Developmental Signals - Bone Morphogenetic Protein
| Embryology - 9 Jun 2026 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
Belongs to the transforming growth factor-beta (TGFB) superfamily, humans have 15 members in this protein signaling family. The proteins are synthesized as prepropeptides, then cleaved, and processed into dimeric proteins.
- TGFB family members: TGFB1, TGFB, TGFB3, bone morphogenetic proteins Bmp-2A, Bmp-2B, Bmp-3, and Bmp-6. mullerian inhibitory substance.
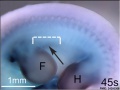
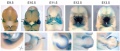
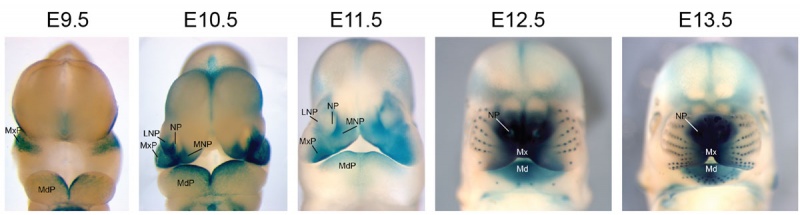
Mouse Bmp4 expression face.[1]
- BMP Mouse Links: Face and limb E9.5-13.5 | Face E9.5-13.5 | Body E11.0 | Body E11.5 | BMP | Mouse Development
Growth Differentiation Factor-6 (Gdf6) is a member of the Bone Morphogenetic Protein (BMP) family of secreted signaling molecules.
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Bone Morphogenetic Protein | BMP | BMP4 | BMP7 | BMP15 |
| Older papers |
|---|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
|
Human BMP Family
| Table - Human Bmp Family | ||||
| Approved Symbol |
Approved Name | Previous Symbols |
Synonyms | Chromosome |
|---|---|---|---|---|
| BMP1 | bone morphogenetic protein 1 | PCOLC | 8p21.3 | |
| BMP2 | bone morphogenetic protein 2 | BMP2A | 20p12.3 | |
| BMP3 | bone morphogenetic protein 3 | 4q21.21 | ||
| BMP4 | bone morphogenetic protein 4 | BMP2B | 14q22.2 | |
| BMP5 | bone morphogenetic protein 5 | 6p12.1 | ||
| BMP6 | bone morphogenetic protein 6 | VGR | VGR1 | 6p24.3 |
| BMP7 | bone morphogenetic protein 7 | OP-1 | 20q13.31 | |
| BMP8A | bone morphogenetic protein 8a | 1p34.3 | ||
| BMP8B | bone morphogenetic protein 8b | BMP8 | OP-2 | 1p34.2 |
| BMP10 | bone morphogenetic protein 10 | 2p13.3 | ||
| BMP15 | bone morphogenetic protein 15 | GDF9B | Xp11.22 | |
| Links: Developmental Signals - Bone Morphogenetic Protein | OMIM BMP2 | HGNC | Bmp Family | Sox Family | Tbx Family | ||||
| Human BMP Family | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Structure
| BMP Superfamily Canonical Signalling | |
|---|---|
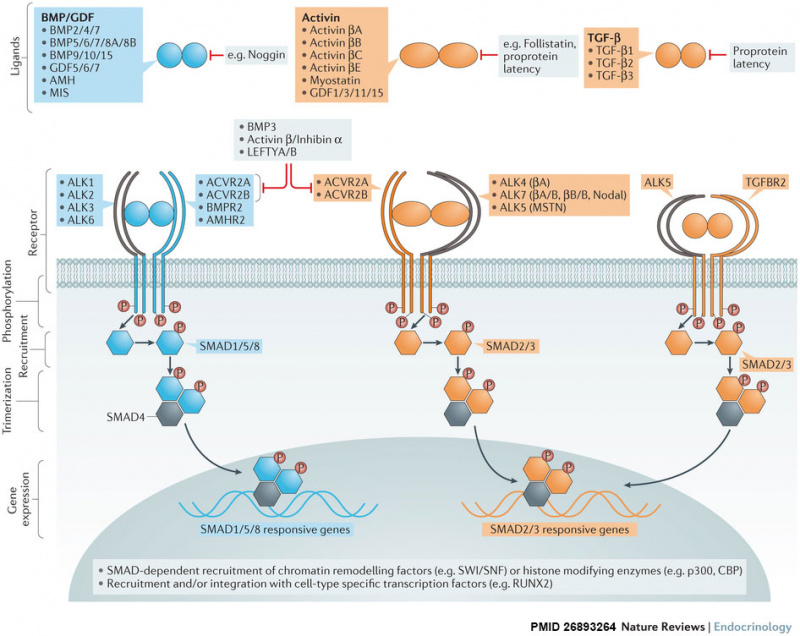
| |
| Over 30 bone morphogenetic protein (BMP) superfamily ligands have been discovered in humans. Most are secreted as mature disulfide-linked dimers, with the exception of TGF-β1, TGF-β2 and TGF-β3, which can be secreted in a latent form and require proteolytic activation. BMPs signal through a multimeric cell surface complex consisting of two type I receptors and two type II receptors.
Activated type I receptors recruit and phosphorylate pathway-specific R-SMADs (SMAD1, SMAD5 and SMAD8 (blue pathway), and SMAD2 and SMAD3 (orange pathway)), which can form trimers with SMAD4 and translocate to the nucleus. SMADs have intrinsic DNA-binding activity and are able to regulate gene expression by recruitment of chromatin-remodelling machinery and integration with tissue-specific transcription factors. SMAD8 is also known as SMAD9. The pathway can be antagonized by many mechanisms including neutralization of ligands by secreted traps such as noggin or follistatin, secretion of latent ligands bound to their propeptides, or via titration of receptors by nonsignalling ligands such as BMP3, activin β/inhibin α dimers or LEFTY monomers. |
|
Gene
Function
BMP4
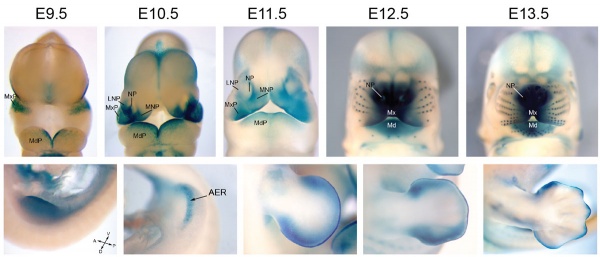
Mouse Bmp4 expression limb and face.[1] |
|
Neural Development
During gastrulation the BMP pathway is antagonised and involved with neural induction. Neural induction signaling through the BMP-regulated Smad1/5 proteins appears to be controlled by fibroblast growth factor (FGF)-regulated Ca2+ entry activating calcineurin (CaN) that in turn dephosphorylates Smad1/5 proteins.[13]
- Links: neural
Genital Development
Bmp4 is an essential growth factor for the initiation of Template:Genital tubercle (GT) outgrowth[6]
- "The external genitalia are appendage organs outgrowing from the posterior body trunk. Murine genital tubercle (GT), anlage of external genitalia, initiates its outgrowth from embryonic day (E) E10.5 as a bud structure. Several growth factors such as fibroblast growth factor (FGF), Wnt and Sonic hedgehog (Shh) are essential for the GT outgrowth. However, the mechanisms of initiation of GT outgrowth are poorly understood. We previously identified bone morphogenetic protein (Bmp) signaling as a negative regulator for GT outgrowth. We show here novel aspects of Bmp4 functions for GT outgrowth. We identified the Bmp4 was already expressed in cloaca region at E9.5, before GT outgrowth. To analyze the function of Bmp4 at early stage for the initiation of GT outgrowth, we utilized the Hoxa3-Cre driver and Bmp4 flox/flox mouse lines. Hoxa3 Cre/+ ; Bmp4 flox/flox mutant mice showed the hypoplasia of GT with reduced expression of outgrowth promoting genes such as Wnt5a, Hoxd13 and p63, whereas Shh expression was not affected. Formation of distal urethral epithelium (DUE) marked by the Fgf8 expression is essential for controlling mesenchymal genes expression in GT and subsequent its outgrowth. Furthermore, Fgf8 expression was dramatically reduced in such mutant mice indicating the defective DUE formation. Hence, current results indicate that Bmp4 is an essential growth factor for the initiation of GT outgrowth independent of Shh signaling. Thus, Bmp4 positively regulates for the formation of DUE. The current study provides new insights into the function of Bmp signaling at early stage for the initiation of GT outgrowth."
- Links: genital
BMP15
Oocyte Development
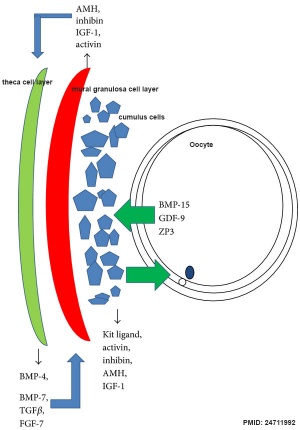
|
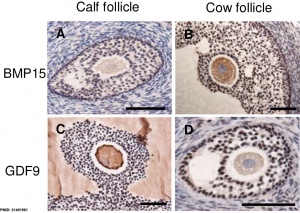
|
| Molecular paracrine interactions involving BMP15 signaling[14] | Localization of BMP15 in calf and cow follicles[15] |
- Links: Oocyte Development
Limb Development
Bmp2, Bmp4 and Bmp7 are co-required in the mouse AER for normal digit patterning but not limb outgrowth[16]
- "Outgrowth and patterning of the vertebrate limb requires a functional apical ectodermal ridge (AER). The AER is a thickening of ectodermal tissue located at the distal end of the limb bud. Loss of this structure, either through genetic or physical manipulations results in truncation of the limb. A number of genes, including Bmps, are expressed in the AER. Previously, it was shown that removal of the BMP receptor Bmpr1a specifically from the AER resulted in complete loss of hindlimbs suggesting that Bmp signaling in the AER is required for limb outgrowth. In this report, we genetically removed the three known AER-expressed Bmp ligands, Bmp2, Bmp4 and Bmp7 from the AER of the limb bud using floxed conditional alleles and the Msx2-cre allele. Surprisingly, only defects in digit patterning and not limb outgrowth were observed. In triple mutants, the anterior and posterior AER was present but loss of the central region of the AER was observed. These data suggest that Bmp ligands expressed in the AER are not required for limb outgrowth but instead play an essential role in maintaining the AER and patterning vertebrate digits."
- Links: Limb Development
Blood Vessel Development
BMP/SMAD signaling pathway regulates angiogenic sprouting and is involved in embryo vascular development.
Signaling Pathway
Identified BMP modulators:[17] Noggin, Chordin, Chordin-like 1, Chordin-like 2, Twisted gastrulation, Dan, BMPER, Sost, Sostdc1, Follistatin, Follistatin-like 1, Follistatin-like 5 and Tolloid.
Receptor
Intracellular Signaling
SNW-domain containing protein 1
(SNW1, SKI-INTERACTING PROTEIN; SKIIP)
A protein that interacts with nuclear receptors and enhances ligand-activated transcription, also called a nuclear receptor co-activator.
Regulator of Spatial BMP Activity, Neural Plate Border Formation, and Neural Crest Specification in Vertebrate Embryos[18]
- "Bone morphogenetic protein (BMP) gradients provide positional information to direct cell fate specification, such as patterning of the vertebrate ectoderm into neural, neural crest, and epidermal tissues, with precise borders segregating these domains. However, little is known about how BMP activity is regulated spatially and temporally during vertebrate development to contribute to embryonic patterning, and more specifically to neural crest formation. Through a large-scale in vivo functional screen in Xenopus for neural crest fate, we identified an essential regulator of BMP activity, SNW1. SNW1 is a nuclear protein known to regulate gene expression. Using antisense morpholinos to deplete SNW1 protein in both Xenopus and zebrafish embryos, we demonstrate that dorsally expressed SNW1 is required for neural crest specification, and this is independent of mesoderm formation and gastrulation morphogenetic movements. By exploiting a combination of immunostaining for phosphorylated Smad1 in Xenopus embryos and a BMP-dependent reporter transgenic zebrafish line, we show that SNW1 regulates a specific domain of BMP activity in the dorsal ectoderm at the neural plate border at post-gastrula stages. We use double in situ hybridizations and immunofluorescence to show how this domain of BMP activity is spatially positioned relative to the neural crest domain and that of SNW1 expression. Further in vivo and in vitro assays using cell culture and tissue explants allow us to conclude that SNW1 acts upstream of the BMP receptors. Finally, we show that the requirement of SNW1 for neural crest specification is through its ability to regulate BMP activity, as we demonstrate that targeted overexpression of BMP to the neural plate border is sufficient to restore neural crest formation in Xenopus SNW1 morphants. We conclude that through its ability to regulate a specific domain of BMP activity in the vertebrate embryo, SNW1 is a critical regulator of neural plate border formation and thus neural crest specification."
Additional Images
OMIM
About OMIM "Online Mendelian Inheritance in Man OMIM is a comprehensive, authoritative, and timely compendium of human genes and genetic phenotypes. The full-text, referenced overviews in OMIM contain information on all known mendelian disorders and over 12,000 genes. OMIM focuses on the relationship between phenotype and genotype. It is updated daily, and the entries contain copious links to other genetics resources." OMIM
- Links: OMIM300247
Abnormalities
Infertility
| Gene abbreviation |
Name | Gene Location | Online Mendelian Inheritance in Man (OMIM) |
HUGO Gene Nomenclature Committee (HGNC) |
GeneCards (GCID) | Diagnosis |
|---|---|---|---|---|---|---|
| BMP15 | Bone morphogenetic protein 15 | Xp11.22 | 300247 | 1068 | GC0XP050910 | Primary ovarian insufficiency |
| Table data source[19] (table 1) Links: fertilization | oocyte | ovary | | Female Infertility Genes | spermatozoa | testis | Male Infertility Genes | Genetic Abnormalities | ART Primary ovarian insufficiency - depletion or dysfunction of ovarian follicles with cessation of menses before age 40 years. | ||||||
| Female Infertility Genes | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
References
- ↑ 1.0 1.1 Jumlongras D, Lachke SA, O'Connell DJ, Aboukhalil A, Li X, Choe SE, Ho JW, Turbe-Doan A, Robertson EA, Olsen BR, Bulyk ML, Amendt BA & Maas RL. (2012). An evolutionarily conserved enhancer regulates Bmp4 expression in developing incisor and limb bud. PLoS ONE , 7, e38568. PMID: 22701669 DOI.
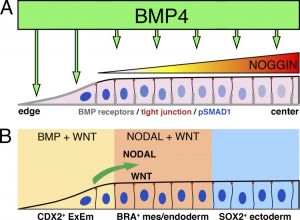
- ↑ 2.0 2.1 Taniguchi K, Heemskerk I & Gumucio DL. (2019). Opening the black box: Stem cell-based modeling of human post-implantation development. J. Cell Biol. , 218, 410-421. PMID: 30552099 DOI.
- ↑ Mao A, Zhang M, Li L, Liu J, Ning G, Cao Y & Wang Q. (2021). Pharyngeal pouches provide a niche microenvironment for arch artery progenitor specification. Development , 148, . PMID: 33334861 DOI.
- ↑ Riepsamen AH, Chan K, Lien S, Sweeten P, Donoghoe MW, Walker G, Fraison EHJ, Stocker WA, Walton KL, Harrison CA, Ledger WL, Robertson DM & Gilchrist RB. (2019). Serum concentrations of oocyte-secreted factors BMP15 and GDF9 during IVF and in women with reproductive pathologies. Endocrinology , , . PMID: 31211369 DOI.
- ↑ Su YH, Chen YC, Ting HC, Fan TP, Lin CY, Wang KT & Yu JK. (2019). BMP controls dorsoventral and neural patterning in indirect-developing hemichordates providing insight into a possible origin of chordates. Proc. Natl. Acad. Sci. U.S.A. , , . PMID: 31189599 DOI.
- ↑ 6.0 6.1 Kajioka D, Suzuki K, Nakada S, Matsushita S, Miyagawa S, Takeo T, Nakagata N & Yamada G. (2019). Bmp4 is an essential growth factor for the initiation of genital tubercle (GT) outgrowth. Congenit Anom (Kyoto) , , . PMID: 30714224 DOI.
- ↑ Row RH, Pegg A, Kinney B, Farr GH, Maves L, Lowell S, Wilson V & Martin BL. (2018). BMP and FGF signaling interact to pattern mesoderm by controlling basic helix-loop-helix transcription factor activity. Elife , 7, . PMID: 29877796 DOI.
- ↑ Mimura S, Suga M, Okada K, Kinehara M, Nikawa H & Furue MK. (2016). Bone morphogenetic protein 4 promotes craniofacial neural crest induction from human pluripotent stem cells. Int. J. Dev. Biol. , 60, 21-8. PMID: 26934293 DOI.
- ↑ Bier E & De Robertis EM. (2015). EMBRYO DEVELOPMENT. BMP gradients: A paradigm for morphogen-mediated developmental patterning. Science , 348, aaa5838. PMID: 26113727 DOI.
- ↑ Xu PF, Houssin N, Ferri-Lagneau KF, Thisse B & Thisse C. (2014). Construction of a vertebrate embryo from two opposing morphogen gradients. Science , 344, 87-9. PMID: 24700857 DOI.
- ↑ Miletich I, Yu WY, Zhang R, Yang K, Caixeta de Andrade S, Pereira SF, Ohazama A, Mock OB, Buchner G, Sealby J, Webster Z, Zhao M, Bei M & Sharpe PT. (2011). Developmental stalling and organ-autonomous regulation of morphogenesis. Proc. Natl. Acad. Sci. U.S.A. , 108, 19270-5. PMID: 22084104 DOI.
- ↑ Salazar VS, Gamer LW & Rosen V. (2016). BMP signalling in skeletal development, disease and repair. Nat Rev Endocrinol , 12, 203-21. PMID: 26893264 DOI.
- ↑ Cho A, Tang Y, Davila J, Deng S, Chen L, Miller E, Wernig M & Graef IA. (2014). Calcineurin signaling regulates neural induction through antagonizing the BMP pathway. Neuron , 82, 109-124. PMID: 24698271 DOI.
- ↑ Chronowska E. (2014). High-throughput analysis of ovarian granulosa cell transcriptome. Biomed Res Int , 2014, 213570. PMID: 24711992 DOI.
- ↑ Hosoe M, Kaneyama K, Ushizawa K, Hayashi KG & Takahashi T. (2011). Quantitative analysis of bone morphogenetic protein 15 (BMP15) and growth differentiation factor 9 (GDF9) gene expression in calf and adult bovine ovaries. Reprod. Biol. Endocrinol. , 9, 33. PMID: 21401961 DOI.
- ↑ Choi KS, Lee C, Maatouk DM & Harfe BD. (2012). Bmp2, Bmp4 and Bmp7 are co-required in the mouse AER for normal digit patterning but not limb outgrowth. PLoS ONE , 7, e37826. PMID: 22662233 DOI.
- ↑ Lorda-Diez CI, Montero JA, Rodriguez-Leon J, Garcia-Porrero JA & Hurle JM. (2013). Expression and functional study of extracellular BMP antagonists during the morphogenesis of the digits and their associated connective tissues. PLoS ONE , 8, e60423. PMID: 23573253 DOI.
- ↑ Wu MY, Ramel MC, Howell M & Hill CS. (2011). SNW1 is a critical regulator of spatial BMP activity, neural plate border formation, and neural crest specification in vertebrate embryos. PLoS Biol. , 9, e1000593. PMID: 21358802 DOI.
- ↑ 19.0 19.1 Harper JC, Aittomäki K, Borry P, Cornel MC, de Wert G, Dondorp W, Geraedts J, Gianaroli L, Ketterson K, Liebaers I, Lundin K, Mertes H, Morris M, Pennings G, Sermon K, Spits C, Soini S, van Montfoort APA, Veiga A, Vermeesch JR, Viville S & Macek M. (2018). Recent developments in genetics and medically assisted reproduction: from research to clinical applications. Eur. J. Hum. Genet. , 26, 12-33. PMID: 29199274 DOI.
Reviews
Schliermann A & Nickel J. (2018). Unraveling the Connection between Fibroblast Growth Factor and Bone Morphogenetic Protein Signaling. Int J Mol Sci , 19, . PMID: 30340367 DOI.
Salazar VS, Gamer LW & Rosen V. (2016). BMP signalling in skeletal development, disease and repair. Nat Rev Endocrinol , 12, 203-21. PMID: 26893264 DOI.
Bier E & De Robertis EM. (2015). EMBRYO DEVELOPMENT. BMP gradients: A paradigm for morphogen-mediated developmental patterning. Science , 348, aaa5838. PMID: 26113727 DOI.
Jain AP, Pundir S & Sharma A. (2013). Bone morphogenetic proteins: The anomalous molecules. J Indian Soc Periodontol , 17, 583-6. PMID: 24174749 DOI.
Ruschke K, Hiepen C, Becker J & Knaus P. (2012). BMPs are mediators in tissue crosstalk of the regenerating musculoskeletal system. Cell Tissue Res. , 347, 521-44. PMID: 22327483 DOI.
Gazzerro E & Canalis E. (2006). Bone morphogenetic proteins and their antagonists. Rev Endocr Metab Disord , 7, 51-65. PMID: 17029022 DOI.
Chen D, Zhao M & Mundy GR. (2004). Bone morphogenetic proteins. Growth Factors , 22, 233-41. PMID: 15621726 DOI.
Articles
Taniguchi K, Heemskerk I & Gumucio DL. (2019). Opening the black box: Stem cell-based modeling of human post-implantation development. J. Cell Biol. , 218, 410-421. PMID: 30552099 DOI.
Cho A, Tang Y, Davila J, Deng S, Chen L, Miller E, Wernig M & Graef IA. (2014). Calcineurin signaling regulates neural induction through antagonizing the BMP pathway. Neuron , 82, 109-124. PMID: 24698271 DOI.
Luo YJ & Su YH. (2012). Opposing nodal and BMP signals regulate left-right asymmetry in the sea urchin larva. PLoS Biol. , 10, e1001402. PMID: 23055827 DOI.
Kuypers E, Collins JJ, Jellema RK, Wolfs TG, Kemp MW, Nitsos I, Pillow JJ, Polglase GR, Newnham JP, Germeraad WT, Kallapur SG, Jobe AH & Kramer BW. (2012). Ovine fetal thymus response to lipopolysaccharide-induced chorioamnionitis and antenatal corticosteroids. PLoS ONE , 7, e38257. PMID: 22693607 DOI.
Online Textbooks
- Molecular Biology of the Cell. 4th edition. Alberts B, Johnson A, Lewis J, et al.New York: Garland Science; 2002. Signal Proteins of the TGF-β Superfamily Act Through Receptor Serine/Threonine Kinases and Smads
- Madame Curie Bioscience Database (Internet). Austin (TX): Landes Bioscience; 2000. TGFβ-dependent Epithelial-Mesenchymal Transition
Search PubMed
- Search PubMed: Bone Morphogenetic Protein
- All Databases: Bone Morphogenetic Protein
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- NCBI - Mouse BMP4 gene
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2026, Haziran 9) Embryology Developmental Signals - Bone Morphogenetic Protein. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Bone_Morphogenetic_Protein
- © Dr Mark Hill 2026, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G