Hutchinson-Gilford Progeria Syndrome
| Embryology - 18 Apr 2026 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
| Educational Use Only - Embryology is an educational resource for learning concepts in embryological development, no clinical information is provided and content should not be used for any other purpose. |
Introduction
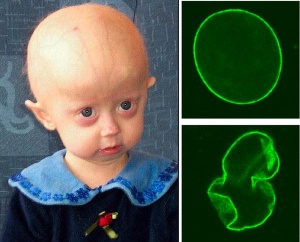
Extremely rare genetic abnormality resulting in a "premature ageing" condition and a range of systemic abnormalities. Lamins A and C are the alternatively spliced products of the LMNA gene and form a key component of the nuclear cytoskeleton, the nuclear lamina.
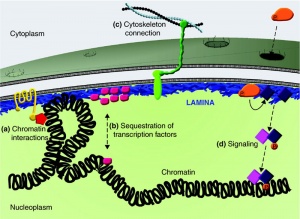
The nuclear cytoskeleton has 2 layers
- outer - less organised surrounds membrane
- inner - nuclear lamina - thin shell (20 nm) underlying the membrane (nuclear envelope)
- associates with both the inner nuclear membrane and underlying chromatin
- can regulate gene expression
- provides anchor sites for nuclear pore complexes
- broken down each cell division
Nuclear Lamins
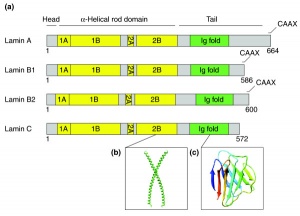
- intermediate filaments
- large family of different filament types
- 10 nm in diameter, forms rope-like networks
- lamins, Class V intermediate filaments
- polypeptide form dimers
- central alpha-helical regions of two polypeptide chains are wound around each other
- assembly
- head-to-tail association of dimers form linear polymers
- side-by-side association of polymers form filaments
- B-type lamins are ubiquitously expressed throughout development.
- A-type lamins in many organisms expression does not appear until midway through embryogenesis (possible role in differentiation)
- Lamin Abnormalities - laminopathies
Some Recent Findings
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Hutchinson-Gilford Progeria Syndrome <pubmed limit=5>Hutchinson-Gilford Progeria Syndrome</pubmed> |
Neonatal progeria: increased ratio of progerin to lamin A leads to progeria of the newborn
Eur J Hum Genet. 2012 Mar 14. doi: 10.1038/ejhg.2012.36. [Epub ahead of print]
Reunert J, Wentzell R, Walter M, Jakubiczka S, Zenker M, Brune T, Rust S, Marquardt T. Source Universitätsklinikum Münster, Klinik für Kinder- und Jugendmedizin - Allgemeine Pädiatrie, Münster, Germany.
Abstract
Hutchinson-Gilford progeria syndrome (HGPS) is an important model disease for premature ageing. Affected children appear healthy at birth, but develop the first symptoms during their first year of life. They die at an average age of 13 years, mostly because of myocardial infarction or stroke. Classical progeria is caused by the heterozygous point mutation c.1824C>T in the LMNA gene, which activates a cryptic splice site. The affected protein cannot be processed correctly to mature lamin A, but is modified into a farnesylated protein truncated by 50 amino acids (progerin). Three more variations in LMNA result in the same mutant protein, but different grades of disease severity. We describe a patient with the heterozygous LMNA mutation c.1821G>A, leading to neonatal progeria with death in the first year of life. Intracellular lamin A was downregulated in the patient's fibroblasts and the ratio of progerin to lamin A was increased when compared with HGPS. It is suggestive that the ratio of farnesylated protein to mature lamin A determines the disease severity in progeria.European Journal of Human Genetics advance online publication, 14 March 2012; doi:10.1038/ejhg.2012.36.
PMID 22419169
Requirements for efficient proteolytic cleavage of prelamin A by ZMPSTE24
PLoS One. 2012;7(2):e32120. Epub 2012 Feb 15.
Barrowman J, Hamblet C, Kane MS, Michaelis S. Source Department of Cell Biology, The Johns Hopkins School of Medicine, Baltimore, Maryland, United States of America.
Abstract
BACKGROUND: The proteolytic maturation of the nuclear protein lamin A by the zinc metalloprotease ZMPSTE24 is critical for human health. The lamin A precursor, prelamin A, undergoes a multi-step maturation process that includes CAAX processing (farnesylation, proteolysis and carboxylmethylation of the C-terminal CAAX motif), followed by ZMPSTE24-mediated cleavage of the last 15 amino acids, including the modified C-terminus. Failure to cleave the prelamin A "tail", due to mutations in either prelamin A or ZMPSTE24, results in a permanently prenylated form of prelamin A that underlies the premature aging disease Hutchinson-Gilford Progeria Syndrome (HGPS) and related progeroid disorders. METHODOLOGY/PRINCIPAL FINDINGS: Here we have investigated the features of the prelamin A substrate that are required for efficient cleavage by ZMPSTE24. We find that the C-terminal 41 amino acids of prelamin A contain sufficient context to allow cleavage of the tail by ZMPSTE24. We have identified several mutations in amino acids immediately surrounding the cleavage site (between Y646 and L647) that interfere with efficient cleavage of the prelamin A tail; these mutations include R644C, L648A and N650A, in addition to the previously reported L647R. Our data suggests that 9 of the 15 residues within the cleaved tail that lie immediately upstream of the CAAX motif are not critical for ZMPSTE24-mediated cleavage, as they can be replaced by the 9 amino acid HA epitope. However, duplication of the same 9 amino acids (to increase the distance between the prenyl group and the cleavage site) impairs the ability of ZMPSTE24 to cleave prelamin A. CONCLUSIONS/SIGNIFICANCE: Our data reveals amino acid preferences flanking the ZMPSTE24 cleavage site of prelamin A and suggests that spacing from the farnesyl-cysteine to the cleavage site is important for optimal ZMPSTE24 cleavage. These studies begin to elucidate the substrate requirements of an enzyme activity critical to human health and longevity.
PMID 22355414
Regulation of prelamin A but not lamin C by miR-9, a brain-specific microRNA
Proc Natl Acad Sci U S A. 2012 Feb 14;109(7):E423-31. Epub 2012 Jan 30.
Jung HJ, Coffinier C, Choe Y, Beigneux AP, Davies BS, Yang SH, Barnes RH 2nd, Hong J, Sun T, Pleasure SJ, Young SG, Fong LG. Source Molecular Biology Institute and Department of Medicine and Human Genetics, University of California, Los Angeles, CA 90095, USA.
Abstract
Lamins A and C, alternatively spliced products of the LMNA gene, are key components of the nuclear lamina. The two isoforms are found in similar amounts in most tissues, but we observed an unexpected pattern of expression in the brain. Western blot and immunohistochemistry studies showed that lamin C is abundant in the mouse brain, whereas lamin A and its precursor prelamin A are restricted to endothelial cells and meningeal cells and are absent in neurons and glia. Prelamin A transcript levels were low in the brain, but this finding could not be explained by alternative splicing. In lamin A-only knockin mice, where alternative splicing is absent and all the output of the gene is channeled into prelamin A transcripts, large amounts of lamin A were found in peripheral tissues, but there was very little lamin A in the brain. Also, in knockin mice expressing exclusively progerin (a toxic form of prelamin A found in Hutchinson-Gilford progeria syndrome), the levels of progerin in the brain were extremely low. Further studies showed that prelamin A expression, but not lamin C expression, is down-regulated by a brain-specific microRNA, miR-9. Expression of miR-9 in cultured cells reduced lamin A expression, and this effect was abolished when the miR-9-binding site in the prelamin A 3' UTR was mutated. The down-regulation of prelamin A expression in the brain could explain why mouse models of Hutchinson-Gilford progeria syndrome are free of central nervous system pathology.
PMID 22308344
References
- ↑ <pubmed>16277559</pubmed>
- ↑ 2.0 2.1 <pubmed>21639948</pubmed>| PMC3219962 | Genome Biology
- ↑ <pubmed>22419169</pubmed>
- ↑ <pubmed>22355414</pubmed>
Reviews
<pubmed>22156746</pubmed> <pubmed>21639948</pubmed>| Genome Biology
Articles
<pubmed></pubmed>
Search PubMed
Search Pubmed: Hutchinson-Gilford Progeria Syndrome | Progeria
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- PubMed Health Progeria
- Progeria Research Foundation
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2026, April 18) Embryology Hutchinson-Gilford Progeria Syndrome. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Hutchinson-Gilford_Progeria_Syndrome
- © Dr Mark Hill 2026, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
