Morula: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 23: | Line 23: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* D'''iversity of human and mouse homeobox gene expression in development and adult tissues'''<ref name="PMID27809766"><pubmed>27809766</pubmed></ref> "8-cell to morula, 12 eutherian-specific homeobox genes not detectable outside of reproductive tissues or the embryo (RHOXF2, RHOXF2B, CPHX1, CPHX2, DPRX, LEUTX, TPRX1, TPRX2, ARGFX, NANOGNB, DUXA, DUXB). [[Developmental Signals - Homeobox|Hox]] | |||
* '''Par-aPKC-dependent and -independent mechanisms cooperatively control cell polarity, Hippo signaling, and cell positioning in 16-cell stage mouse embryos'''<ref name="PMID26450797"><pubmed>26450797</pubmed></ref> "In preimplantation mouse embryos, the Hippo signaling pathway plays a central role in regulating the fates of the trophectoderm (TE) and the inner cell mass (ICM). In early blastocysts with more than 32 cells, the Par-aPKC system controls polarization of the outer cells along the apicobasal axis, and cell polarity suppresses Hippo signaling. Inactivation of Hippo signaling promotes nuclear accumulation of a coactivator protein, Yap, leading to induction of TE-specific genes. However, whether similar mechanisms operate at earlier stages is not known. Here, we show that slightly different mechanisms operate in 16-cell stage embryos. ...These results suggest that cell polarization at the 16-cell stage is regulated by both Par-aPKC-dependent and -independent mechanisms. Asymmetric cell division is involved in cell polarity control, and cell polarity regulates cell positioning and most likely controls Hippo signaling." [[Developmental Signals - Hippo|Hippo]] | [[Mouse Development]] | * '''Par-aPKC-dependent and -independent mechanisms cooperatively control cell polarity, Hippo signaling, and cell positioning in 16-cell stage mouse embryos'''<ref name="PMID26450797"><pubmed>26450797</pubmed></ref> "In preimplantation mouse embryos, the Hippo signaling pathway plays a central role in regulating the fates of the trophectoderm (TE) and the inner cell mass (ICM). In early blastocysts with more than 32 cells, the Par-aPKC system controls polarization of the outer cells along the apicobasal axis, and cell polarity suppresses Hippo signaling. Inactivation of Hippo signaling promotes nuclear accumulation of a coactivator protein, Yap, leading to induction of TE-specific genes. However, whether similar mechanisms operate at earlier stages is not known. Here, we show that slightly different mechanisms operate in 16-cell stage embryos. ...These results suggest that cell polarization at the 16-cell stage is regulated by both Par-aPKC-dependent and -independent mechanisms. Asymmetric cell division is involved in cell polarity control, and cell polarity regulates cell positioning and most likely controls Hippo signaling." [[Developmental Signals - Hippo|Hippo]] | [[Mouse Development]] | ||
* '''Functional genomics of 5- to 8-cell stage human embryos by blastomere single-cell cDNA analysis'''<ref name="PMID21049019">Galán A, Montaner D, Póo ME, Valbuena D, Ruiz V, Aguilar C, Dopazo J, Simón C. '''Functional genomics of 5- to 8-cell stage human embryos by blastomere single-cell cDNA analysis.''' PLoS One. 2010 Oct 26;5(10):e13615. [http://www.ncbi.nlm.nih.gov/pubmed/21049019 PMID21049019] | [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0013615 PLoS One.]</ref> "Forty-nine blastomeres from 5- to 8-cell human embryos have been investigated following an efficient single-cell cDNA amplification protocol to provide a template for high-density microarray analysis. The previously described markers, characteristic of Inner Cell Mass (ICM) (n = 120), stemness (n = 190) and Trophectoderm (TE) (n = 45), were analyzed, and a housekeeping pattern of 46 genes was established. ...In summary, the global single-cell cDNA amplification microarray analysis of the 5- to 8-cell stage human embryos reveals that blastomere fate is not committed to ICM or TE." | * '''Functional genomics of 5- to 8-cell stage human embryos by blastomere single-cell cDNA analysis'''<ref name="PMID21049019">Galán A, Montaner D, Póo ME, Valbuena D, Ruiz V, Aguilar C, Dopazo J, Simón C. '''Functional genomics of 5- to 8-cell stage human embryos by blastomere single-cell cDNA analysis.''' PLoS One. 2010 Oct 26;5(10):e13615. [http://www.ncbi.nlm.nih.gov/pubmed/21049019 PMID21049019] | [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0013615 PLoS One.]</ref> "Forty-nine blastomeres from 5- to 8-cell human embryos have been investigated following an efficient single-cell cDNA amplification protocol to provide a template for high-density microarray analysis. The previously described markers, characteristic of Inner Cell Mass (ICM) (n = 120), stemness (n = 190) and Trophectoderm (TE) (n = 45), were analyzed, and a housekeeping pattern of 46 genes was established. ...In summary, the global single-cell cDNA amplification microarray analysis of the 5- to 8-cell stage human embryos reveals that blastomere fate is not committed to ICM or TE." | ||
Revision as of 11:44, 9 August 2017
| Embryology - 24 Jun 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
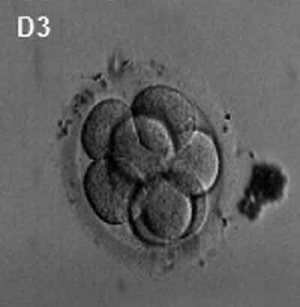
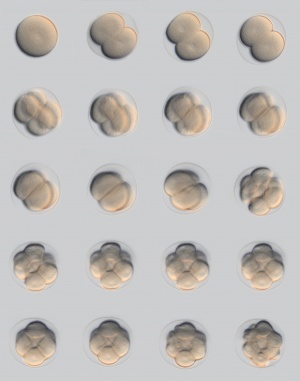
(Latin, morula = mulberry) An early stage in post-fertilization development when cells have rapidly mitotically divided to produce a solid mass of cells (12-15 cells) with a "mulberry" appearance. This stage is followed by formation of a cavity in this cellular mass blastocyst stage.
A key event prior to morula formation is "compaction", where the 8 cell embryo undergoes changes in cell morphology and cell-cell adhesion that initiates the formation of this solid ball of cells.
In humans, morula stage of development occurs during the first days in the first week following fertilization and is described as Carnegie stage 2. This stage is followed by formation of a cavity, the blastocoel, which defines formation of the blastocyst.
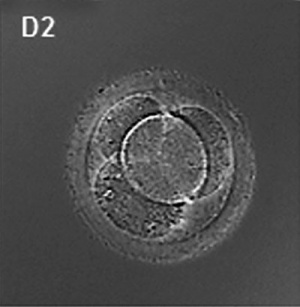
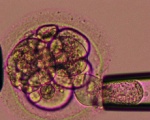 ART Preimplantation blastomere biopsy[2]
|
In Assisted Reproductive Technology, the morula stage is when one of the earliest prenatal diagnostic test can be carried out, by removing a single cell (blastomere) and carrying out genetic diagnosis on its DNA. |
- Links: Carnegie stage 2 | Morula | Mitosis | Blastocyst | Fertilization | Week 1 | Category:Carnegie Stage 2 | Category:Morula
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Morula Development <pubmed limit=5>Morula Development</pubmed> |
Movies
|
|
|
|
|
Compaction
- E-cadherin mediated adhesion initiates at compaction at the 8-cell stage
- regulated post-translationally via protein kinase C and other signalling molecules
Blastomere Division
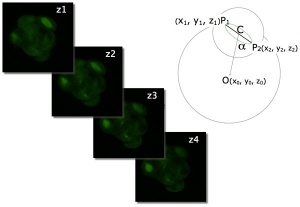
An in vitro study of human blastocyst development[8] showed that those blastomeres that initially divide quickly are more likely to develop to blastocyst stage.
A recent study in mice showed that there was no specific orientation of the mitotic spindle during cell division in the 8 to 16 cell stage transition.[7] This suggests no predetermined cleavage pattern (pre-patterned) at the 8 cell stage and only modulated by the extent of cell rounding up during mitosis. In other species, such as the worm C.elegans and ascidians, have specific patterns of spindle orientation from the zygote stage.
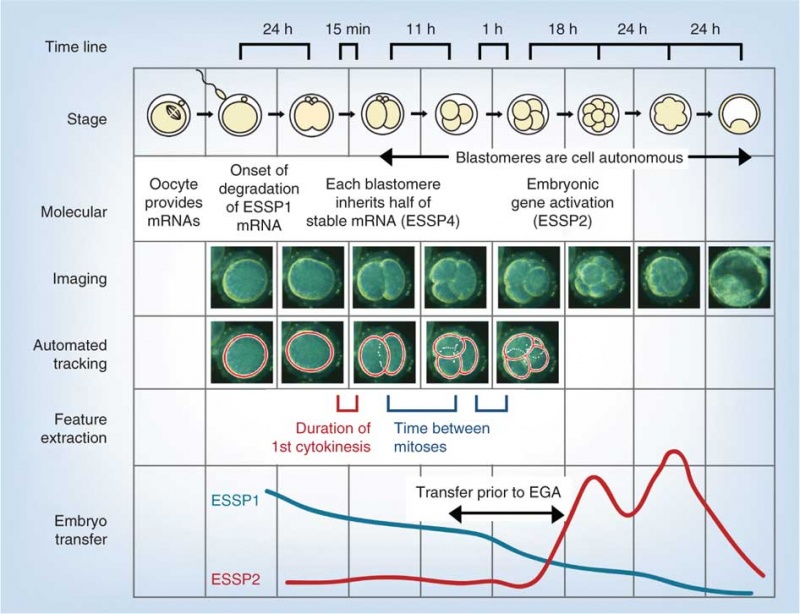
Model Human Morula Development
The following figure is from a recent study[6] using video and genetic analysis of in vitro human development during week 1 following fertilization.
- EGA - embryonic genome activation
- ESSP - embryonic stage–specific pattern, four unique embryonic stage–specific patterns (1-4)
- Links: Figure with legend
Morulas in Other Species
Mouse Morula
- Links: Mouse Development
Sea Urchin Morula
Sea Urchin early embryo cleavage pattern (SDB Gallery Images)
- Links: Sea Urchin Development
Bovine Morula
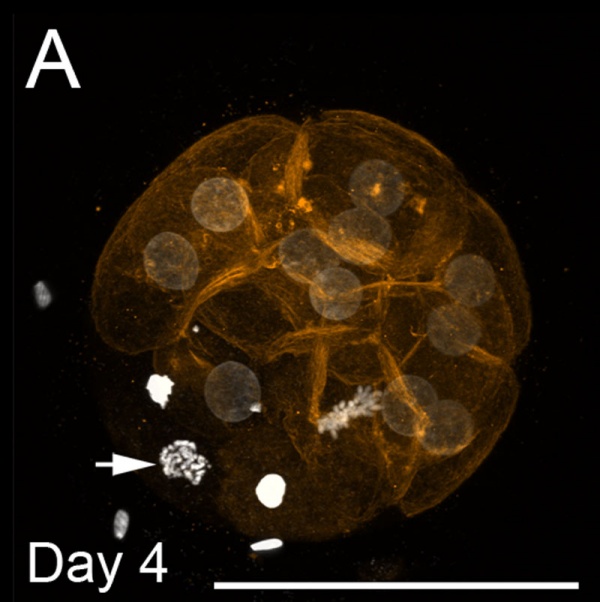
Bovine Morula[9]
- Image shows DNA staining (white) and f-actin filaments (orange) at day 4. Scale bars represent 100 µm.
- Pale staining round nuclei are at interphase.
- Arrow shows single nucleus at prophase.
- A single nucleus is seen at metaphase.
- Condensed bright nuclei are apoptotic.
- Links: Bovine Development | Mitosis
References
- ↑ 1.0 1.1 <pubmed>19924284</pubmed>| PMC2773928 | PLoS One
- ↑ <pubmed>24783200</pubmed>| PMC3982254 | Biomed Res Int.
- ↑ <pubmed>27809766</pubmed>
- ↑ <pubmed>26450797</pubmed>
- ↑ Galán A, Montaner D, Póo ME, Valbuena D, Ruiz V, Aguilar C, Dopazo J, Simón C. Functional genomics of 5- to 8-cell stage human embryos by blastomere single-cell cDNA analysis. PLoS One. 2010 Oct 26;5(10):e13615. PMID21049019 | PLoS One.
- ↑ 6.0 6.1 <pubmed>20890283</pubmed>| Nat Biotechnol.
- ↑ 7.0 7.1 <pubmed>19997595</pubmed>| PLoS One.
- ↑ <pubmed>11821286</pubmed>
- ↑ <pubmed>21811561</pubmed>| PLoS One.
Articles
<pubmed>26416966</pubmed> <pubmed>19289087</pubmed> <pubmed>20157423</pubmed>
Search PubMed
Search Pubmed: morula development | blastomere development |
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, June 24) Embryology Morula. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Morula
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G