Introduction
A protein transcription factor belonging to the evolutionarily conserved forkhead box (FOX) superfamily.
Draft page.
Some Recent Findings
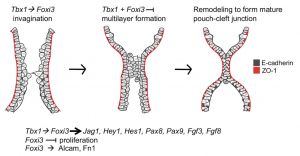
Pharyngeal arch segmentation model Tbx1 and Foxi3
[1]
- Foxa1 and Foxa2 orchestrate development of the urethral tube and division of the embryonic cloaca through an autoregulatory loop with Shh[2] "Congenital anomalies of external genitalia affect approximately 1 in 125 live male births. Development of the genital tubercle, the precursor of the penis and clitoris, is regulated by the urethral plate epithelium, an endodermal signaling center. Signaling activity of the urethral plate is mediated by Sonic hedgehog (SHH), which coordinates outgrowth and patterning of the genital tubercle by controlling cell cycle kinetics and expression of downstream genes. The mechanisms that govern Shh transcription in urethral plate cells are largely unknown. Here we show that deletion of Foxa1 and Foxa2 results in persistent cloaca, an incomplete separation of urinary, genital, and anorectal tracts, and severe hypospadias, a failure of urethral tubulogenesis. Loss of Foxa2 and only one copy of Foxa1 results in urethral fistula, an additional opening of the penile urethra. Foxa1/a2 participate in an autoregulatory feedback loop with Shh, in which FOXA1 and FOXA2 positively regulate transcription of Shh in the urethra, and SHH feeds back to negatively regulate Foxa1 and Foxa2 expression. These findings reveal novel roles for Foxa genes in development of the urethral tube and in division of the embryonic cloaca."
- Tbx1 and Foxi3 genetically interact in the pharyngeal pouch endoderm in a mouse model for 22q11.2 deletion syndrome[1] "We investigated whether Tbx1, the gene for 22q11.2 deletion syndrome (22q11.2DS) and Foxi3, both required for segmentation of the pharyngeal apparatus (PA) to individual arches, genetically interact. We found that all Tbx1+/-;Foxi3+/- double heterozygous mouse embryos had thymus and parathyroid gland defects, similar to those in 22q11.2DS patients.... Several genes expressed in the PA epithelia were downregulated in both Tbx1 and Foxi3 null mutant embryos including Notch pathway genes Jag1, Hes1, and Hey1, suggesting that they may, along with other genes, act downstream to explain the observed genetic interaction. We found Alcam and Fibronectin extracellular matrix proteins were reduced in expression in Foxi3 null but not Tbx1 null embryos, suggesting that some, but not all of the downstream mechanisms are shared." OMIM - Tbx1 | OMIM - Foxi3
- Role of forkhead box gene family in bone metabolism[3] "Bone metabolism is associated with many bone diseases and regulated by multiple signal pathways. Over the past three decades, the functions of a superfamily of evolutionarily conserved transcriptional regulators, known as forkhead box (Fox) family, has been demonstrated to contribute to the bone metabolism. Genetic analysis studies have demonstrated that Fox gene family participate in bone metabolism and that their expression can be regulated by multiple factors. The deregulation of Fox gene family can lead to a series of bone metabolic diseases. In this manuscript, we sketched the biology of the Foxs family, summarized its function of regulating bone metabolism and maintaining bone homeostasis to estimate its potential therapeutic effects in bone diseases, and suggested directions for future exploration in this important field."
- Conserved regulation of neurodevelopmental processes and behavior by FoxP in Drosophila[4] "FOXP proteins form a subfamily of evolutionarily conserved transcription factors involved in the development and functioning of several tissues, including the central nervous system. In humans, mutations in FOXP1 and FOXP2 have been implicated in cognitive deficits including intellectual disability and speech disorders. Drosophila exhibits a single ortholog, called FoxP, but due to a lack of characterized mutants, our understanding of the gene remains poor. Here we show that the dimerization property required for mammalian FOXP function is conserved in Drosophila. In flies, FoxP is enriched in the adult brain, showing strong expression in ~1000 neurons of cholinergic, glutamatergic and GABAergic nature. We generate Drosophila loss-of-function mutants and UAS-FoxP transgenic lines for ectopic expression, and use them to characterize FoxP function in the nervous system. At the cellular level, we demonstrate that Drosophila FoxP is required in larvae for synaptic morphogenesis at axonal terminals of the neuromuscular junction and for dendrite development of dorsal multidendritic sensory neurons. In the developing brain, we find that FoxP plays important roles in α-lobe mushroom body formation. Finally, at a behavioral level, we show that Drosophila FoxP is important for locomotion, habituation learning and social space behavior of adult flies. Our work shows that Drosophila FoxP is important for regulating several neurodevelopmental processes and behaviors that are related to human disease or vertebrate disease model phenotypes. This suggests a high degree of functional conservation with vertebrate FOXP orthologues and established flies as a model system for understanding FOXP related pathologies."
|
| More recent papers
|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
- This search now requires a manual link as the original PubMed extension has been disabled.
- The displayed list of references do not reflect any editorial selection of material based on content or relevance.
- References also appear on this list based upon the date of the actual page viewing.
References listed on the rest of the content page and the associated discussion page (listed under the publication year sub-headings) do include some editorial selection based upon both relevance and availability.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References
Search term: Fox
|
| Older papers
|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
- Immunohistochemical expression analysis of the human fetal lower urogenital tract[5] "We have studied the ontogeny of the developing human Male and Female urogenital tracts from 9 weeks (indifferent stage) to 16 weeks (advanced sex differentiation) of gestation by immunohistochemistry on mid-sagittal sections. Sixteen human fetal pelvises were serial sectioned in the sagittal plane and stained with antibodies to epithelial, muscle, nerve, proliferation and hormone receptor markers. Key findings are: (1) The corpus cavernosum in males and females extends into the glans penis and clitoris, respectively, during the ambisexual stage (9 weeks) and thus appears to be an androgen-independent event. (2) The entire human male (and female) urethra is endodermal in origin based on the presence of FOXA1, KRT 7, uroplakin, and the absence of KRT10 staining. The endoderm of the urethra interfaces with ectodermal epidermis at the site of the urethral meatus. (3) The surface epithelium of the verumontanum is endodermal in origin (FOXA1-positive) with a possible contribution of Pax2-positive epithelial cells implying additional input from the Wolffian duct epithelium. (4) Prostatic ducts arise from the endodermal (FOXA1-positive) urogenital sinus epithelium near the verumontanum. (5) Immunohistochemical staining of mid-sagittal and para-sagittal sections revealed the external anal sphincter, levator ani, bulbospongiosus muscle and the anatomic relationships between these developing skeletal muscles and organs of the Male and Female reproductive tracts."
- Tbx1 is regulated by forkhead proteins in the secondary heart field.[6] "Transcriptional regulation in a tissue-specific and quantitative manner is essential for developmental events, including those involved in cardiovascular morphogenesis. Tbx1 is a T-box-containing transcription factor that is responsible for many of the defects observed in 22q11 deletion syndrome in humans. Tbx1 is expressed in the secondary heart field (SHF) and is essential for cardiac outflow tract (OFT) development....These results suggest that Fox proteins are involved in most, if not all, Tbx1 expression domains and that Tbx1 marks a subset of SHF-derived cells, particularly those that uniquely contribute to the right-sided outflow tract and proximal pulmonary artery." (More? TBX | cardiovascular )
|
Transcription Factor
Abnormalities
Associated with defects in each Fox protein or their signaling pathway.
References
- ↑ 1.0 1.1 Hasten E & Morrow BE. (2019). Tbx1 and Foxi3 genetically interact in the pharyngeal pouch endoderm in a mouse model for 22q11.2 deletion syndrome. PLoS Genet. , 15, e1008301. PMID: 31412026 DOI.
- ↑ Gredler ML, Patterson SE, Seifert AW & Cohn MJ. (2020). Foxa1 and Foxa2 orchestrate development of the urethral tube and division of the embryonic cloaca through an autoregulatory loop with Shh. Dev. Biol. , 465, 23-30. PMID: 32645357 DOI.
- ↑ Huang J, Shen G, Ren H, Zhang Z, Yu X, Zhao W, Shang Q, Cui J, Yu P, Peng J, Liang Z, Yang X & Jiang. (2020). Role of forkhead box gene family in bone metabolism. J. Cell. Physiol. , 235, 1986-1994. PMID: 31549399 DOI.
- ↑ Castells-Nobau A, Eidhof I, Fenckova M, Brenman-Suttner DB, Scheffer-de Gooyert JM, Christine S, Schellevis RL, van der Laan K, Quentin C, van Ninhuijs L, Hofmann F, Ejsmont R, Fisher SE, Kramer JM, Sigrist SJ, Simon AF & Schenck A. (2019). Conserved regulation of neurodevelopmental processes and behavior by FoxP in Drosophila. PLoS ONE , 14, e0211652. PMID: 30753188 DOI.
- ↑ Shen J, Isaacson D, Cao M, Sinclair A, Cunha GR & Baskin L. (2018). Immunohistochemical expression analysis of the human fetal lower urogenital tract. Differentiation , 103, 100-119. PMID: 30287094 DOI.
- ↑ Maeda J, Yamagishi H, McAnally J, Yamagishi C & Srivastava D. (2006). Tbx1 is regulated by forkhead proteins in the secondary heart field. Dev. Dyn. , 235, 701-10. PMID: 16444712 DOI.
Search Bookshelf Fox
Reviews
Golson ML & Kaestner KH. (2016). Fox transcription factors: from development to disease. Development , 143, 4558-4570. PMID: 27965437 DOI.
Ramezani A, Nikravesh H & Faghihloo E. (2019). The roles of FOX proteins in virus-associated cancers. J. Cell. Physiol. , 234, 3347-3361. PMID: 30362516 DOI.
Fortin J, Ongaro L, Li Y, Tran S, Lamba P, Wang Y, Zhou X & Bernard DJ. (2015). Minireview: Activin Signaling in Gonadotropes: What Does the FOX say… to the SMAD?. Mol. Endocrinol. , 29, 963-77. PMID: 25942106 DOI.
Thackray VG. (2014). Fox tales: regulation of gonadotropin gene expression by forkhead transcription factors. Mol. Cell. Endocrinol. , 385, 62-70. PMID: 24099863 DOI.
Articles
Search Pubmed
Search Pubmed Now: Fox
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2026, Mayıs 13) Embryology Developmental Signals - Fox. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Fox
- What Links Here?
- © Dr Mark Hill 2026, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G

