Molecular Development - X Inactivation: Difference between revisions
| Line 17: | Line 17: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* '''Analysis of active and inactive X chromosome architecture reveals the independent organization of 30 nm and large-scale chromatin structures'''<ref name="PMID21070966"><pubmed>21070966</pubmed></ref> "Using a genetic model, we present a high-resolution chromatin fiber analysis of transcriptionally active (Xa) and inactive (Xi) X chromosomes packaged into euchromatin and facultative heterochromatin. Our results show that gene promoters have an open chromatin structure that is enhanced upon transcriptional activation but the Xa and the Xi have similar overall 30 nm chromatin fiber structures." | |||
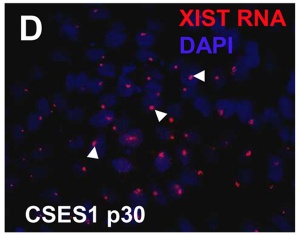
* '''Variations of X chromosome inactivation occur in early passages of female human embryonic stem cells.'''<ref name="PMID20593031" />"Human embryonic stem cells (hESCs) derived from inner cell mass (ICM) of blastocyst stage embryos have been used as a model system to understand XCI initiation and maintenance. Previous studies of undifferentiated female hESCs at intermediate passages have shown three possible states of XCI; 1) cells in a pre-XCI state, 2) cells that already exhibit XCI, or 3) cells that never undergo XCI even upon differentiation. In this study, XCI status was assayed in ten female hESC lines between passage 5 and 15 to determine whether XCI variations occur in early passages of hESCs. Our results show that three different states of XCI already exist in the early passages of hESC. " | * '''Variations of X chromosome inactivation occur in early passages of female human embryonic stem cells.'''<ref name="PMID20593031" />"Human embryonic stem cells (hESCs) derived from inner cell mass (ICM) of blastocyst stage embryos have been used as a model system to understand XCI initiation and maintenance. Previous studies of undifferentiated female hESCs at intermediate passages have shown three possible states of XCI; 1) cells in a pre-XCI state, 2) cells that already exhibit XCI, or 3) cells that never undergo XCI even upon differentiation. In this study, XCI status was assayed in ten female hESC lines between passage 5 and 15 to determine whether XCI variations occur in early passages of hESCs. Our results show that three different states of XCI already exist in the early passages of hESC. " | ||
Revision as of 00:50, 24 April 2011
Introduction
The presence in females of 2 X chromosome raises the issue of gene dosage, in the case of mammals this is regulated by inactivating one of the X chromosomes. To balance expression with the autosomal chromosomes the dosage imbalance is then adjusted by doubling expression of X-linked genes in both sexes.
In some other species compensation occurs by increasing the expression of X in males. The pattern of which X chromosome is inactivated in cells appears to be random, generating 50% cells expressing Father X, 50% cells expressing Mother X (mosaic pattern). The theory of random X inactivation was first suggested in mice in 1961.[2]
The process of inactivation relies on the Xist RNA, a 17 kb non-coding RNA, which accumulates on the future inactive X chromosome.
A second form of X inactivation that occurs only in male meiotic spermatogenesis, meiotic sex chromosome inactivation (MSCI), is not covered in these current notes. MSCI is the process of transcriptional silencing of the X and Y chromosomes.[3]
Some Recent Findings
|
X inactivation Xist
The physical region at the nucleus periphery where the inactive X chromosome is located in a female cell is described as the Barr body.[7]
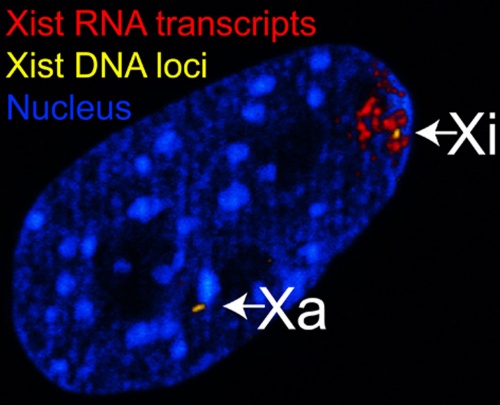
The above figure shows confocal images from a combined RNA-DNA FISH experiment for Xist in female mouse fibroblast cells (Female-biased expression of long non-coding RNAs in domains that escape X-inactivation in mouse.[8]
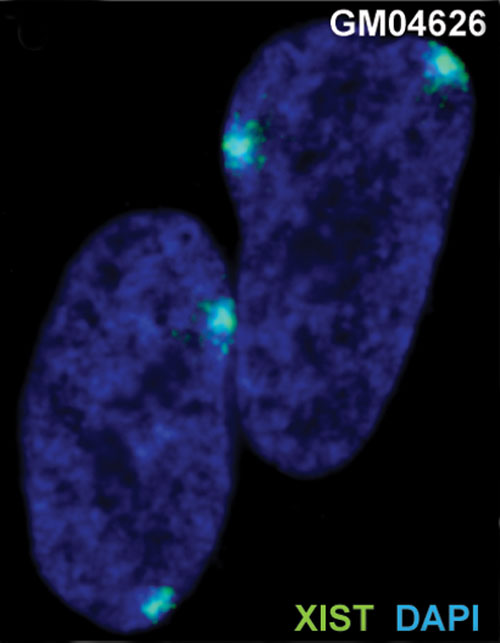
XIST expression in two cell nuclei using RNA FISH probes for the human XIST RNA (green) in the human female fibroblast line GM04626 (karyotype: 47,XXX).[9] As there are 3 copies of the X chromosome in each cell, there are 2 inactive X chromosomes as shown by Xist staining.
- Links: OMIM - Xist
X Inactivation (xist) History
The theory of random X inactivation was first suggested in mice in 1961.[2] The breakthrough was the discovery of the X inactive specific transcript (XIST). [10] This gene is located within the "X inactivation centre" and only expressed by the inactive X chromosome. unlike other genes that encode protein XIST contained no "open reading frames" (ie no codons to encode amino acids). XIST is transcribed but not translated. XIST appears to act as RNA. Current thinking is that it binds to the X Chromosome and is involved in it's translocation to the nuclear periphery. It now appears that XIST appears to initiate X inactivation and it is the methylation of the inactive X genes that maintains inactivity.
Tsix
A gene antisense to Xist, hence the name which is Xist backwards, in embryonic stem cells Xist repression occurs and Tsix is upregulated. This is part of the overall process of maintaining pluripotency in these cells. Tsix upregulation depends on the recruitment of another pluripotent marker Rex1, and of the associated factors Klf4 and c-Myc, by the DXPas34 minisatellite associated with the Tsix promoter.[11]
DXPas34 is made up of 34bp repeats containing Ctcf motifs and is located within a CpG-rich region in the 5-prime end of the mouse Tsix gene. This region also has bidirectional promoter activity, produced overlapping forward and reverse transcripts, and is necessary for both random and imprinted X inactivation in mice. The human sequence also has a 14 kb insertion not found in mouse Tsix.
- Links: OMIM - Tsix
Meiotic Sex Chromosome Inactivation
This second form of X inactivation takes place in the male, during spermatogenesis, as germ cells enter meiosis. This is thought to be a form of meiotic silencing of unsynapsed chromatin that silences chromosomes that fail to pair with their homologous partners.[12]
Placenta
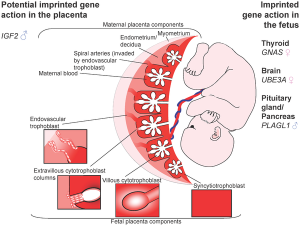
In the mouse placenta, X inactivation is imprinted and the paternal X chromosome is always inactive. In the human placenta, initial studies appear to show a pattern of random X inactivation.[13] They also identified a preferential expression of maternal alleles, indicating that, although imprinted X chromosome inactivation has been lost during evolution, a proliferative advantage may remain for cells that inactivate the paternal X (Xp) in human placenta.
References
- ↑ 1.0 1.1 <pubmed>20593031</pubmed>
- ↑ 2.0 2.1 <pubmed>13764598</pubmed>
- ↑ <pubmed>17329371</pubmed>
- ↑ <pubmed>21070966</pubmed>
- ↑ <pubmed>20532033</pubmed>
- ↑ <pubmed>19887628</pubmed>
- ↑ <pubmed>13865187</pubmed>
- ↑ <pubmed>21047393</pubmed>| BMC Genomics
- ↑ <pubmed>20520737</pubmed>| PLoS One
- ↑ <pubmed>1985261</pubmed>
- ↑ <pubmed>21085182</pubmed>
- ↑ <pubmed>17329371</pubmed>
- ↑ <pubmed>20532033</pubmed>| PLoS One.
Search Pubmed
July 2010 "X Inactivation" - All (3157) Review (519) Free Full Text (1066)
Search Pubmed Now: X Inactivation | Xist | Tsix
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, June 21) Embryology Molecular Development - X Inactivation. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Molecular_Development_-_X_Inactivation
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G