Blastocyst Development
| Embryology - 23 Apr 2026 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
(Greek, blastos = sprout + cystos = cavity) or blastula, the term used to describe the hollow cellular mass that forms in early development.
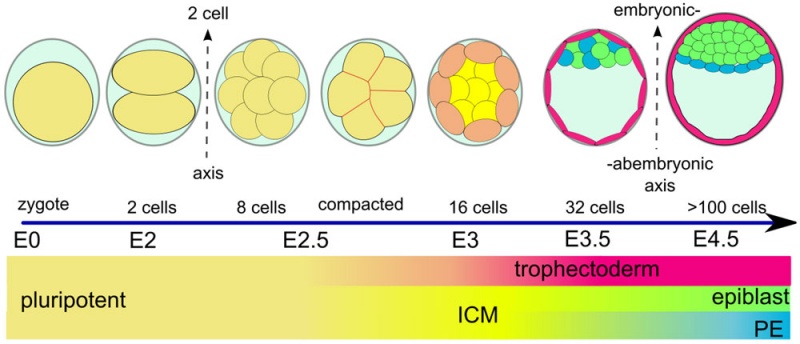
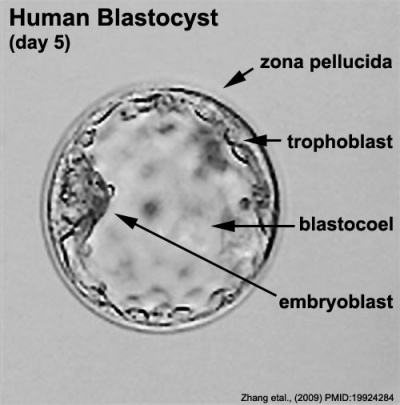
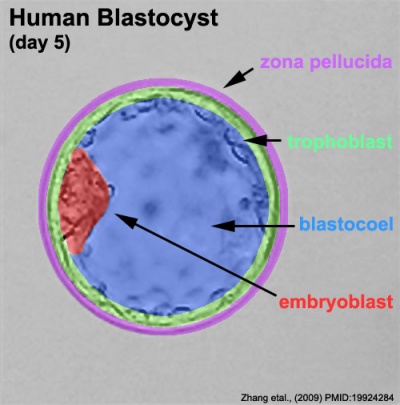
The blastocyst consists of cells forming an outer trophectoderm (TE, trophoblast) layer, an inner cell mass (ICM, embryo blast) and a blastocoel (fluid-filled cavity). The inner cell mass will form the entire embryo, and is the source of true embryonic stem cells capable of forming all cell types within the embryo. In mammals, the trophectoderm will form key cells (trophoblast) of the fatal component of the placenta.
In humans, blastocyst stage of development occurs during the first and second week following fertilization (GA week 3 and 4) and is described initially as Carnegie stage 3. This stage is followed by blastocyst hatching and implantation.
One of the first key developmental patterning decisions in the morula to blastocyst development is TE or ICM cell fate. In the mouse model, Hippo[2](TEAD4) and Notch[3](Cdx2) together appear regulate this early fate decision.
| Blastocyst Links: blastocyst | morula | fertilization | Week 1 | trophoblast | implantation | Human Day 3-6 Movie | Mouse Model Movie | Category:Blastocyst |
| Molecular: Hippo | Notch |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Blastocyst Development | Blastocyst | Blastocoel | Inner Cell Mass | | Trophectoderm |
| Older papers |
|---|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
|
Movies
Human Blastocyst
|
|
|
Model Development
|
|
A recent review{{#pmid28681376|PMID28681376}} of the initial morula to blastocyst formation, based upon animal models, identifies important mechanical steps:
- Compaction - morula blastomeres packing tightly (microfilament cytoskeleton)
- Cleavage planes - spindle direction of dividing cells (mitosis).
- Polarisation - blastomeres apico-basal (Hippo pathway, Yes-associated protein (Yap)
- Cavitation - blastocoel formation with cycles of cavity expansion and collapse.
- epiblast cells - contact the polar trophectoderm
- primitive endoderm - facing the cavity
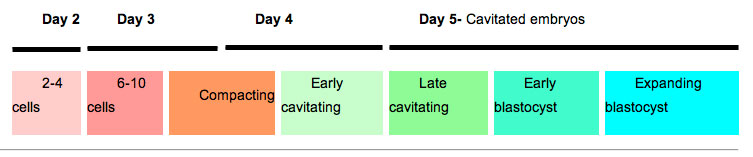
Human Blastocyst Formation
The table below shows human blastocyst in vitro development changes during week 1.[14]
Labeled Blastocyst
Blastocyst Hatching
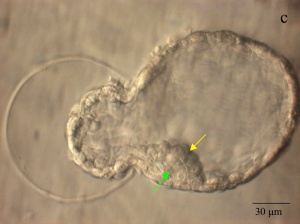
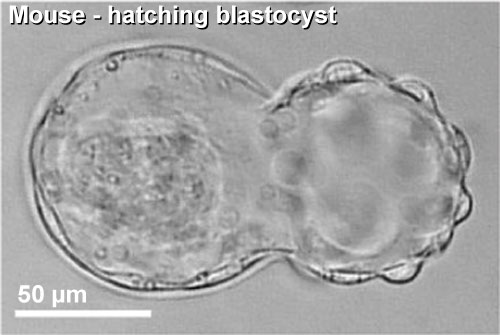
|
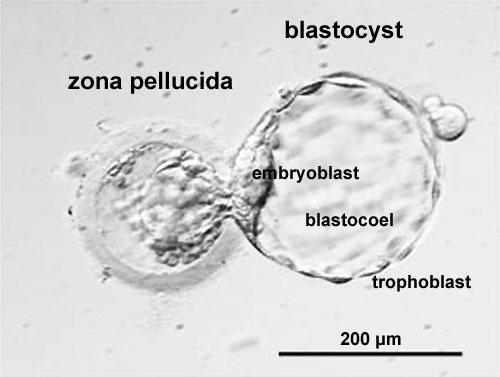
|
| Blastocyst hatching from zona pellucida (mouse) | Blastocyst hatching from zona pellucida (human) |
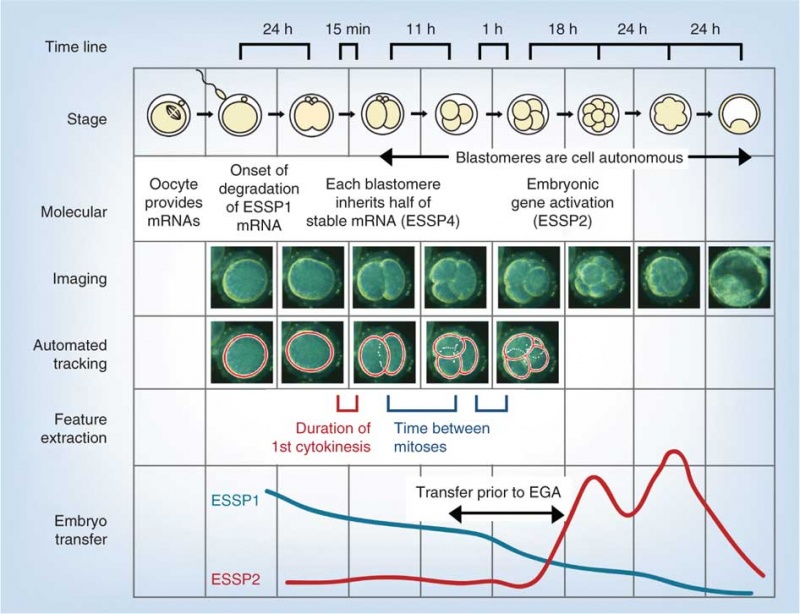
Model Human Blastocyst Development
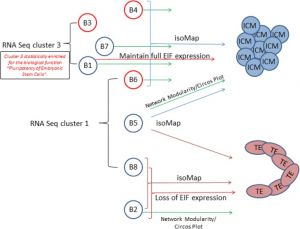
The following figure is from a recent study[11] using video and genetic analysis of in vitro human development during week 1 following fertilization.
- EGA - embryonic genome activation
- ESSP - embryonic stage–specific pattern, four unique embryonic stage–specific patterns (1-4)
- Links: Figure with legend
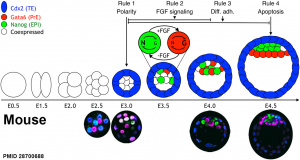
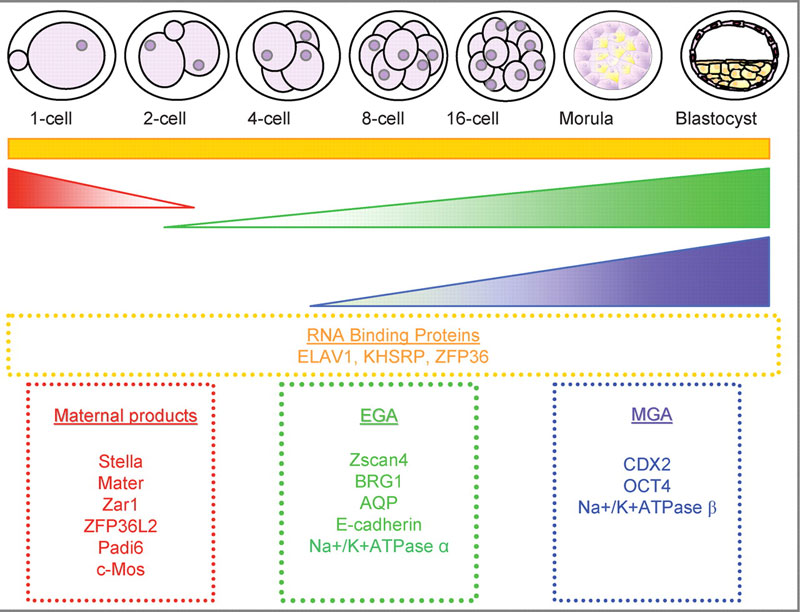
Mouse Blastocyst Gene Expression
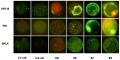
General gene expression patterns are indicated from genomic profiling.[16]
- red - loss of maternal mRNAs
- green - activation of embryonic genome (EGA)
- purple - maternal gene activation (MGA)
- orange - continuous expression
Inner Cell Mass
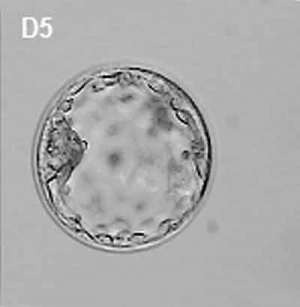
The inner cell mass forms an inner layer of larger cells is also called the "embryoblast" is a cluster of cells located and attached on one wall of the outer trophoblast layer. In week 2 this mass will differentiate into two distinct layers the epiblast and hypoblast.
The hypoblast (or primitive endoderm) is a transient epithelial layer facing towards the blastoceol, it is replaced in week three by the gastrulation migrating endoderm cells.
The epiblast layer will form the entire embryo and undergoes gastrulation in week three to form the 3 germ layers. It also forms an epithelial layer lining the amniotic cavity.
Trophectoderm
The trophectoderm (TE) outer layer of smaller cells is also called the "trophoblast" epithelium, that will later form a key component of the placenta. A key early function is for the transport of sodium (Na+) and chloride (Cl-) ions through this layer into the blastocoel. Later in week 2 this layer will differentiate into two distinct trophoblast layers the syncytiotrophoblast and cytotrophoblast cells and are key to implantation and early placentation.
Differentiation of the early layer has been shown to be regulated by the transcription factors Tead4[17] and then Caudal-related homeobox 2 (Cdx2).
- Links: trophoblast | OMIM -Tead4 | OMIM - Cdx2
Blastocoel
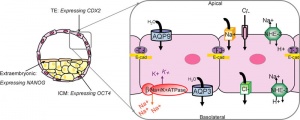
- trophectoderm transports of Na+ and Cl- ions through this layer into the blastocoel
- generates an osmotic gradient driving fluid across this epithelium
- distinct apical and basolateral membrane domains specific for transport
- facilitates transepithelial Na+ and fluid transport for blastocoel formation
- transport is driven by Na, K-adenosine triphosphatase (ATPase) in basolateral membranes of the trophectoderm [18]
Blastocyst Metabolism
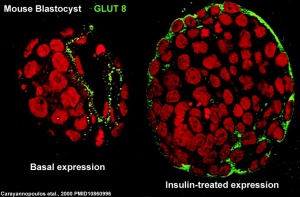
At the blastocyst stage, mammalian development metabolism switches on anaerobic glycolysis metabolism to satisfy metabolic demands of growing blastocyst and formation of the blastocoel. This is thought to be driven by the integral membrane protein family of facilitative glucose transporters (GLUT or SLC2A).
- aerobic - oxidation of lactate and pyruvate via the citric acid cycle (Krebs cycle) and oxidative phosphorylation
- glycolysis- converts glucose into pyruvate
- GLUT - GLUcose Transporter (divided into 3 classes I-III)
- SLC2 - Solute Carrier Family 2
Glucose Transporter Expression
- GLUT1 - from zygote to blastocyst. (all mammalian tissues, basal glucose uptake)
- GLUT2 and GLUT3 - from late eight cell stage to blastocyst. (GLUT2, liver and pancreatic beta cells; GLUT3, all mammalian tissues, basal glucose uptake)
- GLUT4 - not expressed. (muscle and adipose tissue)
- GLUT8 - up-regulated at blastocyst stage. (central nervous system and heart)
- (Data mainly from mouse development, adult tissue expression shown in brackets)
A mouse study,[19] has shown GLUT8 is up-regulated following insulin stimulation, though a more recent GLUT8 knockout mouse shows normal early embryonic development in the absence of this transporter.[20]
- Links: Biochemistry - glucose transporters | GLUT1 | GLUT2 | GLUT8
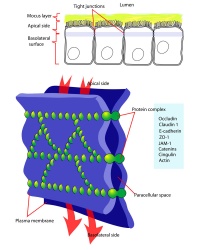
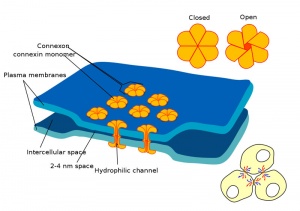
Blastula Cell Communication
Two types of cell junctions have been identified located at different regions in the developing blastocyst.
- Adhesion EM Images: GIT epithelia EM1 | GIT epithelia EM2 | GIT epithelia EM3 | Desmosome EM
- Adhesion Cartoons: Tight junction | Adherens Junction | Desmosome | Gap Junction
Blastocyst Hatching - zona pellucida lost, ZP has sperm entry site, and entire ZP broken down by uterine secretions and possibly blastula secretions.
Uterine Glands - secretions required for blastocyst motility and nutrition
Blastocyst Hatching
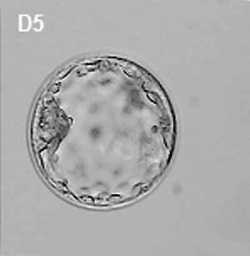
| At about day 5 the human blastocyst "hatches" out of the protective zona pellucida. This hatching allows increased growth, access to uterine nutrient secretions and blastocyst adhesion to the uterine lining. Associated with this hatching process are a series of physical contractions.
|
<mediaplayer width='500' height='450' image="http://php.med.unsw.edu.au/embryology/images/a/af/Human_blastocyst_day_5-6.jpg">File:Human_blastocyst_day_5-6.mp4</mediaplayer>
Human blastocyst contractions (day 5-6)[11] |
Molecular Factors
- TEA DNA- binding domain, these factors bind to the consensus TEA/ATTS cognate binding site[23]
- TEF-3 - renamed Tead1 and Tead4
- Tead3 - is expressed in the placental syncytiotrophoblasts
- E-cadherin - Calcium ion-dependent cell adhesion molecule, a cell membrane adhesive protein required for morula compaction
- epithin - A type II transmembrane serine protease, identified in mouse for compaction of the morula during preimplantation embryonic development. Expressed from 8-cell stage at blastomere contacts and co-localises in the morula with E-cadherin.[24]
- Na, K-adenosine triphosphatase - A sodium potassium pump that generates an osmotic gradient for fluid flow into the blastocoel
- Zonula occludens-1 - (ZO-1) Tight junction protein involved in morula to blastocyst transformation in the mouse [25]
Blastocyst in Other Species
Mouse Blastocyst
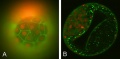
Sox2 expression[26]
Early gene expression[26]
Early gene expression[26]
Early gene expression[26]
Early gene expression[26]
Early mouse development model[27]
- Links: Mouse Development
Bovine Blastocyst
- Links: Bovine Development
References
- ↑ 1.0 1.1 Zhang P, Zucchelli M, Bruce S, Hambiliki F, Stavreus-Evers A, Levkov L, Skottman H, Kerkelä E, Kere J & Hovatta O. (2009). Transcriptome profiling of human pre-implantation development. PLoS ONE , 4, e7844. PMID: 19924284 DOI.
- ↑ Sasaki H. (2015). Position- and polarity-dependent Hippo signaling regulates cell fates in preimplantation mouse embryos. Semin. Cell Dev. Biol. , 47-48, 80-7. PMID: 25986053 DOI.
- ↑ Rayon T, Menchero S, Nieto A, Xenopoulos P, Crespo M, Cockburn K, Cañon S, Sasaki H, Hadjantonakis AK, de la Pompa JL, Rossant J & Manzanares M. (2014). Notch and hippo converge on Cdx2 to specify the trophectoderm lineage in the mouse blastocyst. Dev. Cell , 30, 410-22. PMID: 25127056 DOI.
- ↑ Smith HL, Stevens A, Minogue B, Sneddon S, Shaw L, Wood L, Adeniyi T, Xiao H, Lio P, Kimber SJ & Brison DR. (2019). Systems based analysis of human embryos and gene networks involved in cell lineage allocation. BMC Genomics , 20, 171. PMID: 30836937 DOI.
- ↑ 5.0 5.1 Nissen SB, Perera M, Gonzalez JM, Morgani SM, Jensen MH, Sneppen K, Brickman JM & Trusina A. (2017). Four simple rules that are sufficient to generate the mammalian blastocyst. PLoS Biol. , 15, e2000737. PMID: 28700688 DOI.
- ↑ Kagawa H, Javali A, Khoei HH, Sommer TM, Sestini G, Novatchkova M, Scholte Op Reimer Y, Castel G, Bruneau A, Maenhoudt N, Lammers J, Loubersac S, Freour T, Vankelecom H, David L & Rivron N. (2021). Human blastoids model blastocyst development and implantation. Nature , , . PMID: 34856602 DOI.
- ↑ Gerovska D & Araúzo-Bravo MJ. (2019). Computational analysis of single-cell transcriptomics data elucidates the stabilization of Oct4 expression in the E3.25 mouse preimplantation embryo. Sci Rep , 9, 8930. PMID: 31222057 DOI.
- ↑ Pillai VV, Siqueira LG, Das M, Kei TG, Tu LN, Herren AW, Phinney BS, Cheong SH, Hansen PJ & Selvaraj V. (2019). Physiological profile of undifferentiated bovine blastocyst-derived trophoblasts. Biol Open , 8, . PMID: 30952696 DOI.
- ↑ Maître JL, Turlier H, Illukkumbura R, Eismann B, Niwayama R, Nédélec F & Hiiragi T. (2016). Asymmetric division of contractile domains couples cell positioning and fate specification. Nature , 536, 344-348. PMID: 27487217 DOI.
- ↑ Le Bin GC, Muñoz-Descalzo S, Kurowski A, Leitch H, Lou X, Mansfield W, Etienne-Dumeau C, Grabole N, Mulas C, Niwa H, Hadjantonakis AK & Nichols J. (2014). Oct4 is required for lineage priming in the developing inner cell mass of the mouse blastocyst. Development , 141, 1001-10. PMID: 24504341 DOI.
- ↑ 11.0 11.1 11.2 Wong CC, Loewke KE, Bossert NL, Behr B, De Jonge CJ, Baer TM & Reijo Pera RA. (2010). Non-invasive imaging of human embryos before embryonic genome activation predicts development to the blastocyst stage. Nat. Biotechnol. , 28, 1115-21. PMID: 20890283 DOI.
- ↑ Parks JC, McCallie BR, Janesch AM, Schoolcraft WB & Katz-Jaffe MG. (2011). Blastocyst gene expression correlates with implantation potential. Fertil. Steril. , 95, 1367-72. PMID: 20864103 DOI.
- ↑ Yamanaka Y, Lanner F & Rossant J. (2010). FGF signal-dependent segregation of primitive endoderm and epiblast in the mouse blastocyst. Development , 137, 715-24. PMID: 20147376 DOI.
- ↑ Fong CY & Bongso A. (1999). Comparison of human blastulation rates and total cell number in sequential culture media with and without co-culture. Hum. Reprod. , 14, 774-81. PMID: 10221713
- ↑ Tanaka N, Takeuchi T, Neri QV, Sills ES & Palermo GD. (2006). Laser-assisted blastocyst dissection and subsequent cultivation of embryonic stem cells in a serum/cell free culture system: applications and preliminary results in a murine model. J Transl Med , 4, 20. PMID: 16681851 DOI.
- ↑ 16.0 16.1 Bell CE, Calder MD & Watson AJ. (2008). Genomic RNA profiling and the programme controlling preimplantation mammalian development. Mol. Hum. Reprod. , 14, 691-701. PMID: 19043080 DOI.
- ↑ Nishioka N, Yamamoto S, Kiyonari H, Sato H, Sawada A, Ota M, Nakao K & Sasaki H. (2008). Tead4 is required for specification of trophectoderm in pre-implantation mouse embryos. Mech. Dev. , 125, 270-83. PMID: 18083014 DOI.
- ↑ Kidder GM & Watson AJ. (2005). Roles of Na,K-ATPase in early development and trophectoderm differentiation. Semin. Nephrol. , 25, 352-5. PMID: 16139691 DOI.
- ↑ 19.0 19.1 Carayannopoulos MO, Chi MM, Cui Y, Pingsterhaus JM, McKnight RA, Mueckler M, Devaskar SU & Moley KH. (2000). GLUT8 is a glucose transporter responsible for insulin-stimulated glucose uptake in the blastocyst. Proc. Natl. Acad. Sci. U.S.A. , 97, 7313-8. PMID: 10860996
- ↑ Membrez M, Hummler E, Beermann F, Haefliger JA, Savioz R, Pedrazzini T & Thorens B. (2006). GLUT8 is dispensable for embryonic development but influences hippocampal neurogenesis and heart function. Mol. Cell. Biol. , 26, 4268-76. PMID: 16705176 DOI.
- ↑ <pubmed>14967891</pubmed>
- ↑ Yamazaki K & Kato Y. (1989). Sites of zona pellucida shedding by mouse embryo other than muran trophectoderm. J. Exp. Zool. , 249, 347-9. PMID: 2708952 DOI.
- ↑ Jacquemin P, Hwang JJ, Martial JA, Dollé P & Davidson I. (1996). A novel family of developmentally regulated mammalian transcription factors containing the TEA/ATTS DNA binding domain. J. Biol. Chem. , 271, 21775-85. PMID: 8702974
- ↑ Khang I, Sonn S, Park JH, Rhee K, Park D & Kim K. (2005). Expression of epithin in mouse preimplantation development: its functional role in compaction. Dev. Biol. , 281, 134-44. PMID: 15848395 DOI.
- ↑ Wang H, Ding T, Brown N, Yamamoto Y, Prince LS, Reese J & Paria BC. (2008). Zonula occludens-1 (ZO-1) is involved in morula to blastocyst transformation in the mouse. Dev. Biol. , 318, 112-25. PMID: 18423437 DOI.
- ↑ 26.0 26.1 26.2 26.3 26.4 Keramari M, Razavi J, Ingman KA, Patsch C, Edenhofer F, Ward CM & Kimber SJ. (2010). Sox2 is essential for formation of trophectoderm in the preimplantation embryo. PLoS ONE , 5, e13952. PMID: 21103067 DOI.
- ↑ Krupinski P, Chickarmane V & Peterson C. (2011). Simulating the mammalian blastocyst--molecular and mechanical interactions pattern the embryo. PLoS Comput. Biol. , 7, e1001128. PMID: 21573197 DOI.
Reviews
Maître JL. (2017). Mechanics of blastocyst morphogenesis. Biol. Cell , 109, 323-338. PMID: 28681376 DOI.
Rasmussen TP & Corry GN. (2010). Epigenetic pre-patterning and dynamics during initial stages of mammalian preimplantation development. J. Cell. Physiol. , 225, 333-6. PMID: 20607796 DOI.
Cockburn K & Rossant J. (2010). Making the blastocyst: lessons from the mouse. J. Clin. Invest. , 120, 995-1003. PMID: 20364097 DOI.
Rossant J. (2007). Stem cells and lineage development in the mammalian blastocyst. Reprod. Fertil. Dev. , 19, 111-8. PMID: 17389140
Articles
Smith HL, Stevens A, Minogue B, Sneddon S, Shaw L, Wood L, Adeniyi T, Xiao H, Lio P, Kimber SJ & Brison DR. (2019). Systems based analysis of human embryos and gene networks involved in cell lineage allocation. BMC Genomics , 20, 171. PMID: 30836937 DOI.
Santos J, Pereira CF, Di-Gregorio A, Spruce T, Alder O, Rodriguez T, Azuara V, Merkenschlager M & Fisher AG. (2010). Differences in the epigenetic and reprogramming properties of pluripotent and extra-embryonic stem cells implicate chromatin remodelling as an important early event in the developing mouse embryo. Epigenetics Chromatin , 3, 1. PMID: 20157423 DOI.
Dzamba BJ, Jakab KR, Marsden M, Schwartz MA & DeSimone DW. (2009). Cadherin adhesion, tissue tension, and noncanonical Wnt signaling regulate fibronectin matrix organization. Dev. Cell , 16, 421-32. PMID: 19289087 DOI.
Sheth B, Nowak RL, Anderson R, Kwong WY, Papenbrock T & Fleming TP. (2008). Tight junction protein ZO-2 expression and relative function of ZO-1 and ZO-2 during mouse blastocyst formation. Exp. Cell Res. , 314, 3356-68. PMID: 18817772 DOI.
Nishioka N, Yamamoto S, Kiyonari H, Sato H, Sawada A, Ota M, Nakao K & Sasaki H. (2008). Tead4 is required for specification of trophectoderm in pre-implantation mouse embryos. Mech. Dev. , 125, 270-83. PMID: 18083014 DOI.
Yamanaka Y, Ralston A, Stephenson RO & Rossant J. (2006). Cell and molecular regulation of the mouse blastocyst. Dev. Dyn. , 235, 2301-14. PMID: 16773657 DOI.
Search PubMed
Search April 2010
Search Pubmed: blastocyst development | blastocoel development | inner cell mass development | trophectoderm |
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2026, April 23) Embryology Blastocyst Development. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Blastocyst_Development
- © Dr Mark Hill 2026, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G


![Early gene expression[26]](/embryology/images/thumb/c/ca/Mouse-_preimplantation_gene_expression_02.jpg/120px-Mouse-_preimplantation_gene_expression_02.jpg)
![Early gene expression[26]](/embryology/images/thumb/d/d9/Mouse-_preimplantation_gene_expression_03.jpg/120px-Mouse-_preimplantation_gene_expression_03.jpg)
![Early gene expression[26]](/embryology/images/thumb/5/56/Mouse-_preimplantation_gene_expression_04.jpg/120px-Mouse-_preimplantation_gene_expression_04.jpg)
![Early gene expression[26]](/embryology/images/thumb/7/74/Mouse-_preimplantation_gene_expression_05.jpg/120px-Mouse-_preimplantation_gene_expression_05.jpg)