User:Z3414482: Difference between revisions
No edit summary |
|||
| (17 intermediate revisions by 2 users not shown) | |||
| Line 17: | Line 17: | ||
[[User:Z8600021|Mark Hill]] ([[User talk:Z8600021|talk]]) 12:03, 13 October 2016 (AEDT) Where are your later lab attendance records? | [[User:Z8600021|Mark Hill]] ([[User talk:Z8600021|talk]]) 12:03, 13 October 2016 (AEDT) Where are your later lab attendance records? | ||
[[User:Z3414482|Z3414482]] ([[User talk:Z3414482|talk]]) 13:19, 21 October 2016 (AEDT) | |||
==Lab 1 Assessment== | ==Lab 1 Assessment== | ||
<pubmed>PMC4496430</pubmed> | <pubmed>PMC4496430</pubmed> | ||
| Line 107: | Line 109: | ||
|} | |} | ||
==Lab Assessment 6== | ==Lab Assessment 6== | ||
'''1. Identify a known genetic mutation that is associated with cleft lip or palate.''' | |||
Mutations in T-box transcription factor 22 (TBX22) have been found to be one of the genetic factors causing cleft lip or palate with/without ankyloglossia. Ankyloglossia (“tongue tie”) is a birth defect of a short, tight, lingual frenulum that restricts movement of tongue inhibiting speech articulation. Recent studies have also determined that TBX22 mutation is not only associated with cleft palate and ankyloglossia but also cleft lip and palate and tooth agenesis. | |||
'''2. Identify a recent research article on this gene.''' | |||
PMID 21375406 | |||
'''3. How does this mutation affect developmental signalling in normal development.''' | |||
During embryo development, TBX22 is expressed in the palatal shelves and reaches a peak prior to elevation to a horizontal position above the This gene’s mRNS has been detected also in the base of the tongue (in region of frenulum), interior portion of the nasal septum that fuses to the palatal shelves, the mesenchyme from which tooth buds develop and tooth buds themselves. TBX22 gene are essential for early development and particular mesoderm specification. Currently no gene deletions have been detected for TBX22 gene. However various points of mutation within the gene that cause defects have been identified. Some of these points are: splice site, frameshift and missense changes. | |||
==Lab Assessment 7== | ==Lab Assessment 7== | ||
'''1. What is/are the dystrophin mutation(s)?'''<br /> | |||
Dystrophin mutation is what occurs when there are abnormalities affecting the dystrophin gene, such as large deletions, small point mutations and duplications of one or more exons. Mutations within the dystrophin protein complex leads to many forms of autosomally inherited muscular dystrophy and the most well known disease is the X-linked muscle wasting disease of Duchenne muscular dystrophy. Both Duchenne (DMD) and Becker muscular dystrophy (BMD, a milder form of muscular dystrophy that appears at a later age than Duchenne) occur almost exclusively in males and characterized by progressive muscle weakness and atrophy. Whereas DMD is commonly characterised by complete absence of dystrophin, BMD patients display low levels of truncated dystrophin protein. | |||
<pubmed>26721686</pubmed> | |||
'''2. What is the function of dystrophin?''' | |||
Dystrophin is a 427-kDa cytoskeletal protein produced by the largest known human gene. This protein is most commonly found in skeletal muscles and cardiac muscles although small amounts are also detected in nerve cells of the brain. Both in the skeletal and cardiac muscles, dystrophin play a role in strengthening and protecting the muscle fibres to prevent injury to the cells during its contraction and relaxation. The dystrophin complex allow for the muscle cell’s cytoskeleton to be connected with the network proteins and molecules outside the cell, thus acting as an anchor for the muscle cell. In doing so, the dystrophin complex also most likely participates in cell signalling between the cells and the proteins they interact with. | |||
<pubmed>27458343</pubmed> | |||
'''3. What other tissues/organs are affected by this disorder?''' <br /> | |||
Although little information is currently known about the role of dystrophin nerve cells, recent research have suggested that this protein is important for the normal structure and function of synapses. Furthermore, dystrophin is found to be an essential protein for tissues with a secretory function or those that form barriers between functional compartments such as the blood-brain barrier, choroid plexus or kidney. Thus, DMD or BMD would affect the above mentioned tissues and organs of the human body. | |||
<pubmed>16710609</pubmed> | |||
'''4. What therapies exist for DMD?'''<br /> | |||
There is currently no effective treatment available for this disease although much can be done for the management of the symptoms to improve the quality and length of life of patient. The only medication proven to slow the progression of DMD is steroid, which is not always chosen to be used due to the side effects that it may cause. | |||
<pubmed>27621596</pubmed> | |||
'''5. What animal models are available for muscular dystrophy?''' <br /> | |||
Murine (mdx mice) and dog models are currently available for the study of muscular dystrophy | |||
<pubmed>9334342</pubmed> | |||
==Lab Assessment 8== | ==Lab Assessment 8== | ||
Quiz on urogenital development completed | |||
==Lab Assessment 9== | ==Lab Assessment 9== | ||
These are good brief reviews of project pages, with some specific examples. Some of the assessments could have been more critical, given the existing status of these pages. 7/10 | |||
===Group 2 Peer Review=== | |||
'''1. Strengths''' | |||
Group 2’s wiki page is very clearly structured. The content has been structured and organised in a neat format making it easy for readers who have limited knowledge about the signalling pathway to have a comprehensive overview of the topic. I thought that the use of ‘history’ section and images made the information more entertaining to read and process. The information was not only written in clear sentences but cited well. This would be especially beneficial to readers who might be intrigued to read more about the studies done on animals or even certain diseases that are related to abnormalities in notch signalling. | |||
'''2. Points for improvement''' | |||
Although the overall wiki page was of high quality and looked as if it was almost complete, there were few headings/sub-headings such as “current areas of research” that were left blank. And as diagrams have been used in the content of the page, perhaps using pictures/graphs from recent research studies under sub headings such as “roles in embryonic development” would help the readers to understand better the large volume of information. | |||
'''3. Overall''' | |||
In conclusion, I thought that this wiki page was well put together and close to completion after minor adjustments to the existing content and further additions are made to it. From reading this page, I was able to gain a clear understanding for how the notch signalling pathway plays an important role in embryo development. | |||
===Group 3 Peer Review=== | |||
'''1. Strengths''' | |||
Group 3’s wiki page was entertaining to read and understand the information presented about FGFR pathway because of the variety of elements within the page. I appreciated looking at the various diagrams that were both drawn and provided through external resources. The referencing for the diagrams were well done, allowing readers to easily access research articles that can provide more detailed information about the diagrams. The table also was a great addition to the page that could present the information on subtypes of FGFR in a simple manner. Once the quiz at the end of page is actually completed, I think it would be also a great method for the readers to check their understanding of the information. | |||
'''2. Points for improvement''' | |||
Although there was a lot of information on the page, at times the content seemed a bit disorganised due to the incompletion of certain sections. Also there seemed to be not enough information regarding studies already done/currently being done on animal models to find out more about FGFR pathways. More information on the future of studies in regards to this signalling pathway would also be beneficial. I noticed that there were few typos and sentences that needed revision for correction to grammar/spelling and also information sections that were not adequately referenced e.g. “Bone Development”. | |||
'''3. Overall''' | |||
Group 3’s project has a lot of potential to become of even better quality than it is now. As the current wiki page that I have reviewed is a draft version, I believe that the final wiki page at the completion of this assignment could be a lot more engaging with completion of quiz/addition of more diagrams and provide a more complete information content for FGFR pathway once all the subheadings have been filled out. be a lot more engaging and complete in | |||
===Group 4 Peer Review=== | |||
'''1. Strengths''' | |||
From Group 4’s wiki page, I was particularly impressed by the content provided under the headings of “Mechanisms” and “Animal models”. The information was very detailed and referenced well, thus allowing the readers to easily access external resources should they want further information about a certain statement. A long list of reference could be found at the end of the page despite the wiki page being far from completion, indicating the group’s extensive research put in to create this page. | |||
'''2. Points for improvement''' | |||
I would have like to have seen an introduction for Group 4’s page rather than one single image of the Hedgehog signalling pathway being the only content found under the main heading. As there were no information for the subheadings of “History” nor “Function” provided yet, it was initially quite difficult to gage an idea of how and where this signalling pathway functions, especially in regards to embryogenesis. Some diagrams to assist the information under “mechanisms” would also be helpful for readers to understand better the content as well as a glossary as there are certain technical terms like “autocrine” and “paracrine” that readers might have difficulty understanding. | |||
'''3. Overall''' | |||
Overall, I think that Group 4 is off to a very promising start with their wiki page. Once more information has been added in, especially in terms of introducing the pathway, discussing more about the pathway’s role in embryo development, current and future researches on it with diagrams/tables included, I think that the wiki page would be of very high standard. | |||
===Group 5 Peer Review=== | |||
'''1. Strengths''' | |||
Inclusion of articles throughout the wikipage, Use of table/ Use of pargraphs | |||
One of the positive points about Group 5 was their integration of relevant articles in certain sections of their wiki page (as is the case in “Introduction” and “Function of T-box in cardiac development”) rather than compiling a list of articles as a separate section. I found that this helped myself as a reader to know which resources I could read on to gain further understanding about specific aspects of T-box genes and their signalling. The use of the table as a “summary of the main T-box genes” organised the great volume of information in an easy way for the audience to read (although using dot points would have improved the organisation of it). It was also clear that a lot of research had been put into creating this wiki page. I particularly enjoyed reading about the “Functions of T-box in development” which was very thoroughly researched. | |||
'''2. Points for improvement''' | |||
Referencing sometimes seems a bit off (grouping of article references)/ Glossary/ | |||
There were some sections (such as “Other developmental events”) that were barely referenced or others (such as “TBX22/cleft palate”) where the referencing seemed inaccurate. It would have been to also have seen a glossary as there were a lot of technical words that readers with limited knowledge on this signalling pathway would not have understood as clearly. | |||
'''3. Overall''' | |||
Overall, I thought that Group 5’s wikipage was one of the best that I have seen in this course. The layout was near perfect and information about the signalling pathway was very comprehensive! | |||
===Group 6 Peer Review=== | |||
'''1. Strengths''' | |||
Group 6’s wiki page is off to a promising start. Although there is not much content yet on the page, there are subheadings which indicate that the authors of this page are aware of direction they should be researching towards. | |||
'''2. Points for improvement''' | |||
Firstly the wiki page could improve from a better organisation of the headings/subheadings it currently has. At present, all the subheadings are under the heading of “Introduction” which I believe is not true. Secondly, more content is needed as well as research on the TGF beta signalling pathway. At least compiling a list of useful resources to add to the “Reference list” might help the group get started in knowing which articles to start looking for to gain the information they need for each subheading. And more importantly, this wiki page needs to have a clear method of relating this signalling pathway to embryo development and demonstrate how progress has been made in understanding this signalling pathway through recent studies including those with animal models. Use of more images(with better citation), tables and diagrams would improve the quality of this wiki page also. Finally the current information uploaded on the wiki page needs to be correctly referenced (as it hardly is at this point!) | |||
'''3. Overall''' | |||
I think Group 6 has quite a bit of work to get done to reach completion of their wiki page however they are off to a promising start! | |||
{{Stem Cell Presentations 2016}} | |||
==Lab Assessment 11== | |||
5/5 | |||
'''1. Research paper'''<br /> | |||
<pubmed>26544927</pubmed> | |||
'''2. Summary of main findings from research paper''' <br /> | |||
The aim of the research paper was to show how a 40% increase in ventricular cardiomyocyte number that takes place during preadolescence produces newly synthesized DNA in 57% of the cardiomyocyte nuclei. MHC-nLAC mice that expressed a nuclear-localised β-galactosidase were used to investigate further into this bust of preadolescent cell-cycle activity. Through co-localization of β-galactosidase and BrdU immune reactivity, cumulative ventricular cardiomyocyte DNA synthesis was able to be quantitated in the mice models. The results showed that only low levels of cardiomyocyte DNA synthesis were found in mice characterised by pumps from PN10 through PN19 or from PN12 through PN19. Only low levels of cardiomyocyte DNA synthesis were detected in C57BI/6J inbred mice. However BrdU on the otherhand, was not only detected in in small intestine crypt cells by 24 hr postimplantation of all the different mice model types but also at the end of the labelling period thus confirming its ability to continuously infuse. As the mice that received a single BrdU injeciton on PN14.5, PN15 or PN16 and PN19 also showeed little labeling from cardiomyocyte DNA synthesis, the study concluded that BrdU tyotoxicity and/or the presence of the osmotic mini-pump were not confounding factors. | |||
'''3. Relationship between research paper & review article ''' <br /> | |||
Review article: <pubmed>26932668</pubmed> | |||
The review article describes about the latent regenerative potential of cardiomyocytes that can be found in some animal models but not human heart. The review goes through the cellular and molecular mechanisms that facilitate the proliferation of cardiomyocytes. In doing so, it also shows how and why this capacity is lost in adult mammals normally, and therefore highlights some of the current research areas that might lead to restoration of this ability. <br /> The research article I chose was utilised within the review article to emphasise the ceasing of any proliferative activity within neonatal mammal cardiomyocytes. The research article was underlining the fact that despite certain reports that of a discovery in preadolescent proliferative burst of cardiomyocytes, that this in fact was not true as evidence of it could not be replicated by either cardiomyocyte assays or proliferation marker assays. | |||
Latest revision as of 16:37, 17 November 2016
| Student Information (expand to read) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Individual Assessments | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Please leave this template on top of your student page as I will add your assessment items here. Beginning your online work - Working Online in this course
Click here to email Dr Mark Hill | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 1 Assessment - Researching a Topic | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
In the lab I showed you how to find the PubMed reference database and search it using a topic word. Lab 1 assessment will be for you to use this to find a research reference on "fertilization" and write a brief summary of the main finding of the paper.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 2 Assessment - Uploading an Image | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
OK you are now in a group
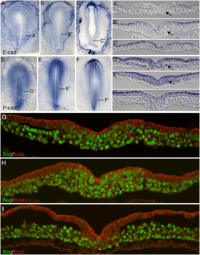
Initially the topic can be as specific or as broad as you want. Chicken embryo E-cad and P-cad gastrulation[1] References
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 4 Assessment - GIT Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
ANAT2341 Quiz Example | Category:Quiz | ANAT2341 Student 2015 Quiz Questions | Design 4 quiz questions based upon gastrointestinal tract. Add the quiz to your own page under Lab 4 assessment and provide a sub-sub-heading on the topic of the quiz. An example is shown below (open this page in view code or edit mode). Note that it is not just how you ask the question, but also how you explain the correct answer. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 5 Assessment - Course Review | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Complete the course review questionnaire and add the fact you have completed to your student page. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 6 Assessment - Cleft Lip and Palate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 7 Assessment - Muscular Dystrophy | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 8 Assessment - Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| A brief quiz was held in the practical class on urogenital development. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 9 Assessment - Peer Assessment | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 10 Assessment - Stem Cells | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
As part of the assessment for this course, you will give a 15 minutes journal club presentation in Lab 10. For this you will in your current student group discuss a recent (published after 2011) original research article (not a review!) on stem cell biology or technology.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 11 Assessment - Heart Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Read the following recent review article on heart repair and from the reference list identify a cited research article and write a brief summary of the paper's main findings. Then describe how the original research result was used in the review article.
<pubmed>26932668</pubmed>Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Lab Attendance
Z3414482 (talk) 14:34, 5 August 2016 (AEST)
Z3414482 (talk) 14:47, 12 August 2016 (AEST)
Z3414482 (talk) 13:15, 19 August 2016 (AEST)
Z3414482 (talk) 13:52, 26 August 2016 (AEST)
Z3414482 (talk) 13:43, 2 September 2016 (AEST)
Z3414482 (talk) 13:24, 9 September 2016 (AEST)
Mark Hill (talk) 12:03, 13 October 2016 (AEDT) Where are your later lab attendance records?
Z3414482 (talk) 13:19, 21 October 2016 (AEDT)
Lab 1 Assessment
<pubmed>PMC4496430</pubmed> Summary This research article studies the effectiveness of artificial oocyte activation (AOA) with a calcium ionophore for fertilization under two circumstances. The first overall circumstance, under which the effectiveness of this procedure was tested for, was when there had been total fertilization failure in past IVF cycles. Secondly, the other more specific cases AOA was tested under, were those that showed severe male factor infertility with non-motile spermatozoa even after pentoxifylline (PF) treatement. AOA is a useful method in avoiding total fertilization failure in human in vitro fetilizaiton-embryo transfer (IVF-ET). AOA attained through calcium ionophore can induce calcium oscillation in oocytes and initate the fertilization process.
For the purpose of this study, between January 2006 to June 2013, 29 intracytoplasmic sperm injection (ICSI) – AOA were performed. All cases were characterised by male factor infertility, however out of the 29 cases further division was made depending on sperm motility after PF treatment. From the data collected, it was concluded that regardless of whether sperm motility was restored after PF treatment or not, oocyte activation is a useful method to ensure fertilization in TESE-ICSI cycles.
| Mark Hill 18 August 2016 - OK so this looks at mechanisms of how the oocyte can be artificially activated giving us insights into the normal in vivo mechanism. Good summary. | Assessment 5/5 |
Lab 2 Assessment
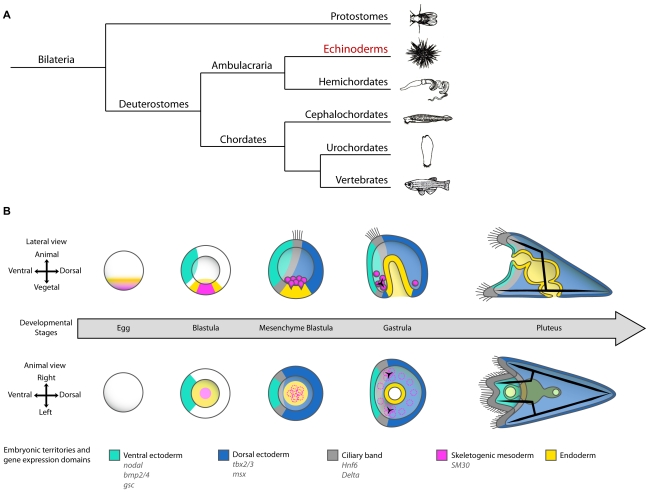
Position of echinoderms in phylogeny of bilateria and establishment of D/V polarity during early development of the sea urchin embryo.
| Mark Hill 29 August 2016 - You do not need to include the Reference, Copyright and Student Image template on your page. But you did need to have a link to the reference with the image legend, as shown below.
Position of echinoderms in phylogeny of bilateria and establishment of D/V polarity during early development of the sea urchin embryo.[1] |
Assessment 4/5 |
Reference
<pubmed>19956794</pubmed>
Copyright
Copyright Lapraz et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
- Note - This image was originally uploaded as part of an undergraduate science student project and may contain inaccuracies in either description or acknowledgements. Students have been advised in writing concerning the reuse of content and may accidentally have misunderstood the original terms of use. If image reuse on this non-commercial educational site infringes your existing copyright, please contact the site editor for immediate removal.
Lab 4 Assessment
| Mark Hill 13 October 2016 - These seem good quiz questions. Note that question 2 your answer says option 2 but it is actually 3 that you have shown as the correct answer. | Assessment 5/5 |
Lab 5 Assessment
Questionnaire done
| Mark Hill 13 October 2016 - Questionnaire on course structure. | Assessment 5/5 |
Lab Assessment 6
1. Identify a known genetic mutation that is associated with cleft lip or palate.
Mutations in T-box transcription factor 22 (TBX22) have been found to be one of the genetic factors causing cleft lip or palate with/without ankyloglossia. Ankyloglossia (“tongue tie”) is a birth defect of a short, tight, lingual frenulum that restricts movement of tongue inhibiting speech articulation. Recent studies have also determined that TBX22 mutation is not only associated with cleft palate and ankyloglossia but also cleft lip and palate and tooth agenesis.
2. Identify a recent research article on this gene.
PMID 21375406
3. How does this mutation affect developmental signalling in normal development.
During embryo development, TBX22 is expressed in the palatal shelves and reaches a peak prior to elevation to a horizontal position above the This gene’s mRNS has been detected also in the base of the tongue (in region of frenulum), interior portion of the nasal septum that fuses to the palatal shelves, the mesenchyme from which tooth buds develop and tooth buds themselves. TBX22 gene are essential for early development and particular mesoderm specification. Currently no gene deletions have been detected for TBX22 gene. However various points of mutation within the gene that cause defects have been identified. Some of these points are: splice site, frameshift and missense changes.
Lab Assessment 7
1. What is/are the dystrophin mutation(s)?
Dystrophin mutation is what occurs when there are abnormalities affecting the dystrophin gene, such as large deletions, small point mutations and duplications of one or more exons. Mutations within the dystrophin protein complex leads to many forms of autosomally inherited muscular dystrophy and the most well known disease is the X-linked muscle wasting disease of Duchenne muscular dystrophy. Both Duchenne (DMD) and Becker muscular dystrophy (BMD, a milder form of muscular dystrophy that appears at a later age than Duchenne) occur almost exclusively in males and characterized by progressive muscle weakness and atrophy. Whereas DMD is commonly characterised by complete absence of dystrophin, BMD patients display low levels of truncated dystrophin protein.
<pubmed>26721686</pubmed>
2. What is the function of dystrophin?
Dystrophin is a 427-kDa cytoskeletal protein produced by the largest known human gene. This protein is most commonly found in skeletal muscles and cardiac muscles although small amounts are also detected in nerve cells of the brain. Both in the skeletal and cardiac muscles, dystrophin play a role in strengthening and protecting the muscle fibres to prevent injury to the cells during its contraction and relaxation. The dystrophin complex allow for the muscle cell’s cytoskeleton to be connected with the network proteins and molecules outside the cell, thus acting as an anchor for the muscle cell. In doing so, the dystrophin complex also most likely participates in cell signalling between the cells and the proteins they interact with.
<pubmed>27458343</pubmed>
3. What other tissues/organs are affected by this disorder?
Although little information is currently known about the role of dystrophin nerve cells, recent research have suggested that this protein is important for the normal structure and function of synapses. Furthermore, dystrophin is found to be an essential protein for tissues with a secretory function or those that form barriers between functional compartments such as the blood-brain barrier, choroid plexus or kidney. Thus, DMD or BMD would affect the above mentioned tissues and organs of the human body.
<pubmed>16710609</pubmed>
4. What therapies exist for DMD?
There is currently no effective treatment available for this disease although much can be done for the management of the symptoms to improve the quality and length of life of patient. The only medication proven to slow the progression of DMD is steroid, which is not always chosen to be used due to the side effects that it may cause.
<pubmed>27621596</pubmed>
5. What animal models are available for muscular dystrophy?
Murine (mdx mice) and dog models are currently available for the study of muscular dystrophy
<pubmed>9334342</pubmed>
Lab Assessment 8
Quiz on urogenital development completed
Lab Assessment 9
These are good brief reviews of project pages, with some specific examples. Some of the assessments could have been more critical, given the existing status of these pages. 7/10
Group 2 Peer Review
1. Strengths
Group 2’s wiki page is very clearly structured. The content has been structured and organised in a neat format making it easy for readers who have limited knowledge about the signalling pathway to have a comprehensive overview of the topic. I thought that the use of ‘history’ section and images made the information more entertaining to read and process. The information was not only written in clear sentences but cited well. This would be especially beneficial to readers who might be intrigued to read more about the studies done on animals or even certain diseases that are related to abnormalities in notch signalling.
2. Points for improvement
Although the overall wiki page was of high quality and looked as if it was almost complete, there were few headings/sub-headings such as “current areas of research” that were left blank. And as diagrams have been used in the content of the page, perhaps using pictures/graphs from recent research studies under sub headings such as “roles in embryonic development” would help the readers to understand better the large volume of information.
3. Overall
In conclusion, I thought that this wiki page was well put together and close to completion after minor adjustments to the existing content and further additions are made to it. From reading this page, I was able to gain a clear understanding for how the notch signalling pathway plays an important role in embryo development.
Group 3 Peer Review
1. Strengths
Group 3’s wiki page was entertaining to read and understand the information presented about FGFR pathway because of the variety of elements within the page. I appreciated looking at the various diagrams that were both drawn and provided through external resources. The referencing for the diagrams were well done, allowing readers to easily access research articles that can provide more detailed information about the diagrams. The table also was a great addition to the page that could present the information on subtypes of FGFR in a simple manner. Once the quiz at the end of page is actually completed, I think it would be also a great method for the readers to check their understanding of the information.
2. Points for improvement
Although there was a lot of information on the page, at times the content seemed a bit disorganised due to the incompletion of certain sections. Also there seemed to be not enough information regarding studies already done/currently being done on animal models to find out more about FGFR pathways. More information on the future of studies in regards to this signalling pathway would also be beneficial. I noticed that there were few typos and sentences that needed revision for correction to grammar/spelling and also information sections that were not adequately referenced e.g. “Bone Development”.
3. Overall
Group 3’s project has a lot of potential to become of even better quality than it is now. As the current wiki page that I have reviewed is a draft version, I believe that the final wiki page at the completion of this assignment could be a lot more engaging with completion of quiz/addition of more diagrams and provide a more complete information content for FGFR pathway once all the subheadings have been filled out. be a lot more engaging and complete in
Group 4 Peer Review
1. Strengths
From Group 4’s wiki page, I was particularly impressed by the content provided under the headings of “Mechanisms” and “Animal models”. The information was very detailed and referenced well, thus allowing the readers to easily access external resources should they want further information about a certain statement. A long list of reference could be found at the end of the page despite the wiki page being far from completion, indicating the group’s extensive research put in to create this page.
2. Points for improvement
I would have like to have seen an introduction for Group 4’s page rather than one single image of the Hedgehog signalling pathway being the only content found under the main heading. As there were no information for the subheadings of “History” nor “Function” provided yet, it was initially quite difficult to gage an idea of how and where this signalling pathway functions, especially in regards to embryogenesis. Some diagrams to assist the information under “mechanisms” would also be helpful for readers to understand better the content as well as a glossary as there are certain technical terms like “autocrine” and “paracrine” that readers might have difficulty understanding.
3. Overall Overall, I think that Group 4 is off to a very promising start with their wiki page. Once more information has been added in, especially in terms of introducing the pathway, discussing more about the pathway’s role in embryo development, current and future researches on it with diagrams/tables included, I think that the wiki page would be of very high standard.
Group 5 Peer Review
1. Strengths
Inclusion of articles throughout the wikipage, Use of table/ Use of pargraphs One of the positive points about Group 5 was their integration of relevant articles in certain sections of their wiki page (as is the case in “Introduction” and “Function of T-box in cardiac development”) rather than compiling a list of articles as a separate section. I found that this helped myself as a reader to know which resources I could read on to gain further understanding about specific aspects of T-box genes and their signalling. The use of the table as a “summary of the main T-box genes” organised the great volume of information in an easy way for the audience to read (although using dot points would have improved the organisation of it). It was also clear that a lot of research had been put into creating this wiki page. I particularly enjoyed reading about the “Functions of T-box in development” which was very thoroughly researched.
2. Points for improvement
Referencing sometimes seems a bit off (grouping of article references)/ Glossary/ There were some sections (such as “Other developmental events”) that were barely referenced or others (such as “TBX22/cleft palate”) where the referencing seemed inaccurate. It would have been to also have seen a glossary as there were a lot of technical words that readers with limited knowledge on this signalling pathway would not have understood as clearly.
3. Overall
Overall, I thought that Group 5’s wikipage was one of the best that I have seen in this course. The layout was near perfect and information about the signalling pathway was very comprehensive!
Group 6 Peer Review
1. Strengths
Group 6’s wiki page is off to a promising start. Although there is not much content yet on the page, there are subheadings which indicate that the authors of this page are aware of direction they should be researching towards.
2. Points for improvement
Firstly the wiki page could improve from a better organisation of the headings/subheadings it currently has. At present, all the subheadings are under the heading of “Introduction” which I believe is not true. Secondly, more content is needed as well as research on the TGF beta signalling pathway. At least compiling a list of useful resources to add to the “Reference list” might help the group get started in knowing which articles to start looking for to gain the information they need for each subheading. And more importantly, this wiki page needs to have a clear method of relating this signalling pathway to embryo development and demonstrate how progress has been made in understanding this signalling pathway through recent studies including those with animal models. Use of more images(with better citation), tables and diagrams would improve the quality of this wiki page also. Finally the current information uploaded on the wiki page needs to be correctly referenced (as it hardly is at this point!)
3. Overall
I think Group 6 has quite a bit of work to get done to reach completion of their wiki page however they are off to a promising start!
| Lab 10 - Stem Cell Presentations 2016 | |
|---|---|
| Group Mark | Assessor General Comments |
|
Group 1: 15/20 Group 2: 19/20 Group 3: 20/20 Group 4: 19/20 Group 5: 16/20 Group 6: 16/20 |
The students put great effort in their presentation and we heard a nice variety of studies in stem cell biology and regenerative medicine today. The interaction after the presentation was great.
As general feedback I would like to advise students to:
|
Lab Assessment 11
5/5
1. Research paper
<pubmed>26544927</pubmed>
2. Summary of main findings from research paper
The aim of the research paper was to show how a 40% increase in ventricular cardiomyocyte number that takes place during preadolescence produces newly synthesized DNA in 57% of the cardiomyocyte nuclei. MHC-nLAC mice that expressed a nuclear-localised β-galactosidase were used to investigate further into this bust of preadolescent cell-cycle activity. Through co-localization of β-galactosidase and BrdU immune reactivity, cumulative ventricular cardiomyocyte DNA synthesis was able to be quantitated in the mice models. The results showed that only low levels of cardiomyocyte DNA synthesis were found in mice characterised by pumps from PN10 through PN19 or from PN12 through PN19. Only low levels of cardiomyocyte DNA synthesis were detected in C57BI/6J inbred mice. However BrdU on the otherhand, was not only detected in in small intestine crypt cells by 24 hr postimplantation of all the different mice model types but also at the end of the labelling period thus confirming its ability to continuously infuse. As the mice that received a single BrdU injeciton on PN14.5, PN15 or PN16 and PN19 also showeed little labeling from cardiomyocyte DNA synthesis, the study concluded that BrdU tyotoxicity and/or the presence of the osmotic mini-pump were not confounding factors.
3. Relationship between research paper & review article
Review article: <pubmed>26932668</pubmed>
The review article describes about the latent regenerative potential of cardiomyocytes that can be found in some animal models but not human heart. The review goes through the cellular and molecular mechanisms that facilitate the proliferation of cardiomyocytes. In doing so, it also shows how and why this capacity is lost in adult mammals normally, and therefore highlights some of the current research areas that might lead to restoration of this ability.
The research article I chose was utilised within the review article to emphasise the ceasing of any proliferative activity within neonatal mammal cardiomyocytes. The research article was underlining the fact that despite certain reports that of a discovery in preadolescent proliferative burst of cardiomyocytes, that this in fact was not true as evidence of it could not be replicated by either cardiomyocyte assays or proliferation marker assays.
- ↑ <pubmed>19956794</pubmed>