Worm Development: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 14: | Line 14: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* '''The {{Hox}} Gene egl-5 Acts as a Terminal Selector for VD13 Development via Wnt Signaling'''{{#pmid:32138237|PMID32138237}} "Nervous systems are comprised of diverse cell types that differ functionally and morphologically. During development, extrinsic signals, e.g., growth factors, can activate intrinsic programs, usually orchestrated by networks of transcription factors. Within that network, transcription factors that drive the specification of features specific to a limited number of cells are often referred to as terminal selectors. While we still have an incomplete view of how individual neurons within organisms become specified, reporters limited to a subset of neurons in a nervous system can facilitate the discovery of cell specification programs. We have identified a fluorescent reporter that labels VD13, the most posterior of the 19 inhibitory GABA (γ-amino butyric acid)-ergic motorneurons, and two additional neurons, LUAL and LUAR. Loss of function in multiple Wnt signaling genes resulted in an incompletely penetrant loss of the marker, selectively in VD13, but not the LUAs, even though other aspects of GABAergic specification in VD13 were normal. The posterior Hox gene, egl-5, was necessary for expression of our marker in VD13, and ectopic expression of egl-5 in more anterior GABAergic neurons induced expression of the marker. These results suggest egl-5 is a terminal selector of VD13, subsequent to GABAergic specification." {{Hox}} {{neural}} | |||
* '''Homeobox Genes of Caenorhabditis elegans and Spatio-Temporal Expression'''<ref name="PMID26024448"><pubmed>26024448</pubmed>| [http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0126947 PLoS One.]</ref> "We show that, out of 103 homeobox genes, 70 are co-orthologous to human homeobox genes. 14 are highly divergent, lacking an obvious ortholog even in other Caenorhabditis species. One of these homeobox genes encodes 12 homeodomains, while three other highly divergent homeobox genes encode a novel type of double homeodomain, termed HOCHOB. To understand how transcription factors regulate cell fate during development, precise spatio-temporal expression data need to be obtained. Using a new imaging framework that we developed, Endrov, we have generated spatio-temporal expression profiles during embryogenesis of over 60 homeobox genes, as well as a number of other developmental control genes using GFP reporters." [[Developmental Signals - Homeobox] | * '''Homeobox Genes of Caenorhabditis elegans and Spatio-Temporal Expression'''<ref name="PMID26024448"><pubmed>26024448</pubmed>| [http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0126947 PLoS One.]</ref> "We show that, out of 103 homeobox genes, 70 are co-orthologous to human homeobox genes. 14 are highly divergent, lacking an obvious ortholog even in other Caenorhabditis species. One of these homeobox genes encodes 12 homeodomains, while three other highly divergent homeobox genes encode a novel type of double homeodomain, termed HOCHOB. To understand how transcription factors regulate cell fate during development, precise spatio-temporal expression data need to be obtained. Using a new imaging framework that we developed, Endrov, we have generated spatio-temporal expression profiles during embryogenesis of over 60 homeobox genes, as well as a number of other developmental control genes using GFP reporters." [[Developmental Signals - Homeobox] | ||
* '''Basic Caenorhabditis elegans Methods: Synchronization and Observation'''<ref><pubmed>22710399</pubmed>| [http://www.jove.com/video/4019/basic-caenorhabditis-elegans-methods-synchronization-and-observation J Vis Exp.]</ref> "Research into the molecular and developmental biology of the nematode Caenorhabditis elegans was begun in the early seventies by Sydney Brenner and it has since been used extensively as a model organism (1). C. elegans possesses key attributes such as simplicity, transparency and short life cycle that have made it a suitable experimental system for fundamental biological studies for many years (2). ...Because of its transparency, C. elegans structures can be distinguished under the microscope using Differential Interference Contrast microscopy, also known as Nomarski microscopy. The use of a fluorescent DNA binder, DAPI (4',6-diamidino-2-phenylindole), for instance, can lead to the specific identification and localization of individual cells, as well as subcellular structures/defects associated to them." | * '''Basic Caenorhabditis elegans Methods: Synchronization and Observation'''<ref><pubmed>22710399</pubmed>| [http://www.jove.com/video/4019/basic-caenorhabditis-elegans-methods-synchronization-and-observation J Vis Exp.]</ref> "Research into the molecular and developmental biology of the nematode Caenorhabditis elegans was begun in the early seventies by Sydney Brenner and it has since been used extensively as a model organism (1). C. elegans possesses key attributes such as simplicity, transparency and short life cycle that have made it a suitable experimental system for fundamental biological studies for many years (2). ...Because of its transparency, C. elegans structures can be distinguished under the microscope using Differential Interference Contrast microscopy, also known as Nomarski microscopy. The use of a fluorescent DNA binder, DAPI (4',6-diamidino-2-phenylindole), for instance, can lead to the specific identification and localization of individual cells, as well as subcellular structures/defects associated to them." | ||
| Line 19: | Line 21: | ||
|} | |} | ||
{| class="wikitable mw-collapsible mw-collapsed" | {| class="wikitable mw-collapsible mw-collapsed" | ||
! More recent papers | ! More recent papers | ||
|- | |- | ||
| [[File:Mark_Hill.jpg|90px|left]] {{Most_Recent_Refs}} | | [[File:Mark_Hill.jpg|90px|left]] {{Most_Recent_Refs}} | ||
Search term: [http://www.ncbi.nlm.nih.gov/pubmed/?term=Worm+Embryology ''Worm Embryology''] | Search term: [http://www.ncbi.nlm.nih.gov/pubmed/?term=Caenorhabditis+elegans+Development ''Caenorhabditis elegans Development''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Worm+Development ''Worm Development''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Worm+Embryology ''Worm Embryology''] | ||
|} | |||
==Adult Anatomy== | ==Adult Anatomy== | ||
| Line 100: | Line 101: | ||
{{ | {{Animals}} | ||
{{Glossary}} | |||
{{Footer}} | |||
[[Category:Worm]] | [[Category:Worm]] | ||
Revision as of 11:55, 30 April 2020
| Embryology - 21 May 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
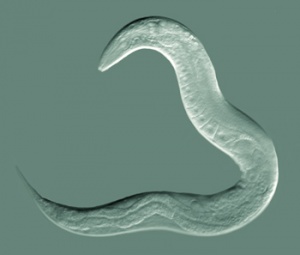
Initially used in the 1960's by Sydney Brenner to study the genetics of development and neurobiology. Early embryological studies of the nematode worm (roundworm) Caenorhabditis elegans (C.Elegans, so called because of its "elegant" curving movement) characterized the fate of each and every cell in the worm through all stages of development. This worm was the first to have its entire genome sequenced and also used recently in space experiments (see below).
The USA space shuttle Atlantis in November 2009 launched Caenorhabditis elegans into space as part of an experiment to study RNA interference and protein phosphorylation in a space environment.
- "RNA interference and protein phosphorylation in space environment using the nematode Caenorhabditis elegans (CERISE) is an experiment that addresses two scientific objectives. The first is to evaluate the effect of microgravity on ribonucleic acid (RNA) interference. The second is to study how the space environment effects protein phosphorylation (addition of a phosphate molecule) and signal transduction in the muscle fibers of gene knock-downed Caenorhabditis elegans."
| Animal Development: axolotl | bat | cat | chicken | cow | dog | dolphin | echidna | fly | frog | goat | grasshopper | guinea pig | hamster | horse | kangaroo | koala | lizard | medaka | mouse | opossum | pig | platypus | rabbit | rat | salamander | sea squirt | sea urchin | sheep | worm | zebrafish | life cycles | development timetable | development models | K12 |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Caenorhabditis elegans Development | Worm Development | Worm Embryology |
Adult Anatomy
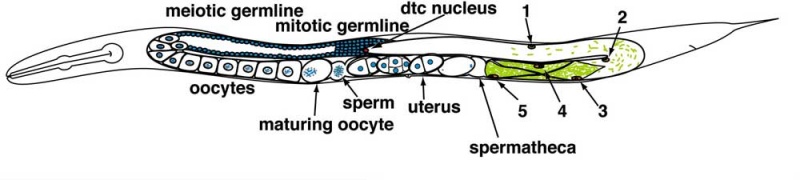
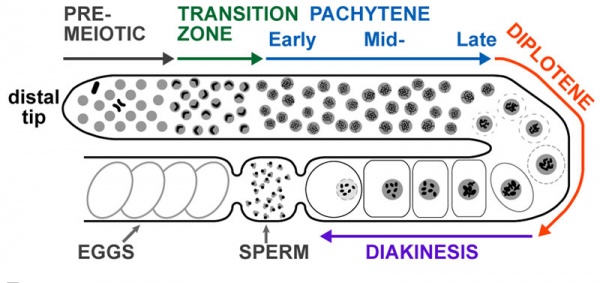
Adult Hermaphrodite Gonad
Adult hermaphrodite gonad arm[5] - A drawing representation of an adult hermaphrodite gonad arm. The progression of germ cell proliferation and meiosis are indicated by the arrows starting from the distal tip region of the gonad arm.
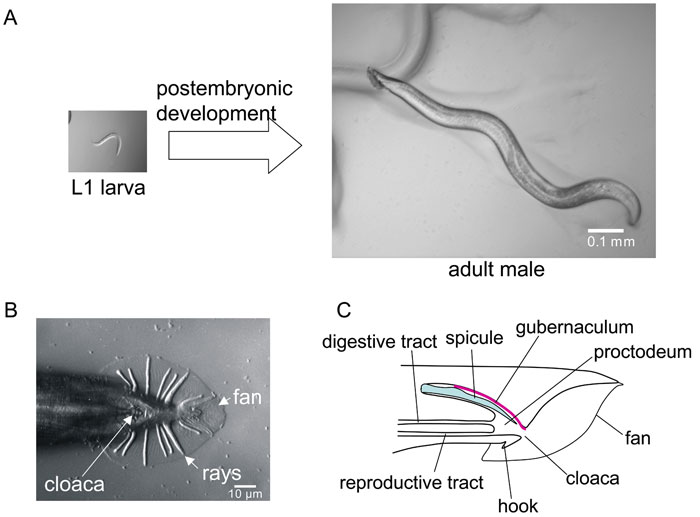
Male Development
The features that differentiate the C. elegans male from the hermaphrodite arise during postembryonic development.[6]
RNA interference
The two researchers, Andrew Z. Fire and Craig C. Mello[7], were investigating how gene expression is regulated in C. elegans and identified the novel regulation method of RNA interference (RNAi), gene silencing by double-stranded RNA. This discovery was awarded the 2006 Nobel Prize in Physiology or Medicine.
- Links: 2006 Nobel Press Release
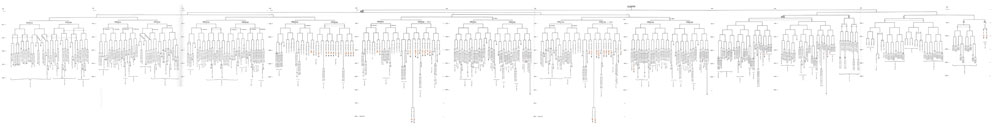
Embryonic Cell Lineages
The overview diagram above shows the fate of each individual cell in the developing c. elegans.
- Zygote (P0 cell) divides into two daughter cells (AB and P1 cells).
- These two daughter cells then divide into the next generation.
- the "X" indicates cells that die by apoptosis during development.
Note the above image is not at a readable resolution, to view see large readable version (10,389 × 1,336 pixels). Embryonic cell lineage developed by J .E. Sulston, E. Schierenberg, J. G. White, J. N. Thomson.
- Links: Apoptosis | Worm Atlas - Cell Lineages
Gastrointestinal Tract
The worm digestive tract consists of a pharynx, intestine, and rectum and contains only about 100 cells. Development is regulated by similar transcription factors found for other species (FoxA and GATA factors).[8]
- FoxA - pharynx and rectum
- GATA - intestine
References
- ↑ Kurland M, O'Meara B, Tucker DK & Ackley BD. (2020). The Hox Gene egl-5 Acts as a Terminal Selector for VD13 Development via Wnt Signaling. J Dev Biol , 8, . PMID: 32138237 DOI.
- ↑ <pubmed>26024448</pubmed>| PLoS One.
- ↑ <pubmed>22710399</pubmed>| J Vis Exp.
- ↑ <pubmed>20232378</pubmed>
- ↑ <pubmed>20661436</pubmed>| PLoS Genetics
- ↑ <pubmed>18050419</pubmed> Worm Book - Male development
- ↑ <pubmed>12857879</pubmed>| PMC165691
- ↑ <pubmed>20570129</pubmed>
Reviews
WormBook - a comprehensive, open-access collection of original, peer-reviewed chapters covering topics related to the biology of Caenorhabditis elegans and other nematodes.
- Asymmetric cell division and axis formation in the embryo
- Embryological variation during nematode development
Articles
Search Pubmed
July 2010 "c elegans Development" All (5126) Review (898) Free Full Text (2363)
Search Pubmed: Worm Development | Caenorhabditis elegans Development | c elegans Development
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- WormBook open-access collection of chapters covering topics related to the biology of Caenorhabditis elegans (C. elegans) and other nematodes.
- Caenorhabditis Genome Sequencing Projects
- Caenorhabditis elegans WWW Server
- The Hall lab - aim to serve the C. elegans community by creating a digital database of TEM images & wild type tissues- an Atlas of worm anatomy.
| Animal Development: axolotl | bat | cat | chicken | cow | dog | dolphin | echidna | fly | frog | goat | grasshopper | guinea pig | hamster | horse | kangaroo | koala | lizard | medaka | mouse | opossum | pig | platypus | rabbit | rat | salamander | sea squirt | sea urchin | sheep | worm | zebrafish | life cycles | development timetable | development models | K12 |
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, May 21) Embryology Worm Development. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Worm_Development
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G