic helix-loop-helix family member e40 | STRA13, BHLHB2 | DEC1, bHLHe40, SHARP2, Clast5 | 3p26.1 | |
| BHLHE41 | basic helix-loop-helix family member e41 | BHLHB3 | DEC2, SHARP-1, SHARP1, bHLHe41 | 12p12.1 | |
| NCOA3 | nuclear receptor coactivator 3 | SRC-3, KAT13B, CAGH16, ACTR, p/CIP, bHLHe42, TRAM-1, AIB1, TNRC16, SRC3, RAC3 | 20q13.12 | ||
| EPAS1 | endothelial PAS domain protein 1 | HLF, HIF2A, bHLHe73, MOP2, PASD2 | 2p21 | ||
| NCOA1 | nuclear receptor coactivator 1 | RIP160, bHLHe74, NCoA-1, F-SRC-1, SRC1, KAT13A | 2p23.3 | ||
| NCOA2 | nuclear receptor coactivator 2 | TIF2, bHLHe75, NCoA-2, GRIP1, KAT13C | 8q13.3 | ||
| AHR | aryl hydrocarbon receptor | bHLHe76 | 7p21.1 | ||
| AHRR | aryl-hydrocarbon receptor repressor | AHH, AHHR | KIAA1234, bHLHe77 | 5p15.33 | |
| HIF1A | hypoxia inducible factor 1 subunit alpha | bHLHe78, HIF1, HIF-1alpha, MOP1, PASD8 | 14q23.2 | ||
| NPAS4 | neuronal PAS domain protein 4 | bHLHe79, Le-PAS, NXF, PASD10 | 11q13.2 | ||
| SOHLH1 | spermatogenesis and oogenesis specific basic helix-loop-helix 1 | C9orf157 | NOHLH, bA100C15.3, SPATA27, bHLHe80, TEB2 | 9q34.3 | |
| SOHLH2 | spermatogenesis and oogenesis specific basic helix-loop-helix 2 | TEB1, SPATA28, bHLHe81, FLJ20449 | 13q13.3 | ||
| TCFL5 | transcription factor like 5 | CHA, E2BP-1, bHLHe82, Figlb | 20q13.33 | ||
| Links: Developmental Signals - basic Helix-Loop-Helix | HGNC | Bmp Family | Fgf Family | Sox Family | Tbx Family | |||||
| Human bHLH Family | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Functions
Developmental patterning signal.
Neural
Hearing
Mouse organ of Corti development requires Neurod1[2]
Axial Skeleton
Limb
Other
Signaling Pathway
References
- ↑ Baker NE & Brown NL. (2018). All in the family: proneural bHLH genes and neuronal diversity. Development , 145, . PMID: 29720483 DOI.
- ↑ Jahan I, Pan N, Kersigo J & Fritzsch B. (2010). Neurod1 suppresses hair cell differentiation in ear ganglia and regulates hair cell subtype development in the cochlea. PLoS ONE , 5, e11661. PMID: 20661473 DOI.
Reviews
Articles
Search Pubmed
Search Bookshelf bHLH
Search Pubmed Now: bHLH basic helix loop helix
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, June 21) Embryology Developmental Signals - basic Helix-Loop-Helix. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_basic_Helix-Loop-Helix
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G