Developmental Mechanism - Apoptosis
| Embryology - 5 May 2026 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
This single term "apoptosis" describes the way in which the majority of cells die within our adult body are removed every day, "Programmed Cell Death". In development, apoptosis begins in the early blastocyst and is a developmental mechanism found throughout tissues in the embryo and fetus developmental stages. In addition to the many developmental roles this process is used in multicellular organisms to remove cells that are: aged, superfluous, infected, contain genetic errors or are transformed.
While the cellular morphological changes associated with this process are the same in all cells, there are many different signaling pathways that can "trigger" this process. They fall generally into two signalling classes either intrinsic or extrinsic to the cell. To describe "programmed cell death" as apoptosis was originally used in 1972 by Kerr, Wyllie and Currie.[2]
Cell Biology Lecture - Cell Death
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Development Apoptosis |
| Older papers |
|---|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page. |
Developmental Examples
Blastocyst Development
| Removing cells from the developing inner cell mass.
|
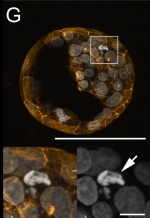
Cell death in bovine blastocyst[5] |
Cloacal Membrane Development
The initial cloaca is the common early endoderm lined space of the hindgut that will later become partitioned by a septum into a dorsal gastrointestinal component (rectum) and ventral renal/genital component (urogenital sinus). Located at the inferior end of the cloaca is the cloacal membrane, that also forms part of the embryo surface. The cloacal membrane is formed during gastrulation by ectoderm and endoderm without a middle (intervening) layer of mesoderm, and later degenerates by apoptosis[6][7] after cloacal septation in mammals.
- Links: cloacal membrane
Digit Development
| Removing cells between digits (fingers and toes) of the upper and lower limbs.
|
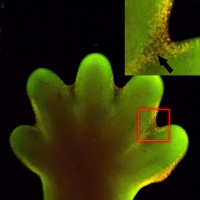
Interdigital apoptosis in the mous hindlimb.[8] |
Bone Development
During ossification removing chondrocytes.
- Links: bone
Neural development
Removing excess or inappropriately connected neurons.
- Migratory neural crest Cells Phagocytose Dead Cells in the Developing Nervous System{{#pmid:31495570|PMID31495570}} "During neural tube closure and spinal cord development, many cells die in both the central and peripheral nervous systems (CNS and PNS, respectively). However, myeloid-derived professional phagocytes have not yet colonized the trunk region during early neurogenesis. How apoptotic cells are removed from this region during these stages remains largely unknown. Using live imaging in zebrafish, we demonstrate that neural crest cells (NCCs) respond rapidly to dying cells and phagocytose cellular debris around the neural tube. Additionally, NCCs have the ability to enter the CNS through motor exit point transition zones and clear debris in the spinal cord. Surprisingly, NCCs phagocytosis mechanistically resembles macrophage phagocytosis and their recruitment toward cellular debris is mediated by interleukin-1β. Taken together, our results reveal a role for NCCs in phagocytosis of debris in the developing nervous system before the presence of professional phagocytes."
- Links: neural
Ovary Development
Removing excess primordial follicles from the ovary cortex.
- Links: ovary
Mammary Development
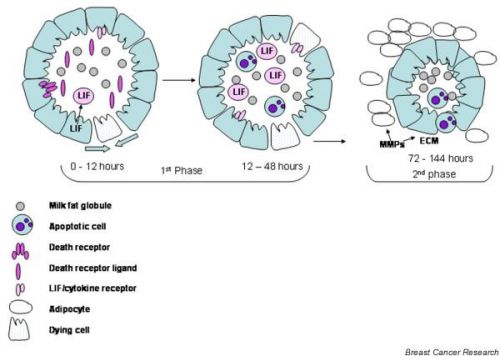
Mammary involution[9]
|
|
Full involution returns the gland to a pre-pregnant state. ECM, extracellular matrix.
- Links: mammary gland
Renal Development
In the rat model of renal development, apoptotic death has been found in both the nephrogenic region and medullary papilla.[10] (See recent reviewPubmedParser error: The PubmedParser extension received invalid XML data. ())
- Links: {{Renal))
Genital Development
Reciprocal Spatiotemporally Controlled apoptosis Regulates Wolffian Duct Cloaca Fusion[11]
- "The epithelial Wolffian duct (WD) inserts into the cloaca (primitive bladder) before metanephric kidney development, thereby establishing the initial plumbing for eventual joining of the ureters and bladder. Defects in this process cause common anomalies in the spectrum of congenital anomalies of the kidney and urinary tract (CAKUT). However, developmental, cellular, and molecular mechanisms of WD-cloaca fusion are poorly understood. Through systematic analysis of early WD tip development in mice, we discovered that a novel process of spatiotemporally regulated apoptosis in WD and cloaca was necessary for WD-cloaca fusion. Aberrant RET tyrosine kinase signaling through tyrosine (Y) 1062, to which PI3K- or ERK-activating proteins dock, or Y1015, to which PLCγ docks, has been shown to cause CAKUT-like defects. Cloacal apoptosis did not occur in RetY1062F mutants, in which WDs did not reach the cloaca, or in RetY1015F mutants, in which WD tips reached the cloaca but did not fuse. Moreover, inhibition of ERK or apoptosis prevented WD-cloaca fusion in cultures, and WD-specific genetic deletion of YAP attenuated cloacal apoptosis and WD-cloacal fusion in vivo Thus, cloacal apoptosis requires direct contact and signals from the WD tip and is necessary for WD-cloacal fusion. These findings may explain the mechanisms of many CAKUT."
Worm Development
The overview diagram above shows the fate of each individual cell in the developing c. elegans.
- the "X" indicates cells that die by apoptosis during development.
- Zygote (P0 cell) divides into two daughter cells (AB and P1 cells).
- These two daughter cells then divide into the next generation.
Note the above image is not at a readable resolution, to view see large readable version (10,389 × 1,336 pixels). Embryonic cell lineage developed by J .E. Sulston, E. Schierenberg, J. G. White, J. N. Thomson.
Nobel Prize 2002
The 2002 Nobel Prize in Physiology or Medicine went to three researchers who originally identified this mechanism in the genetic regulation of organ development and programmed cell death.
- Sydney Brenner (b 1927), Berkeley, CA, USA, established C. elegans as a novel experimental model organism. This provided a unique opportunity to link genetic analysis to cell division, differentiation and organ development – and to follow these processes under the microscope. Brenner's discoveries, carried out in Cambridge, UK, laid the foundation for this year's Prize.
- John Sulston (b 1942), Cambridge, England, mapped a cell lineage where every cell division and differentiation could be followed in the development of a tissue in C. elegans. He showed that specific cells undergo programmed cell death as an integral part of the normal differentiation process, and he identified the first mutation of a gene participating in the cell death process.
- Robert Horvitz (b 1947), Cambridge, MA, USA, has discovered and characterized key genes controlling cell death in C. elegans. He has shown how these genes interact with each other in the cell death process and that corresponding genes exist in humans.
- Links: Nobel Prize 2002
Adult Examples
Bacterial Infection
| <html5media height="620" width="620">File:Cell apoptosis 02.mp4</html5media> | This movie from an in vitro experimental bacterial infection of Hela cells with Neisseria gonorrhoeae shows the cells dying by apoptosis.
|
Apoptotic Cell Morphology
The following cellular changes occur in sequence during apoptosis.
- loss of cell membrane phospholipid asymmetry
- Condensation of chromatin
- Reduction in nuclear size JCB - Nucleus changes
- Internucleosomal DNA cleavage TUNEL staining
- DNA ladder
- shrinkage of the cell
- Cleavage of cytoskeletal proteins PNAS - Actin cleavage by ICE-like proteases
- note actin also binds DNase 1, cleavage may release this enzyme to further cleave DNA
- membrane blebbing
- breakdown of the cell into membrane-bound apoptotic bodies (apoptosomes)
- bodies then phagocytosed by other cells
Experimentally a number of different techniques have been developed and are now used to identify these changes.
- Links:
Apoptosis Regulators
Regulators can initiate or block apoptosis, the regulators shown block apoptosis.
| Regulator → | Adaptor → | Effector | |||
|---|---|---|---|---|---|
| C. elegans | Ced-9 → | Ced-4 → | Ced-3 → | Death | |
| Vertebrates | Bcl-2 → | Apaf-1 → | Caspase-9 → | Caspase-3 → | Death |
p63
p63 (tumor protein p63) gene (3q28) encodes a transcription factor with multiple isotypes that transactivate p53 reporter genes and induce apoptosis. For example, during palate development p63 can exert spatio-temporal control of palatal epithelial cell fate to prevent cleft palate [12]
- Links: OMIM p63
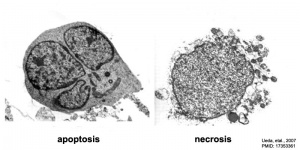
Necrosis
Necrosis (Greek, nekros = corpse) is not a form of developmental cell death, but a pathological cell death from extrinsic injury (tissue damage) and is irreversible. Necrosis is induced by tumor necrosis factor, double-stranded RNA, viral infection or bacterial infection toxins.
In the early stages, cell and organelles (mitochondria) swell (oncosis) (Greek, onkos = 'swelling') previously described as a separate form of cell death. This is due to disruption of plasma membrane leading to cell contents leak out leading to inflammation and necrosis.
In the late stages, there is a loss of cell membrane integrity, finally cell disintegration. This cell lysis can also trigger an inflammatory response, leading to further inflammation and damage, and triggering a cycle of death
- Bacteria - S. aureus - pore-forming secreted toxins can induce necrosis
- Prothymosin-alpha 1 - (ProT) inhibits necrosis, switches cells from necrosis to apoptosis
References
- ↑ Ueda H, Fujita R, Yoshida A, Matsunaga H & Ueda M. (2007). Identification of prothymosin-alpha1, the necrosis-apoptosis switch molecule in cortical neuronal cultures. J. Cell Biol. , 176, 853-62. PMID: 17353361 DOI.
- ↑ Kerr JF, Wyllie AH & Currie AR. (1972). Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br. J. Cancer , 26, 239-57. PMID: 4561027
- ↑ Shinotsuka N, Yamaguchi Y, Nakazato K, Matsumoto Y, Mochizuki A & Miura M. (2018). Caspases and matrix metalloproteases facilitate collective behavior of non-neural ectoderm after hindbrain neuropore closure. BMC Dev. Biol. , 18, 17. PMID: 30064364 DOI.
- ↑ Koto A & Miura M. (2011). Who lives and who dies: Role of apoptosis in quashing developmental errors. Commun Integr Biol , 4, 495-7. PMID: 21966582 DOI.
- ↑ Leidenfrost S, Boelhauve M, Reichenbach M, Güngör T, Reichenbach HD, Sinowatz F, Wolf E & Habermann FA. (2011). Cell arrest and cell death in mammalian preimplantation development: lessons from the bovine model. PLoS ONE , 6, e22121. PMID: 21811561 DOI.
- ↑ Miller SA, Clark C, Cooney R, Crary E & Payzant W. (1998). Apoptosis fenestrates chick cloacal membrane and occluded rectal cord and may have a minor role in removal of pharyngeal membranes. Ann. N. Y. Acad. Sci. , 857, 268-71. PMID: 9917854
- ↑ Sasaki C, Yamaguchi K & Akita K. (2004). Spatiotemporal distribution of apoptosis during normal cloacal development in mice. Anat Rec A Discov Mol Cell Evol Biol , 279, 761-7. PMID: 15278947 DOI.
- ↑ Bandyopadhyay A, Tsuji K, Cox K, Harfe BD, Rosen V & Tabin CJ. (2006). Genetic analysis of the roles of BMP2, BMP4, and BMP7 in limb patterning and skeletogenesis. PLoS Genet. , 2, e216. PMID: 17194222 DOI.
- ↑ Watson CJ. (2006). Involution: apoptosis and tissue remodelling that convert the mammary gland from milk factory to a quiescent organ. Breast Cancer Res. , 8, 203. PMID: 16677411 DOI.
- ↑ Coles HS, Burne JF & Raff MC. (1993). Large-scale normal cell death in the developing rat kidney and its reduction by epidermal growth factor. Development , 118, 777-84. PMID: 8076517
- ↑ Hoshi M, Reginensi A, Joens MS, Fitzpatrick JAJ, McNeill H & Jain S. (2018). Reciprocal Spatiotemporally Controlled Apoptosis Regulates Wolffian Duct Cloaca Fusion. J. Am. Soc. Nephrol. , 29, 775-783. PMID: 29326158 DOI.
- ↑ Richardson R, Mitchell K, Hammond NL, Mollo MR, Kouwenhoven EN, Wyatt ND, Donaldson IJ, Zeef L, Burgis T, Blance R, van Heeringen SJ, Stunnenberg HG, Zhou H, Missero C, Romano RA, Sinha S, Dixon MJ & Dixon J. (2017). p63 exerts spatio-temporal control of palatal epithelial cell fate to prevent cleft palate. PLoS Genet. , 13, e1006828. PMID: 28604778 DOI.
Textbooks
Molecular Biology of the Cell
Alberts, Bruce; Johnson, Alexander; Lewis, Julian; Raff, Martin; Roberts, Keith; Walter, Peter New York and London: Garland Science; c2002
- Molecular Biology of the Cell 4th ed. - IV. Internal Organization of the Cell Chapter 17. The Cell Cycle and Programmed Cell Death
- The Cell Cycle and Programmed Cell Death
- Programmed Cell Death (Apoptosis)
- Figure 17-37. Cell death
- Figure 17-47. Two ways in which survival factors suppress apoptosis
- Figure 17-42. Cell-cycle arrest or apoptosis induced by excessive stimulation of mitogenic pathways
- Figure 17-38. The caspase cascade involved in apoptosis
Molecular Cell Biology
Lodish, Harvey; Berk, Arnold; Zipursky, S. Lawrence; Matsudaira, Paul; Baltimore, David; Darnell, James E. New York: W. H. Freeman & Co.; c1999
- Molecular Cell Biology - Chapter 23. Cell Interactions in Development
- Cell Death and Its Regulation
- Figure 23-50. Current models of the intracellular pathways leading to cell death by apoptosis or to trophic factor–mediated cell survival in mammalian cells
- Movie: Apoptosis
The Cell- A Molecular Approach
Cooper, Geoffrey M. Sunderland (MA): Sinauer Associates, Inc.; c2000
- The Cell - A Molecular Approach - IV. Cell Regulation Chapter 13. Cell Signaling
- Regulation of Programmed Cell Death
Reviews
Lin W & Xu G. (2019). Autophagy: A Role in the Apoptosis, Survival, Inflammation, and Development of the Retina. Ophthalmic Res. , 61, 65-72. PMID: 29694961 DOI.
Tuzlak S, Kaufmann T & Villunger A. (2016). Interrogating the relevance of mitochondrial apoptosis for vertebrate development and postnatal tissue homeostasis. Genes Dev. , 30, 2133-2151. PMID: 27798841 DOI.
Pérez-Garijo A & Steller H. (2015). Spreading the word: non-autonomous effects of apoptosis during development, regeneration and disease. Development , 142, 3253-62. PMID: 26443630 DOI.
Domingos PM & Steller H. (2007). Pathways regulating apoptosis during patterning and development. Curr. Opin. Genet. Dev. , 17, 294-9. PMID: 17629474 DOI.
Del Riccio V, van Tuyl M & Post M. (2004). Apoptosis in lung development and neonatal lung injury. Pediatr. Res. , 55, 183-9. PMID: 14630991 DOI.
Articles
Search PubMed
Search Pubmed: developmental apoptosis | developmental cell death | apoptosis
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- UNSW Cell Biology - Lecture - Cell Death
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2026, Mayıs 5) Embryology Developmental Mechanism - Apoptosis. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Mechanism_-_Apoptosis
- © Dr Mark Hill 2026, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G