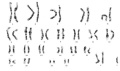Molecular Development - X Inactivation
| Embryology - 21 May 2026 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
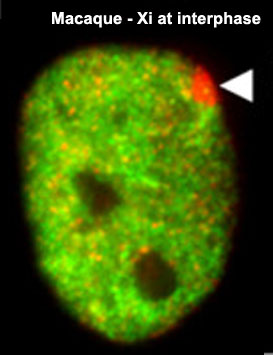
The presence in females of two X chromosome raises the issue of gene dosage, in the case of mammals this is regulated by inactivating one of the X chromosomes. To balance expression with the autosomal chromosomes the dosage imbalance is then adjusted by doubling expression of X-linked genes in both sexes.
In some other species compensation occurs by increasing the expression of X in males. The pattern of which X chromosome is inactivated in cells appears to be random, generating 50% cells expressing Father X, 50% cells expressing Mother X (mosaic pattern). The theory of random X inactivation was first suggested in mice in 1961.[3]
The process of inactivation relies on the Xist RNA, a 17 kb non-coding RNA, which accumulates on the future inactive X chromosome.
A second form of X inactivation that occurs only in male meiotic spermatogenesis, meiotic sex chromosome inactivation (MSCI), is not covered in these current notes. MSCI is the process of transcriptional silencing of the X and Y chromosomes.[4] X inactivation is not an encoded function and occurs randomly throughout female tissues, this represents a form of epigenetics.
Some species have several copies of sex chromosomes, for example the platypus karyotype (2 n = 52) consists of 21 autosomes and 10 sex chromosomes (5X's and 5Y's in male and 5 X-pairs in female). In mice, the paternal X chromosome is silenced. In rabbits, maternal or paternal selection occurs downstream of Xist expression.
| Historic Embryology |
|
Mary Lyon (1925-2014) was a UK geneticist who proposed in 1961 the theory of X inactivation, where one of the two X chromosomes in the cells of female mammals is randomly inactivated during early development. In deference to her, this process is also referred to as "Lyonisation". She also worked on other X-linked genetic diseases, such as Duchenne muscular dystrophy and haemophilia. |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: X inactivation | Xist | Tsix |
| Older papers |
|---|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
|
XIST
The initiation of X chromosome inactivation requires cis accumulation of the large non-translated XIST RNA, which covers the X chromosome, followed by epigenetic changes on the future inactive X chromosome.
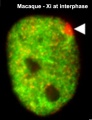
The physical region at the nucleus periphery where the inactive X chromosome is located in a female cell is described as the Barr body.[14]
|
|
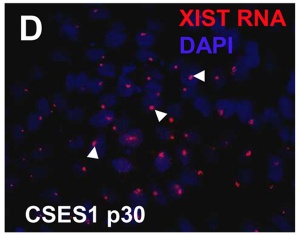
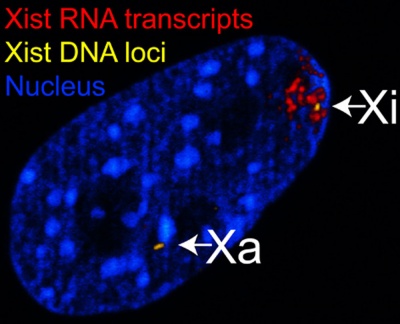
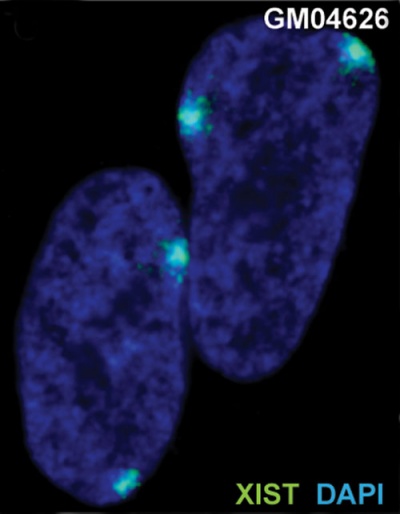
| The above figure shows confocal images from a combined RNA-DNA FISH experiment for Xist in female mouse fibroblast cells (Female-biased expression of long non-coding RNAs in domains that escape X-inactivation in mouse.[15] | XIST expression in two cell nuclei using RNA FISH probes for the human XIST RNA (green) in the human female fibroblast line GM04626 (karyotype: 47,XXX).[16] As there are 3 copies of the X chromosome in each cell, there are 2 inactive X chromosomes as shown by Xist staining. |
- Links: OMIM - Xist
X Inactivation Model
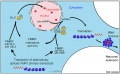
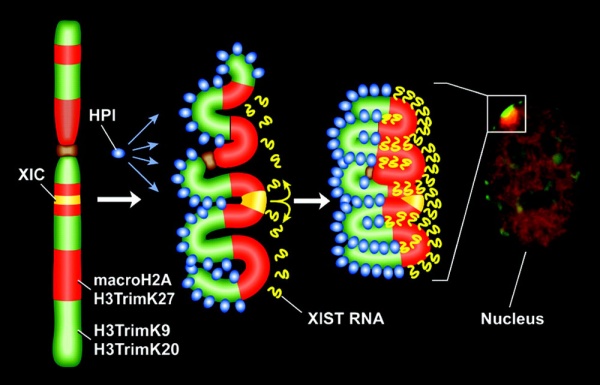
Model for XIST RNA spread from X inactivation center[17]
Above model showing how heterochromatin of the Xi could transition between metaphase and interphase to be organized into the two nonoverlapping heterochromatin territories and to explain how XIST RNA could rapidly spread in cis outward from the X inactivation center (XIC) along only part of the Xi.
Barr Body
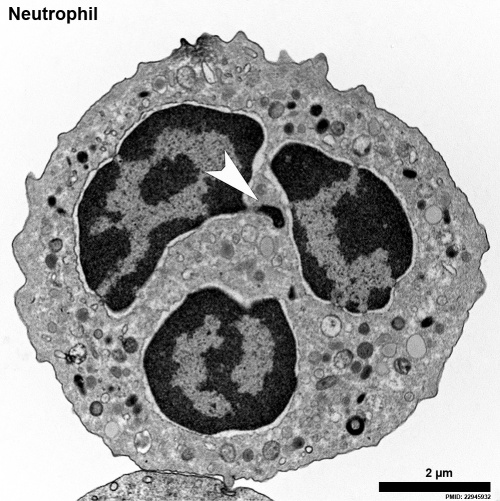
A female human neutrophil - Barr body (White arrowhead)[18]
This neutrophil comes from a female donor and one inactivated x chromosome can be found as an extranuclear stretch of heterochromatin. These structures are termed Barr bodies, and in neutrophils “drum sticks.”
| Historic Embryology |
|
[Murray L. Barr (1908-1995) was a Canadian researcher who, along with E. Bertram, first identified the inactivated x chromosome (Barr body) as an extension from the nucleus in female cells. "An interesting and useful variant is the study of blood films. In females only, there is an appendage with special characteristics and probably containing the sex chromatin, attached to a lobe of the nucleus in a small proportion of neutrophilic leukocytes."[19] |
X Inactivation (xist) History
The theory of random X inactivation was first suggested in mice in 1961.[3] The breakthrough was the discovery of the X inactive specific transcript (XIST).[20] This gene is located within the "X inactivation centre" and only expressed by the inactive X chromosome. Unlike other genes that encode protein XIST contained no "open reading frames" (ie no codons to encode amino acids).
XIST is transcribed but not translated. XIST appears to act as RNA. Current thinking is that it binds to the X Chromosome and is involved in it's translocation to the nuclear periphery. It now appears that XIST appears to initiate X inactivation and it is the methylation of the inactive X genes that maintains inactivity.
- Links: Mary Lyon - X Inactivation
Tsix
A gene antisense to Xist, hence the name which is Xist backwards, in embryonic stem cells Xist repression occurs and Tsix is upregulated. This is part of the overall process of maintaining pluripotency in these cells. Tsix upregulation depends on the recruitment of another pluripotent marker Rex1, and of the associated factors Klf4 and c-Myc, by the DXPas34 minisatellite associated with the Tsix promoter.[21]
DXPas34 is made up of 34bp repeats containing Ctcf motifs and is located within a CpG-rich region in the 5-prime end of the mouse Tsix gene. This region also has bidirectional promoter activity, produced overlapping forward and reverse transcripts, and is necessary for both random and imprinted X inactivation in mice. The human sequence also has a 14 kb insertion not found in mouse Tsix.
- Links: OMIM - Tsix
Meiotic Sex Chromosome Inactivation
Both the X and Y chromosome are transcriptionally silenced during spermatogenesis, at primary spermatocyte stage onward, by a process called meiotic sex chromosome inactivation (MSCI). This second form of X inactivation takes place in the male, during spermatogenesis, as germ cells enter meiosis. This is thought to be a form of meiotic silencing of unsynapsed chromatin that silences chromosomes that fail to pair with their homologous partners.[4]
- Links: Meiosis | Y Chromosome
Placenta
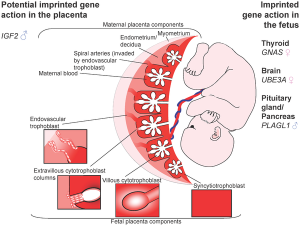
In the mouse placenta, X inactivation is imprinted and the paternal X chromosome is always inactive. In the human placenta, initial studies appear to show a pattern of random X inactivation.[12] They also identified a preferential expression of maternal alleles, indicating that, although imprinted X chromosome inactivation has been lost during evolution, a proliferative advantage may remain for cells that inactivate the paternal X (Xp) in human placenta.
- Links: Placenta Development | Mouse Development
Cancer
Recently meta-analysis of postnatal up-regulation of XIST has been shown to associate with poor prognosis in human cancers.[22][23]
References
- ↑ 1.0 1.1 Dvash T, Lavon N & Fan G. (2010). Variations of X chromosome inactivation occur in early passages of female human embryonic stem cells. PLoS ONE , 5, e11330. PMID: 20593031 DOI.
- ↑ McLaughlin CR & Chadwick BP. (2011). Characterization of DXZ4 conservation in primates implies important functional roles for CTCF binding, array expression and tandem repeat organization on the X chromosome. Genome Biol. , 12, R37. PMID: 21489251 DOI.
- ↑ 3.0 3.1 LYON MF. (1961). Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature , 190, 372-3. PMID: 13764598
- ↑ 4.0 4.1 Turner JM. (2007). Meiotic sex chromosome inactivation. Development , 134, 1823-31. PMID: 17329371 DOI.
- ↑ Colognori D, Sunwoo H, Wang D, Wang CY & Lee JT. (2020). Xist Repeats A and B Account for Two Distinct Phases of X Inactivation Establishment. Dev. Cell , , . PMID: 32531209 DOI.
- ↑ van Bemmel JG, Galupa R, Gard C, Servant N, Picard C, Davies J, Szempruch AJ, Zhan Y, Żylicz JJ, Nora EP, Lameiras S, de Wit E, Gentien D, Baulande S, Giorgetti L, Guttman M, Hughes JR, Higgs DR, Gribnau J & Heard E. (2019). The bipartite TAD organization of the X-inactivation center ensures opposing developmental regulation of Tsix and Xist. Nat. Genet. , 51, 1024-1034. PMID: 31133748 DOI.
- ↑ Żylicz JJ, Bousard A, Žumer K, Dossin F, Mohammad E, da Rocha ST, Schwalb B, Syx L, Dingli F, Loew D, Cramer P & Heard E. (2018). The Implication of Early Chromatin Changes in X Chromosome Inactivation. Cell , , . PMID: 30595450 DOI.
- ↑ Sangrithi MN & Turner JMA. (2018). Mammalian X Chromosome Dosage Compensation: Perspectives From the Germ Line. Bioessays , , . PMID: 29756331 DOI.
- ↑ Wu EX, Stanar P & Ma S. (2014). X-chromosome inactivation in female newborns conceived by assisted reproductive technologies. Fertil. Steril. , 101, 1718-23. PMID: 24726222 DOI.
- ↑ Gendrel AV & Heard E. (2011). Fifty years of X-inactivation research. Development , 138, 5049-55. PMID: 22069183 DOI.
- ↑ Naughton C, Sproul D, Hamilton C & Gilbert N. (2010). Analysis of active and inactive X chromosome architecture reveals the independent organization of 30 nm and large-scale chromatin structures. Mol. Cell , 40, 397-409. PMID: 21070966 DOI.
- ↑ 12.0 12.1 Moreira de Mello JC, de Araújo ES, Stabellini R, Fraga AM, de Souza JE, Sumita DR, Camargo AA & Pereira LV. (2010). Random X inactivation and extensive mosaicism in human placenta revealed by analysis of allele-specific gene expression along the X chromosome. PLoS ONE , 5, e10947. PMID: 20532033 DOI.
- ↑ Schoeftner S, Blanco R, Lopez de Silanes I, Muñoz P, Gómez-López G, Flores JM & Blasco MA. (2009). Telomere shortening relaxes X chromosome inactivation and forces global transcriptome alterations. Proc. Natl. Acad. Sci. U.S.A. , 106, 19393-8. PMID: 19887628 DOI.
- ↑ BARR ML & CARR DH. (1962). Correlations between sex chromatin and sex chromosomes. Acta Cytol. , 6, 34-45. PMID: 13865187
- ↑ Reinius B, Shi C, Hengshuo L, Sandhu KS, Radomska KJ, Rosen GD, Lu L, Kullander K, Williams RW & Jazin E. (2010). Female-biased expression of long non-coding RNAs in domains that escape X-inactivation in mouse. BMC Genomics , 11, 614. PMID: 21047393 DOI.
- ↑ Thorogood NP & Brown CJ. (2010). Active chromatin marks are retained on X chromosomes lacking gene or repeat silencing despite XIST/Xist expression in somatic cell hybrids. PLoS ONE , 5, e10787. PMID: 20520737 DOI.
- ↑ Chadwick BP & Willard HF. (2004). Multiple spatially distinct types of facultative heterochromatin on the human inactive X chromosome. Proc. Natl. Acad. Sci. U.S.A. , 101, 17450-5. PMID: 15574503 DOI.
- ↑ Brinkmann V & Zychlinsky A. (2012). Neutrophil extracellular traps: is immunity the second function of chromatin?. J. Cell Biol. , 198, 773-83. PMID: 22945932 DOI.
- ↑ BARR ML. (1956). The sex chromatin and its bearing on errors of sex development. Can Med Assoc J , 74, 419-22. PMID: 13304780
- ↑ Brown CJ, Ballabio A, Rupert JL, Lafreniere RG, Grompe M, Tonlorenzi R & Willard HF. (1991). A gene from the region of the human X inactivation centre is expressed exclusively from the inactive X chromosome. Nature , 349, 38-44. PMID: 1985261 DOI.
- ↑ Navarro P, Oldfield A, Legoupi J, Festuccia N, Dubois A, Attia M, Schoorlemmer J, Rougeulle C, Chambers I & Avner P. (2010). Molecular coupling of Tsix regulation and pluripotency. Nature , 468, 457-60. PMID: 21085182 DOI.
- ↑ Liu JL, Zhang WQ, Zhao M & Huang MY. (2019). Upregulation of long noncoding RNA XIST is associated with poor prognosis in human cancers. J. Cell. Physiol. , 234, 6594-6600. PMID: 30341910 DOI.
- ↑ Chen J, Yang X, Gong D, Cui Y, Hu J, Li H, Liu P, Li C, Cheng X, Liu L, Chen H & Zu X. (2019). Long noncoding RNA X-inactive specific transcript as a prognostic factor in cancer patients: A meta-analysis based on retrospective studies. Medicine (Baltimore) , 98, e14095. PMID: 30653128 DOI.
Reviews
Strehle M & Guttman M. (2020). Xist drives spatial compartmentalization of DNA and protein to orchestrate initiation and maintenance of X inactivation. Curr. Opin. Cell Biol. , 64, 139-147. PMID: 32535328 DOI.
Patrat C, Ouimette JF & Rougeulle C. (2020). X chromosome inactivation in human development. Development , 147, . PMID: 31900287 DOI.
Berletch JB, Yang F & Disteche CM. (2010). Escape from X inactivation in mice and humans. Genome Biol. , 11, 213. PMID: 20573260 DOI.
Agrelo R & Wutz A. (2010). X inactivation and disease. Semin. Cell Dev. Biol. , 21, 194-200. PMID: 19815084 DOI.
Bondy CA & Cheng C. (2009). Monosomy for the X chromosome. Chromosome Res. , 17, 649-58. PMID: 19802705 DOI.
Articles
Morey C & Avner P. (2011). The demoiselle of X-inactivation: 50 years old and as trendy and mesmerising as ever. PLoS Genet. , 7, e1002212. PMID: 21811421 DOI.
Dvash T, Lavon N & Fan G. (2010). Variations of X chromosome inactivation occur in early passages of female human embryonic stem cells. PLoS ONE , 5, e11330. PMID: 20593031 DOI.
Search Pubmed
July 2010 "X Inactivation" - All (3157) Review (519) Free Full Text (1066)
Search Pubmed Now: X Inactivation | Xist | Tsix | Meiotic Sex Chromosome Inactivation
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2026, Mayıs 21) Embryology Molecular Development - X Inactivation. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Molecular_Development_-_X_Inactivation
- © Dr Mark Hill 2026, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G