Molecular Development - Epigenetics: Difference between revisions
| Line 31: | Line 31: | ||
| Maternal and paternal resource allocation<ref name="PMID20617174" /> | | Maternal and paternal resource allocation<ref name="PMID20617174" /> | ||
|} | |} | ||
==Activation-Induced cytidine Deaminase== | |||
[[File:Mammalian active DNA demethylation.jpg|thumb|Mammalian active DNA demethylation<ref name="PMID20236475"><pubmed>20236475</pubmed></ref>]] | |||
Activation-Induced cytidine Deaminase (AID) is an enzyme required for demethylation (removal of CpG methylation). Within the genome, DNA methylation is associated with epigenetic mechanisms and occurs at cytosine residues that are followed by guanines.<ref name="PMID20236475"><pubmed>20236475</pubmed></ref> This enzyme is also found expressed in primordial germ cells. | |||
:'''Links:''' [[Primordial Germ Cell Development]] | |||
==References== | ==References== | ||
Revision as of 13:41, 19 November 2010
Introduction
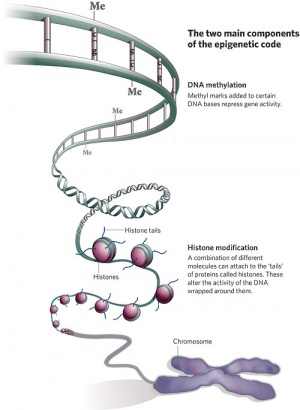
In terms of molecular mechanisms, the field of epigenetics has begun to florish with some recent important findings. Epigenetics as the name implies, is the inheritance mechanisms that lie outside the DNA sequence of our genes. One of the initial discoveries was the effects of DNA methylation upon gene expression and then modifications of nucleosomal histones. This DNA methylation, usually associated with 5-methylcytosine (m5C), leads to transcriptional silencing in vertebrates.
Molecular mechanisms of development is an exciting area and requires a variety of different skills. This page introduces only a few examples and should give you a feel for the topic. Note that each section of system notes has a page covering molecular development in that system.
Some Recent Findings
|
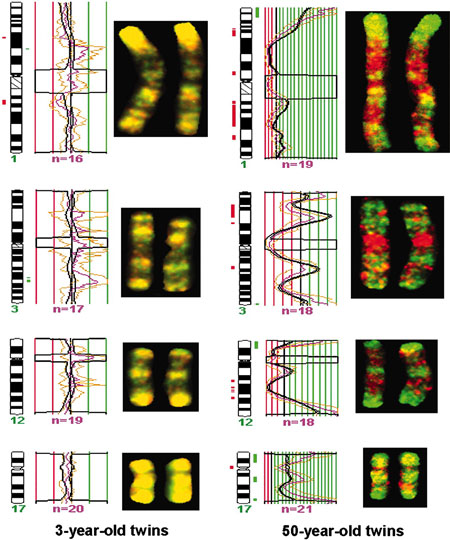
DNA Methylation Changes with Age
|
| DNA Methylation, young and old monozygous twins.[4] |
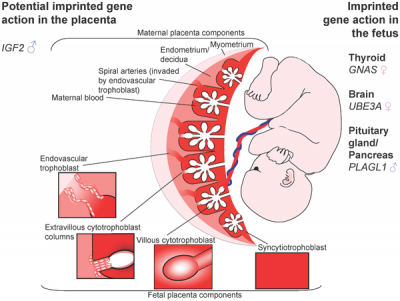
Potential Imprinted Genes
|
|
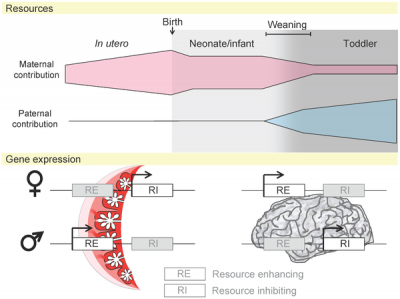
| Placenta potential imprinted genes[5] | Maternal and paternal resource allocation[5] |
Activation-Induced cytidine Deaminase
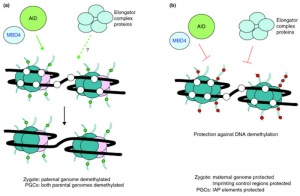
Activation-Induced cytidine Deaminase (AID) is an enzyme required for demethylation (removal of CpG methylation). Within the genome, DNA methylation is associated with epigenetic mechanisms and occurs at cytosine residues that are followed by guanines.[6] This enzyme is also found expressed in primordial germ cells.
References
Search Pubmed
June 2010 "epigenetics" - All (37097) Review (6256) Free Full Text (15778)
Search Pubmed Now: epigenetics
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, June 3) Embryology Molecular Development - Epigenetics. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Molecular_Development_-_Epigenetics
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G