Cell Division - Mitosis: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| (11 intermediate revisions by the same user not shown) | |||
| Line 2: | Line 2: | ||
== Introduction == | == Introduction == | ||
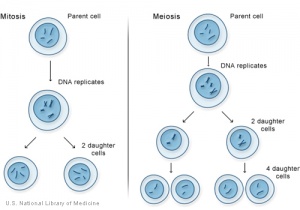
[[Image:Mitosis meiosis1.jpg|thumb|alt=cartoon of mitosis and meiosis|Mitosis and meiosis]] | [[Image:Mitosis meiosis1.jpg|thumb|alt=cartoon of mitosis and meiosis|Mitosis and meiosis]] | ||
Normal cell division in all cells, except germ cells, occurs by 2 mechanical processes that initially divide the nucleus then the cell cytoplasm. This process produces two (daughter) cells that should be genetically identical to the parent cell. | Normal cell division in all cells, except germ cells, occurs by 2 mechanical processes that initially divide the nucleus then the cell cytoplasm. This process produces two (daughter) cells that should be genetically identical to the parent cell. Note that DNA duplication (replication) occurs during interphase (S phase), before mitosis and not during mitosis. | ||
| Line 27: | Line 27: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* ''' | * '''Review - Mosaicism in Preimplantation Human Embryos: When Chromosomal Abnormalities Are the Norm'''{{#pmid:28457629|PMID28457629}} "Along with errors in meiosis, mitotic errors during post-zygotic cell division contribute to pervasive aneuploidy in human embryos. Relatively little is known, however, about the genesis of these errors or their fitness consequences. Rapid technological advances are helping to close this gap, revealing diverse molecular mechanisms contributing to mitotic error. These include altered cell cycle checkpoints, aberrations of the centrosome, and failed chromatid cohesion, mirroring findings from cancer biology. Recent studies are challenging the idea that mitotic error is abnormal, emphasizing that the fitness impacts of mosaicism depend on its scope and severity. In light of these findings, technical and philosophical limitations of various screening approaches are discussed, along with avenues for future research." | ||
|} | |} | ||
{| class="wikitable mw-collapsible mw-collapsed" | {| class="wikitable mw-collapsible mw-collapsed" | ||
| Line 34: | Line 34: | ||
| [[File:Mark_Hill.jpg|90px|left]] {{Most_Recent_Refs}} | | [[File:Mark_Hill.jpg|90px|left]] {{Most_Recent_Refs}} | ||
Search term: [http://www.ncbi.nlm.nih.gov/pubmed/?term=Mitosis ''Mitosis''] | Search term: [http://www.ncbi.nlm.nih.gov/pubmed/?term=Mitosis ''Mitosis''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Prophase ''Prophase''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Prometaphase ''Prometaphase''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Metaphase ''Metaphase''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Anaphase ''Anaphase''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Telophase ''Telophase''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Cytokinesis ''Cytokinesis''] | ||
|} | |||
{| class="wikitable mw-collapsible mw-collapsed" | |||
! Older papers | |||
|- | |||
| {{Older papers}} | |||
* '''Golgi apparatus self-organizes into the characteristic shape via postmitotic reassembly dynamics'''{{#pmid:28461510|PMID28461510}} "The Golgi apparatus is a membrane-bounded organelle with the characteristic shape of a series of stacked flat cisternae. During mitosis in mammalian cells, the Golgi apparatus is once fragmented into small vesicles and then reassembled to form the characteristic shape again in each daughter cell. The mechanism and details of the reassembly process remain elusive. ...We show that the characteristic Golgi shape is spontaneously organized from the assembly of vesicles by proper tuning of the two additional mechanisms, i.e., the Golgi reassembly process is modeled as self-organization. We also demonstrate that the fine Golgi shape forms via a balance of three reaction speeds: vesicle aggregation, membrane fusion, and shape relaxation. Moreover, the membrane fusion activity decreases thickness and the number of stacked cisternae of the emerging shapes." | |||
* '''The nucleoporin ELYS/Mel28 regulates nuclear envelope subdomain formation in HeLa cells'''{{#pmid:22555603|PMID22555603}} "In open mitosis, the nuclear envelope (NE) reassembles at the end of each mitosis. This process involves the reformation of the nuclear pore complex (NPC), the inner and outer nuclear membranes, and the nuclear lamina. In human cells, cell cycle-dependent NE subdomains exist, characterized as A-type lamin-rich/NPC-free or B-type lamin-rich/NPC-rich, which are initially formed as core or noncore regions on mitotic chromosomes, respectively. Although postmitotic NE formation has been extensively studied, little is known about the coordination of NPC and NE assembly. ...Our data show, that ELYS/Mel28 plays a role in NE subdomain formation in late mitosis." | |||
|} | |} | ||
==Movies== | ==Movies== | ||
| Line 45: | Line 52: | ||
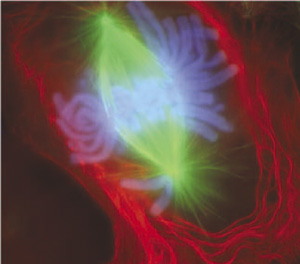
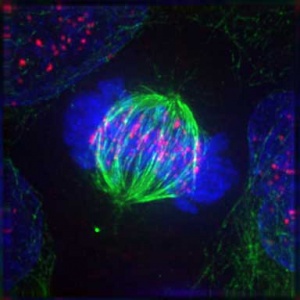
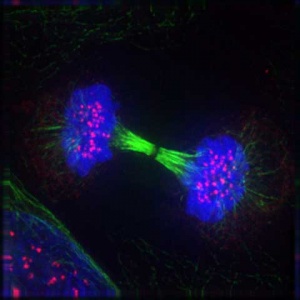
'''Mitosis''' | '''Mitosis''' | ||
This movie shows a cell dividing by mitosis with a fluorescently labelled protein that is located at the kinetochores and along the axes of the chromosome arms. This allows you to see the chromosomes and the linking region (kinetochore) between chromosome pairs and the mitotic spindle microtubules. | This movie shows a cell dividing by mitosis with a fluorescently labelled protein that is located at the kinetochores and along the axes of the chromosome arms. This allows you to see the chromosomes and the linking region (kinetochore) between chromosome pairs and the mitotic spindle microtubules.{{#pmid:12105179|PMID12105179}} | ||
{{Mitosis movie 1}} | {{Mitosis movie 1}} | ||
| Line 62: | Line 69: | ||
:'''Links:''' [http://www.ncbi.nlm.nih.gov/books/bv.fcgi?&rid=mcb.figgrp.5500 MCB Movie - The stages of mitosis and cytokinesis in an animal cell] | :'''Links:''' [http://www.ncbi.nlm.nih.gov/books/bv.fcgi?&rid=mcb.figgrp.5500 MCB Movie - The stages of mitosis and cytokinesis in an animal cell] | ||
==Early Mitosis== | |||
{| | |||
| In early development to the {{morula}} stage, cells are undergoing rapid cell mitotic divisions and do not have the usual G1 and G2 checkpoint regulation.{{#pmid:19288185|PMID19288185}} | |||
During M-phase, cells still maintain the spindle assembly checkpoint, but not apoptosis activation.{{#pmid:28159965|PMID28159965}} The spindle assembly checkpoint appears to also be the key to mitotic cell cycle progression during these early cell divisions.{{#pmid:21720555|PMID21720555}} | |||
| <html5media height="250" width="260">File:Mouse_zygote_division.mp4</html5media> | |||
Early {{mouse}} {{mitosis}} | |||
|} | |||
==Cell Changes== | ==Cell Changes== | ||
| Line 135: | Line 154: | ||
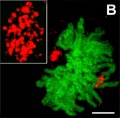
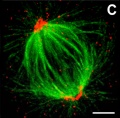
===Anaphase=== | ===Anaphase=== | ||
[[ | [[File:Chromosome_motility_anaphase.jpg|thumb|Chromosome motility anaphase]] | ||
* Separation of sister Kinetochores | * Separation of sister Kinetochores | ||
* shortening of Kinetochore microtubules pulls chromosome to spindle pole. | * shortening of Kinetochore microtubules pulls chromosome to spindle pole. | ||
| Line 185: | Line 204: | ||
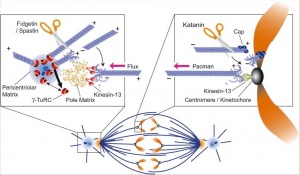
[[File:Spindle assembly motors 01.jpg|800px]] | [[File:Spindle assembly motors 01.jpg|800px]] | ||
Spindle assembly motors | Spindle assembly motors{{#pmid:26668328|PMID26668328}} | ||
Microtubule (MT)-bound motors promote bipolar spindle formation, whereas chromosome-associated motors drive proper kinetochore orientation and chromosome movement to the equator. | Microtubule (MT)-bound motors promote bipolar spindle formation, whereas chromosome-associated motors drive proper kinetochore orientation and chromosome movement to the equator. | ||
| Line 212: | Line 231: | ||
[[File:Mitosis_peroxisomes_01.jpg|600px]] | [[File:Mitosis_peroxisomes_01.jpg|600px]] | ||
Peroxisome (red) location at Interphase (a) and during Mitosis (b and c) | Peroxisome (red) location at Interphase (a) and during Mitosis (b and c){{#pmid:19194514|PMID19194514}} | ||
<gallery> | <gallery> | ||
File:Mitosis_peroxisomes_02.jpg|Interphase | File:Mitosis_peroxisomes_02.jpg|Interphase | ||
| Line 223: | Line 242: | ||
===Golgi=== | ===Golgi=== | ||
* 2 processes - disassembly and reassembly | * 2 processes - disassembly and reassembly{{#pmid:18156178|PMID18156178}} | ||
* Golgi stack undergoes a continuous fragmentation process | * Golgi stack undergoes a continuous fragmentation process | ||
* fragments are distributed into daughter cells | * fragments are distributed into daughter cells | ||
Latest revision as of 10:41, 2 August 2019
| Embryology - 26 Jun 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
Normal cell division in all cells, except germ cells, occurs by 2 mechanical processes that initially divide the nucleus then the cell cytoplasm. This process produces two (daughter) cells that should be genetically identical to the parent cell. Note that DNA duplication (replication) occurs during interphase (S phase), before mitosis and not during mitosis.
Germ cells, oocyte and spermatozoa, undergo meiotic cell division.
- Mitosis segregation of chromosomes and formation of 2 nuclei
- Cytokinesis splitting of the cell as a whole into 2 daughter cells
- Recent Nobel Prizes- 2001 Cell Cycle, 2002 Cell Death
| Mitosis of the single zygote produces how many cells in the adult? |
|---|
| Not a straightforward calculation! Cells through development and in the adult divide and die influenced by cell type, genetics and environment. In the adult, most tissues though would have a relatively contestant balance between loss and gain in cell numbers. A recent paper has attempted to estimate the number for humans[1] "In particular, the reported total cell number of a human being ranges between 1012 and 1016 and it is widely mentioned without a proper reference. ...A current estimation of human total cell number calculated for a variety of organs and cell types is presented. These partial data correspond to a total number of 3.72 × 1013." |
| Cell Division Links: meiosis | mitosis | Lecture - Cell Division and Fertilization | spermatozoa | oocyte | fertilization | zygote | Genetics |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Mitosis | Prophase | Prometaphase | Metaphase | Anaphase | Telophase | Cytokinesis |
| Older papers |
|---|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
|
Movies
| <html5media height="420" width="400">File:Mitosis 01.mp4</html5media> |
Mitosis This movie shows a cell dividing by mitosis with a fluorescently labelled protein that is located at the kinetochores and along the axes of the chromosome arms. This allows you to see the chromosomes and the linking region (kinetochore) between chromosome pairs and the mitotic spindle microtubules.[5]
|
|
|
|
|
|
Early Mitosis
| In early development to the morula stage, cells are undergoing rapid cell mitotic divisions and do not have the usual G1 and G2 checkpoint regulation.[6]
|
<html5media height="250" width="260">File:Mouse_zygote_division.mp4</html5media> |
Cell Changes
- Nucleus
- Chromosome condensation
- Nuclear envelope breakdown
- Cytoplasm
- Cytoskeleton reorganization
- Spindle formation (MT) Contractile ring (MF)
- Organelle redistribution
- Mitosis Energy
- Cell division uses up a lot of energy, so cells ensure they have enough resources to complete the job before committing to it.
Mitosis Phases
- Based on light microscopy of living cells light and electron microscopy of fixed and stained cells
- 5 Phases - prophase, prometaphase, metaphase, anaphase, and telophase
- Cytokinesis 6th stage overlaps the end of mitosis
MBC The stages of mitosis and cytokinesis in an animal cell
Interphase
- not a mitotic phase (discussed in cell cycle)
- Chromosomes dispersed in nucleus
- Gene expression
- Cytoskeleton and cell organelles - Distributed and functioning
- Mitochondria undergo independent proliferation/division
Chromosome Changes
Prophase
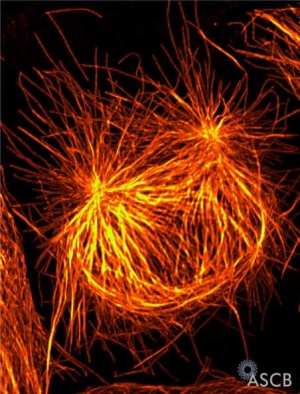
- Chromosome DNA has been earlier duplicated (S Phase)
- Chromosomes begin condensing
- Chromosome pairs (chromatids) held together at centromere
- Microtubules disassemble
- Mitotic spindle begins to form
Spindle Apparatus
- 3 sets of microtubules - (+) ends point away from centrosome at each pole.
- astral microtubules - anchor the pole end in position
- kinetochore microtubules - connected to chromosomes
- polar microtubules - form the structure of the spindle apparatus
Spindle Apparatus EM | Spindle Apparatus | MBC Movie- Microtubule dynamics during mitosis
At end of prophase nuclear envelope breaks down
Prometaphase
- Microtubules now enter nuclear region
- Nuclear envelope forms vesicles around mitotic spindle
- Kinetochores form on centromere attach to some MTs of spindle
Dynamic instability and the capture of chromosomes
Centromeric attachment of microtubules
At end of prometaphase chromosomes move to metaphase plate
Metaphase
- Kinetochore MTs align chromosomes in one midpoint plane.
- Astrin is a spindle-associated protein required for chromosome alignment at the metaphase plate.[10]
Proposed alternative mechanisms for chromosome congression
Metaphase ends when sister kinetochores separate
Anaphase
- Separation of sister Kinetochores
- shortening of Kinetochore microtubules pulls chromosome to spindle pole.
- Katanin is a microtubule-severing complex involved with this stage of microtubule dynamics.[11]
Anaphase ends as nuclear envelope (membrane) begins to reform.
Telophase
- Chromosomes arrive at spindle poles
- Kinetochore MTs lost
- Condensed chromosomes begin expanding
- Continues through cytokinesis
Links: Figure 19-41 Microtubule dynamics during mitosis | Figure 19-34. The stages of mitosis and cytokinesis in an animal cell | Cytokinetic abscission: cellular dynamics at the midbody
Cleavage of Zygote
Mouse zygote mitosis[12]
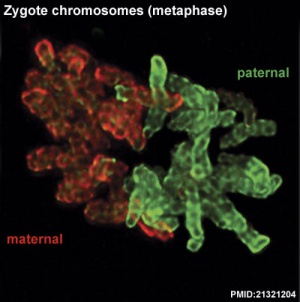
|
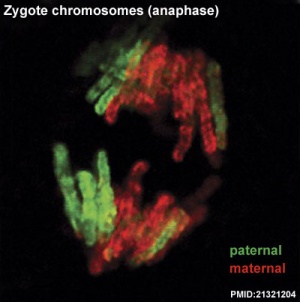
|
| First metaphase | First anaphase |
Cleavage of the zygote forms 2 blastomeres and is cleavage with no cytoplasm synthesis.
- special "embryonic" cell cycle S phases and M phases alternate without any intervening G1 or G2 phases (MSMSMSMS, adult MG1SG2) therefore individual cell volume decreases
Cell division within these cells is initially synchronous (at the same time), then becomes asynchronously (at different times).
- slow- centre cells, larger fast- peripheral cells
- Links: Zygote | Cell Division - Mitosis | Movie - Early Cell Division | Movie - Week 1 Cell Cleavage | Carnegie stage 1
Cytokinesis
- Division of cytoplasmic contents
- Contractile ring forms at midpoint under membrane
- Microfilament ring - contracts forming cleavage furrow
- myosin II is the motor
- Eventually fully divides cytoplasm
Links: Cytokinesis | Cytokinesis in Plants
Mitotic Spindle
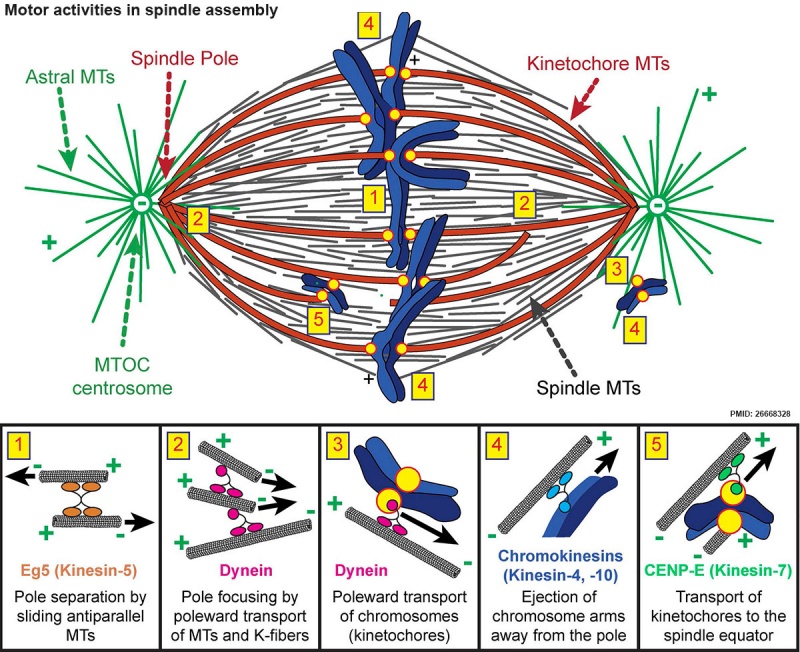
Spindle assembly motors[13]
Microtubule (MT)-bound motors promote bipolar spindle formation, whereas chromosome-associated motors drive proper kinetochore orientation and chromosome movement to the equator.
| Box 1 | Box 2 | Box 3 | Box 4 | Box 5 |
|---|---|---|---|---|
| Motor-dependent mechanisms establish bipolarity as Eg5 (kinesin-5) motors slide antiparallel microtubules apart with their minus ends leading and their plus ends directed toward the spindle equator. | Minus end–directed motors such as dynein move microtubules poleward with their minus ends leading, thereby incorporating K-fibers into the spindle and focusing spindle poles. | Kinetochore-associated dynein transports chromosomes along astral microtubules toward the spindle poles from the periphery. | Plus end–directed chromokinesins (kinesin-4 and -10) eject chromosome arms outward. | CENP-E (kinesin-7) transports unattached kinetochores toward the equator along spindle microtubules. MTOC, microtubule organizing centre. |
Cell Organelles
Mitochondria
- Divide independently of cell mitosis
- distributed into daughter cells
Peroxisomes
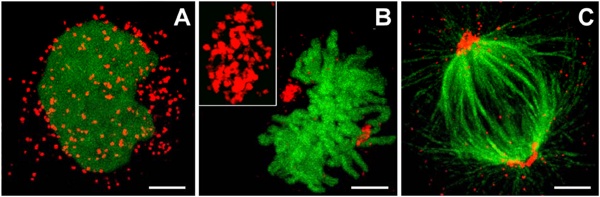
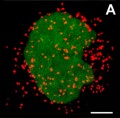
- localise at spindle poles
Peroxisome (red) location at Interphase (a) and during Mitosis (b and c)[14]
Endoplasmic Reticulum
- Associated with nuclear membrane.
Golgi
- 2 processes - disassembly and reassembly[15]
- Golgi stack undergoes a continuous fragmentation process
- fragments are distributed into daughter cells
- are reassembled into new Golgi stacks
Disassembly
- Unstacking - mediated by two mitotic kinases (cdc2 and plk)
- Vesiculation - mediated by COPI budding machinery ARF1 and the coatomer complex
Reassembly
- Fusion - formation of single cisternae by membrane fusion
- Restacking - requires dephosphorylation of Golgi stacking proteins by protein phosphatase PP2A
References
- ↑ <pubmed>23829164</pubmed>
- ↑ McCoy RC. (2017). Mosaicism in Preimplantation Human Embryos: When Chromosomal Abnormalities Are the Norm. Trends Genet. , 33, 448-463. PMID: 28457629 DOI.
- ↑ Tachikawa M & Mochizuki A. (2017). Golgi apparatus self-organizes into the characteristic shape via postmitotic reassembly dynamics. Proc. Natl. Acad. Sci. U.S.A. , 114, 5177-5182. PMID: 28461510 DOI.
- ↑ Clever M, Funakoshi T, Mimura Y, Takagi M & Imamoto N. (2012). The nucleoporin ELYS/Mel28 regulates nuclear envelope subdomain formation in HeLa cells. Nucleus , 3, 187-99. PMID: 22555603 DOI.
- ↑ Tavormina PA, Côme MG, Hudson JR, Mo YY, Beck WT & Gorbsky GJ. (2002). Rapid exchange of mammalian topoisomerase II alpha at kinetochores and chromosome arms in mitosis. J. Cell Biol. , 158, 23-9. PMID: 12105179 DOI.
- ↑ Kiessling AA, Bletsa R, Desmarais B, Mara C, Kallianidis K & Loutradis D. (2009). Evidence that human blastomere cleavage is under unique cell cycle control. J. Assist. Reprod. Genet. , 26, 187-95. PMID: 19288185 DOI.
- ↑ Jacobs K, Van de Velde H, De Paepe C, Sermon K & Spits C. (2017). Mitotic spindle disruption in human preimplantation embryos activates the spindle assembly checkpoint but not apoptosis until Day 5 of development. Mol. Hum. Reprod. , 23, 321-329. PMID: 28159965 DOI.
- ↑ Wei Y, Multi S, Yang CR, Ma J, Zhang QH, Wang ZB, Li M, Wei L, Ge ZJ, Zhang CH, Ouyang YC, Hou Y, Schatten H & Sun QY. (2011). Spindle assembly checkpoint regulates mitotic cell cycle progression during preimplantation embryo development. PLoS ONE , 6, e21557. PMID: 21720555 DOI.
- ↑ Russan NM. Let's Build a Spindle. ASCB Image & Video Library. 2008;CYT-190. Available at: http://cellimages.ascb.org/u?/p4041coll12,521
- ↑ <pubmed>21402792</pubmed>
- ↑ <pubmed>17452528</pubmed>
- ↑ <pubmed>21321204</pubmed>| PMC2132672 | PNAS
- ↑ Heald R & Khodjakov A. (2015). Thirty years of search and capture: The complex simplicity of mitotic spindle assembly. J. Cell Biol. , 211, 1103-11. PMID: 26668328 DOI.
- ↑ Kredel S, Oswald F, Nienhaus K, Deuschle K, Röcker C, Wolff M, Heilker R, Nienhaus GU & Wiedenmann J. (2009). mRuby, a bright monomeric red fluorescent protein for labeling of subcellular structures. PLoS ONE , 4, e4391. PMID: 19194514 DOI.
- ↑ Tang D, Mar K, Warren G & Wang Y. (2008). Molecular mechanism of mitotic Golgi disassembly and reassembly revealed by a defined reconstitution assay. J. Biol. Chem. , 283, 6085-94. PMID: 18156178 DOI.
Reviews
Articles
Search Pubmed
Search Pubmed: mitosis
NCBI - Policies and Guidelines | PubMed | Help:Reference Tutorial
Additional Images
Terms
| Cell Division Terms (expand to view) | ||
|---|---|---|
meiosis | mitosis
| ||
|
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- Nature - Cell Division Milestones
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, June 26) Embryology Cell Division - Mitosis. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Cell_Division_-_Mitosis
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G