Developmental Signals - Nodal: Difference between revisions
mNo edit summary |
No edit summary |
||
| Line 44: | Line 44: | ||
===Left-Right Axis Development=== | ===Left-Right Axis Development=== | ||
[[File:Mouse left-right axis 02.jpg|600px]] | {| | ||
|[[File:Mouse left-right axis 02.jpg|600px]] | |||
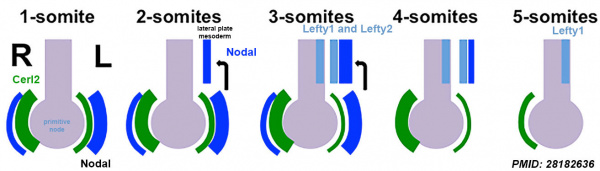
Model ([[:Category:Mouse E8.0|E8.0]] Left-Right Axis<ref name | | Model ([[:Category:Mouse E8.0|E8.0]] Left-Right Axis<ref name=PMID28182636><pubmed>28182636</pubmed></ref> Nodal activity and expression (dark blue) | ||
* 1-somite stage | * Gastrula stage - Initially perinodal crown cells symmetrically express Nodal, Wnt and their antagonist Cerberus like-2 (Cerl2). | ||
* 2-somite stage wild type embryos, Nodal induce Nodal expression in the L-LPM | * 1-somite stage Cerl2 expression (green) becomes asymmetric with reduced expression on the left side of the node in response to fluid flow. With reduced expression of its antagonist on the left, Nodal activity and expression (dark blue) increases on the left side and decreases on the right. | ||
* 2-somite stage wild type embryos, Nodal induce Nodal expression in the L-LPM (left lateral plate mesoderm) | |||
* 3-somite stage robust expression of Nodal along with Lefty1 and Lefty2 (light blue) is detected in the L-LPM and Lefty1 in the midline (light blue). | * 3-somite stage robust expression of Nodal along with Lefty1 and Lefty2 (light blue) is detected in the L-LPM and Lefty1 in the midline (light blue). | ||
* 4-somite stage embryos Nodal expression is reduced but Lefty1 and Lefty2 are strongly expressed in the L-LPM. Lefty1 expression in the midline and Cerl2 around the node inhibits Nodal signaling in the R-LPM. | * 4-somite stage embryos Nodal expression is reduced but Lefty1 and Lefty2 are strongly expressed in the L-LPM. Lefty1 expression in the midline and Cerl2 around the node inhibits Nodal signaling in the R-LPM. | ||
|} | |||
:'''Links:''' [[:File:Mouse left-right axis 02.jpg|wildtype alone]] | [[Developmental Signals - Nodal|Nodal]] | [[Developmental Mechanism - Axes Formation|Axes Formation]] | [[Gastrulation]] | [[:Category:Mouse E8.0|Mouse E8.0]] | [[Mouse Development]] | :'''Links:''' [[:File:Mouse left-right axis 02.jpg|wildtype alone]] | [[Developmental Signals - Nodal|Nodal]] | [[Developmental Mechanism - Axes Formation|Axes Formation]] | [[Gastrulation]] | [[:Category:Mouse E8.0|Mouse E8.0]] | [[Mouse Development]] | ||
Revision as of 12:52, 30 April 2017
| Embryology - 27 Jun 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
Nodal is a member of the TGF-beta family and together with Lefty both are involved in the initial left-right (L-R) patterning of the axis of the embryo during gastrulation. This patterning signal is later used for many cell fate developmental processes. This patterning role was first established in mouse and zebrafish models, the human homolog was first identified in 1997.[1]
Left-right (L-R) asymmetry include the position on the left side of the heart and spleen and the development of the curvature of the stomach.
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Nodal <pubmed limit=5>Nodal</pubmed> |
| Older papers |
|---|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
|
Nodal Signaling
Nodal represents a family of transmembrane receptors passing only once through the plasma membrane. Nodal acts through SMAD2 dependent and independent intracellular pathways.
Nodal Ligands
Functions
Developmental patterning signal.
Left-Right Axis Development
|
Model (E8.0 Left-Right Axis[3] Nodal activity and expression (dark blue)
|
- Links: wildtype alone | Nodal | Axes Formation | Gastrulation | Mouse E8.0 | Mouse Development
(Theiler Stage 12a First Somites unturned embryo with first appearance of somite pairs 1-4 somites. The allantois extends further into the exocoelom and the maxillary components of the 1st branchial arch become prominent. The preotic sulcus is visible in the 2-3 somite embryo. The cardiogenic plate begins to form and the foregut pocket is clearly visible. Embryonic age = 8 dpc (range 7.5-8.75 dpc) 1-7 somite pairs
Stomach Development
- Stomach curvature is generated by left-right asymmetric gut morphogenesis[4] "Left-right (LR) asymmetry is a fundamental feature of internal anatomy, yet the emergence of morphological asymmetry remains one of the least understood phases of organogenesis. Asymmetric rotation of the intestine is directed by forces outside the gut, but the morphogenetic events that generate anatomical asymmetry in other regions of the digestive tract remain unknown. Here, we show in mouse and Xenopus that the mechanisms that drive the curvature of the stomach are intrinsic to the gut tube itself. The left wall of the primitive stomach expands more than the right wall, as the left epithelium becomes more polarized and undergoes radial rearrangement. These asymmetries exist across several species, and are dependent on LR patterning genes, including Foxj1, Nodal and Pitx2 Our findings have implications for how LR patterning manifests distinct types of morphological asymmetries in different contexts."
Abnormalities
- Links:
References
Reviews
<pubmed></pubmed> <pubmed></pubmed> <pubmed></pubmed> <pubmed></pubmed> <pubmed></pubmed> <pubmed></pubmed>
Search Pubmed
Search Bookshelf Nodal
Search Pubmed Now: Nodal Signaling
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- Online Mendelian Inheritance in Man (OMIM) NODAL | LEFTY1 | LEFTY2 | NODAL MODULATOR 1; NOMO1 | NODAL MODULATOR 2; NOMO2 | NODAL MODULATOR 3; NOMO3 |
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, June 27) Embryology Developmental Signals - Nodal. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Nodal
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
