User:Z5015686: Difference between revisions
| Line 113: | Line 113: | ||
'''Identify a recent research article on this gene:''' | '''Identify a recent research article on this gene:''' <pubmed>23029012</pubmed> | ||
<pubmed>23029012</pubmed> | |||
'''How does this mutation affect developmental signaling in normal development:''' | '''How does this mutation affect developmental signaling in normal development:''' | ||
Revision as of 00:40, 23 September 2016
| Student Information (expand to read) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Individual Assessments | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Please leave this template on top of your student page as I will add your assessment items here. Beginning your online work - Working Online in this course
Click here to email Dr Mark Hill | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 1 Assessment - Researching a Topic | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
In the lab I showed you how to find the PubMed reference database and search it using a topic word. Lab 1 assessment will be for you to use this to find a research reference on "fertilization" and write a brief summary of the main finding of the paper.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 2 Assessment - Uploading an Image | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
OK you are now in a group
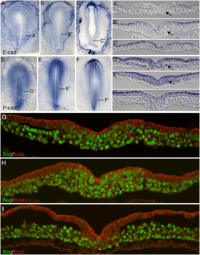
Initially the topic can be as specific or as broad as you want. Chicken embryo E-cad and P-cad gastrulation[1] References
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 4 Assessment - GIT Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
ANAT2341 Quiz Example | Category:Quiz | ANAT2341 Student 2015 Quiz Questions | Design 4 quiz questions based upon gastrointestinal tract. Add the quiz to your own page under Lab 4 assessment and provide a sub-sub-heading on the topic of the quiz. An example is shown below (open this page in view code or edit mode). Note that it is not just how you ask the question, but also how you explain the correct answer. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 5 Assessment - Course Review | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Complete the course review questionnaire and add the fact you have completed to your student page. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 6 Assessment - Cleft Lip and Palate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 7 Assessment - Muscular Dystrophy | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 8 Assessment - Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| A brief quiz was held in the practical class on urogenital development. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 9 Assessment - Peer Assessment | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 10 Assessment - Stem Cells | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
As part of the assessment for this course, you will give a 15 minutes journal club presentation in Lab 10. For this you will in your current student group discuss a recent (published after 2011) original research article (not a review!) on stem cell biology or technology.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 11 Assessment - Heart Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Read the following recent review article on heart repair and from the reference list identify a cited research article and write a brief summary of the paper's main findings. Then describe how the original research result was used in the review article.
<pubmed>26932668</pubmed>Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Lab Attendance
Z5015686 (talk) 14:34, 5 August 2016 (AEST)
Z5015686 (talk) 14:40, 12 August 2016 (AEST)
Z5015686 (talk) 13:15, 19 August 2016 (AEST)
Z5015686 (talk) 13:10, 26 August 2016 (AEST)
Z5015686 (talk) 13:07, 2 September 2016 (AEST)
Z5015686 (talk) 13:07, 9 September 2016 (AEST)
Z5015686 (talk) 13:33, 16 September 2016 (AEST)
Lab 1 Assessment
<pubmed> PMC4770082 </pubmed>
Short Summary Of Findings
The results of this paper suggest that human oocyte developmental potential can be predicted by the quality and maturation of the oocyte prior to fertilisation, (at the 2PN, pronucleus phase). The experiential design outlined in the paper involved measuring the mechanical properties of mouse and human zygotes using minimally invasive technologies (such as micropipette aspiration) to determine which were most predictive of viability (viability was defined as embryos that would most likely survive to blastocyst stage of development.) The results showed that individual parameters had limited predictive power on viability, however when considered together there was a greater distinction between viable and non-viable embryos. Through the use of statistical analysis it was found that their method of classification to predict embryo blastocyst formation that was based on these mechanical properties had >90% precision, 95% specificity and 75% sensitivity. Mice received embryos that were predicted to be either viable or non-viable based on their mechanical properties, which positively correlated to the mice who later had live births. The experimenters then investigated firstly whether there was a correlation between the viable and non-viable embryos and their gene expression, and secondly how/why these mechanical parameters correlated with viability. Interestingly they found that non-viable embryos had a reduced/different expression of some genes that are important for processes including, but not limited to, regulating cell cycle, oocyte maturation, chromosome segregation, DNA repair and telomere maintenance, thus suggesting that zygote gene expression correlates with viability. They also found that non-viable oocytes might undergo suboptimal fertilisation. Some genes that were identified to be differentially expressed in viable and non-viable embryos are important for fertilisation, including some whose products are found on the oocyte plasma membrane and in its zona pellucida, where if expressed incorrectly could potentially inhibit sperm-egg binding. Additionally, a reduced expression for a gene coding for a sperm protein was identified in non-viable zygotes, as well as a receptor that is involved in initiating the calcium oscillations that leads to cortical granule release and zona-hardening (which assists in the prevention of polyspermy.) Therefore in conclusion, this research demonstrates a way to accurately predict embryo viability early on in development, at the pronucleus stage, suggesting that embryo developmental potential is determined pre-fertilisation. This research has relevant applications in embryo selection process in IVF clinics.
| Mark Hill 18 August 2016 - You have added the citation correctly and written a good summary of the article's main findings. I guess the question is what provides the zygote viscoelastic properties and sperm gene expression? | Assessment 5/5 |
Lab 2 Assessment
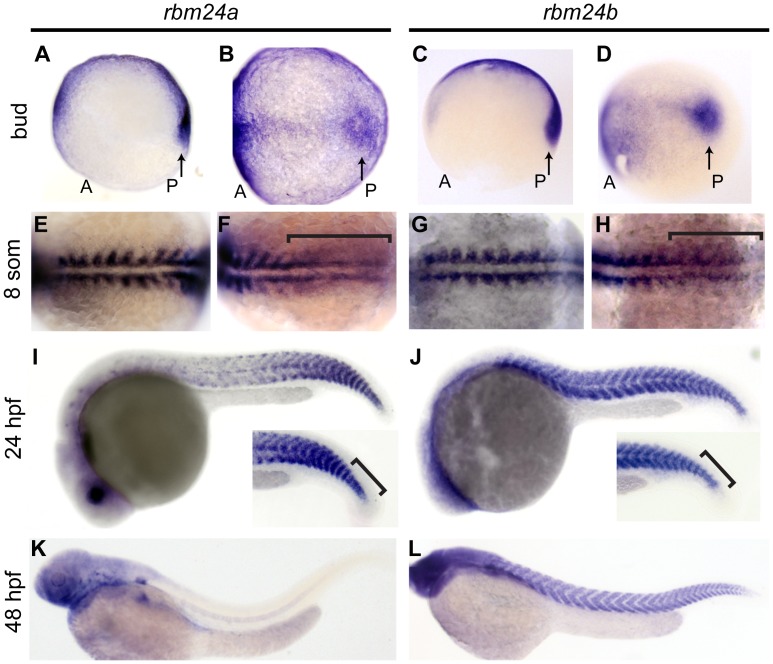
Rbm24a and rbm24b are expressed throughout somitogenesis[1]
| Mark Hill 29 August 2016 - All information Reference, Copyright and Student Image template correctly included with the file and referenced on your page here. | Assessment 5/5 |
Lab 3 Assessment
| Mark Hill 31 August 2016 - Lab 3 Assessment Quiz - Mesoderm and Ectoderm development. | Assessment 4/5 |
Lab 4 Assessment
Gastrointestinal Quiz
Lab 5 Assessment
Completed Course Feedback Questionnaire
Z5015686 (talk) 13:15, 9 September 2016 (AEST)
Lab 6 Assessment
Identify a known genetic mutation that is associated with cleft lip or palate: Mutations in the Interferon Regulatory Factor 6 (IRF6) protein-coding gene (located on chromosome 1) account for the majority of cases of Van der Woude syndrome (VDWS), an autosomal dominant genetic disorder, which has been found to be associated with both cleft lip and cleft palate.
Identify a recent research article on this gene: <pubmed>23029012</pubmed>
How does this mutation affect developmental signaling in normal development: For the most part the underlying mechanism behind the mutation of the IRF6 gene and the development of cleft lip and palate is largely unknown. However, animal studies involving Irf6 mutant mice have offered an explanation to why this gene could contribute to the development of these abnormalities. These mice presented with hyper-proliferative epidermis failing to undergo terminal differentiation, leading to epithelial adhesions that are able to occlude the oral cavity. IRF6 is also thought to be involved in keratinocyte proliferation and differentiation as well as the formation of the oral periderm. [2] Recent research suggests that IRF6 gene interacts with other genes, specifically the Transforming Growth Factor Alpha (TGFA) gene (involved in activating a signalling pathway responsible for cell proliferation, differentiation and development) and may account for up to 10% of cleft lip and cleft palate cases. Interestingly, IRFA knockout mice didn’t express Tgfa in tissues in the palate. [3] In summary it is thought that mutations in the IRF6 gene are thought to affect developmental signalling directly or through associations with other genes, however more research is required.
Lab 7 Assessment
What is/are the dystrophin mutation(s)? The dystrophin gene is the largest known human gene and is located on locus Xp21. Mutations of this gene (such as selections, point mutations and duplications) affect the structure/function of the protein dystrophin, and is responsible for causing both Duchenne (DMD) and Becker (BMD) muscular dystrophies[4], which have a prevalence 4.78 and 1.53 per 100,000 males respectively.[5] Although they have similar signs and symptoms, they vary in their severity, onset age and rate at which the disease develops - with DND being in general, the more common and severe of the two, appearing earlier in childhood in the from of muscle weakness and rapidly develops, affected individuals have impaired development of normal motor functions. Both are associated with the heart condition cardiomyopathy (weakened cardiac muscles).[6]
What is the function of dystrophin?
Dystrophin is an important cytoskeletal protein, and is a crucial component of the larger dystrophin-glycoprotein complex (DGC) which functions to both stabilise and signal interactions between the cytoskeleton, membrane and extracellular matrix, essentially have a central role in mediating muscle stability. Dystrophin has four main functional domains (actin binding amino terminal, central rod, cysteine-rich domains and carboxyl terminus) which help mediate the complexes interactions with cellular components, for example mediates interactions with actin filaments through the actin binding domain, and interactions with microtubules through the rod domain.[7]
What other tissues/organs are affected by this disorder? This disorder is known to result in both cardiac failure and respiratory failure due to the weakening of muscles (as healthy muscle fibres are lost and replaced by fibrosis and fat, and thus have reduced function.)[8]
What therapies exist for DMD? Diagnosis of DMD can be confirmed through DNA tests, muscle biopsy (testing for presence or relative size of dystrophin) and even prenatal tests, and although there is no current cure for this disease, some treatments are available to help control age of onset in the hope to maximise affected individuals quality of life. Pharmacological treatments include corticosteriods (including prednisolone and deflazacort) which have shown some benefits in patients such as an improvement in strength, pulmonary function, timed motor function and delaying age at loss of ambulation and cardiomyopathy onset - however, these medications are not without their own set of side effects.[9] Current research is looking into the possibility gene therapies which aim to restore dystrophin expression such as the use of viral vectors (acting as vehicles for DMD gene)[10], and antisense oligonucleotide mediated exon skipping. [11]
What animal models are available for muscular dystrophy? Historically the most popular animal model for muscular dystrophy over the years has been the MDX mouse, the results of which have been shown to be promising and now a larger animal model of canine DMD (cDMD) is being used.[12]
References
- ↑ <pubmed>25170925</pubmed> PLOSONE
- ↑ <pubmed>21331089</pubmed> [1]
- ↑ <pubmed>23029012</pubmed> [2]
- ↑ <pubmed> 4767260</pubmed> [3]
- ↑ <pubmed>24780148</pubmed>[4]
- ↑ <pubmed> 4767260</pubmed> [5]
- ↑ <pubmed> 4767260</pubmed> [6]
- ↑ <pubmed>4767260</pubmed> [7]
- ↑ <pubmed>26833937</pubmed> [8]
- ↑ <pubmed>27215286</pubmed> [9]
- ↑ <pubmed>23829870</pubmed> [10]
- ↑ <pubmed> 25740330</pubmed> [11]
fertilization PMID 27486480