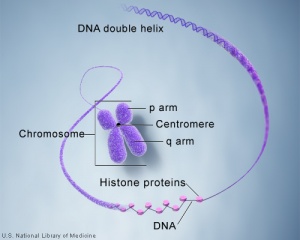
Molecular Development - Epigenetics
Introduction
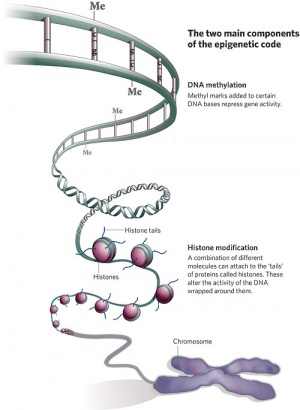
In terms of molecular mechanisms, the field of epigenetics has begun to florish with some recent important findings. Epigenetics as the name implies, is the inheritance mechanisms that lie outside the DNA sequence of our genes and include DNA methylation, histone modification, and those of the microRNA machinery. One of the initial discoveries was the effects of DNA methylation upon gene expression and then modifications of nucleosomal histones. DNA methylation is usually associated with 5-methylcytosine (m5C) and leads to transcriptional silencing in vertebrates. Epigenetic modifications can be transmitted from one cell generation to the next (mitotic inheritance) and can also be transmitted down organismal generations (meiotic inheritance). Recently the term “methylome” has been coined to refer to the methylation profile of the whole genome.
Molecular mechanisms of development is an exciting research area and requires a variety of different skills. This page introduces only a few examples and should give you a feel for the topic. Note that each section of system notes has a page covering molecular development in that system.
Some Recent Findings
|
- Links: References | More References
DNA Methylation
Enzymes that lead to DNA methylation are described as methyltransferases (DNMTs) and fall into two categories.
- DNMT1 - copies the pattern of DNA methylation during cell replication (methylation maintenance).
- DNMT3a and DNMT3b - are responsible for the de novo DNA methylation.
DNA Methylation Changes with Age
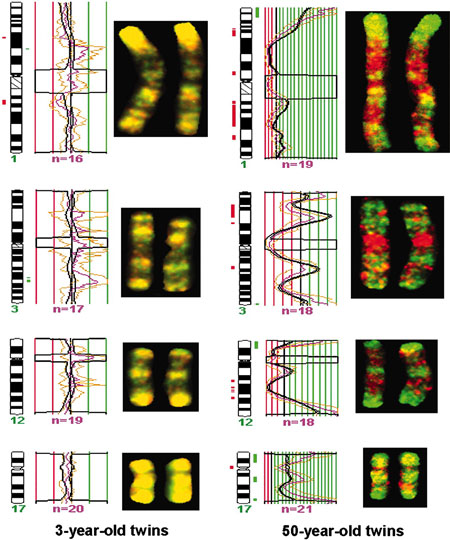
|
| DNA Methylation, young and old monozygous twins.[5] |
Developmental Methylation Changes
Within the embryonic genome DNA methylation occurs at regions of cytosine residues followed by guanines (CpG) and is a main epigenetic mechanism regulating early gene expression and later genomic imprinting, see review.[6] There are extensive changes in DNA methylation state that occur during development and relate to imprinted genes, the total number of genes imprinted in currently unknown, with 100 identified imprinted genes in the mouse and about 50 known in the human. This imprinting will also differ between the embryo and the placenta.[7]
Overall a developmental demethylation is followed by an eventual remethylation of about 70% of all CpGs, except for the primordial germ cell population. The primordial germ cells will form the germ line cell population in the embryo gonad. These cells appear to undergo epigenetic reprogramming through an independent genome-wide of erasure of imprints and epimutations through cytidine deaminases.[8]
The zygote initially has a male and female pronucleus that will fuse to form the diploid nucleus. Each of the male and female pronuclei have different patterns of methylation and undergo different demethylation processes. The male pronucleus undergoes an extensive loss of DNA methylation, with imprinted genes resisting this process. The female pronucleus has less active demethylation requiring several mitotic rounds to complete this process.[9]
Primordial Germ Cells
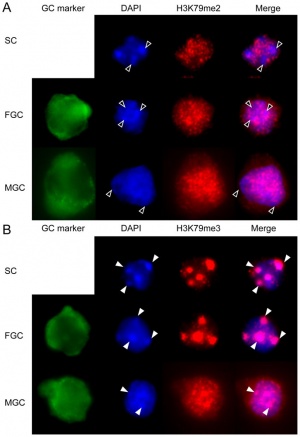
Mouse primordial germ cell DNA methylation[10] |
Demethylation
|
DNA Demethylation
Activation-Induced cytidine Deaminase
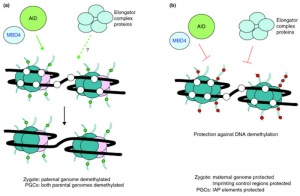
Activation-Induced cytidine Deaminase (AID) is an enzyme required for demethylation (removal of CpG methylation). Within the genome, DNA methylation is associated with epigenetic mechanisms and occurs at cytosine residues that are followed by guanines.[9] This enzyme is also found expressed in primordial germ cells.
Histone Modification
Histones are a family of proteins involved in the bundling of genomic DNA into chromatin. The unit size of chromatin DNA + associated histones is described as the nucleosome. A group of histone modifying enzymes can modify histone protein NH2-terminal tails by: acetylation, methylation, phosphorylation, sumoylation, or ubiquitination. These modifications of histone proteins can in turn determine the accessibility of the DNA to the transcription machinery.
Histone Acetylation
The lysine residues on histone tails can have acetyl groups either removed (by histone deacetyltransferases, HDACs) or added (by histone acetyl transferases, HATs).
- Normally the lysine residues on histone tails bear a positive charge that can bind negatively charged DNA to form a condensed structure with low transcriptional activity.
- Histone acetylation removes these positive charges allowing a less condensed structure with higher transcriptional activity.
Potential Imprinted Genes
|
|
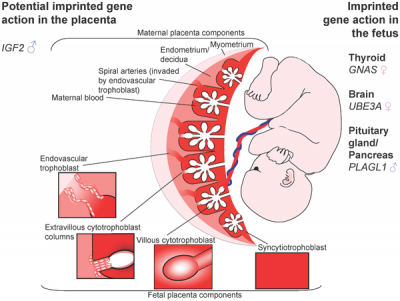
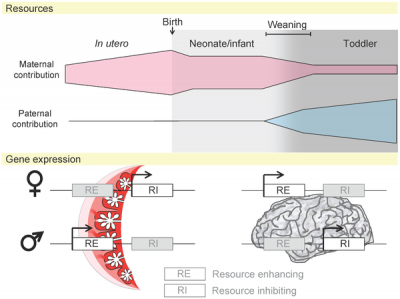
| Placenta potential imprinted genes[7] | Maternal and paternal resource allocation[7] |
References
- ↑ <pubmed>16688142</pubmed>
- ↑ <pubmed>21297937</pubmed>
- ↑ PLoS Collection - Epigenetics 2010
- ↑ <pubmed>20644535</pubmed>
- ↑ <pubmed>16009939</pubmed>
- ↑ <pubmed>20236475</pubmed>
- ↑ 7.0 7.1 7.2 <pubmed>20617174</pubmed>
- ↑ <pubmed>20098412</pubmed>
- ↑ 9.0 9.1 9.2 <pubmed>20236475</pubmed>
- ↑ <pubmed>21886830</pubmed>| PLoS One.
Search Pubmed
June 2010 "epigenetics" - All (37097) Review (6256) Free Full Text (15778)
Search Pubmed Now: epigenetics
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, June 16) Embryology Molecular Development - Epigenetics. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Molecular_Development_-_Epigenetics
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G