File:VIT Gene Evolution.jpg
VIT_Gene_Evolution.jpg (668 × 405 pixels, file size: 76 KB, MIME type: image/jpeg)
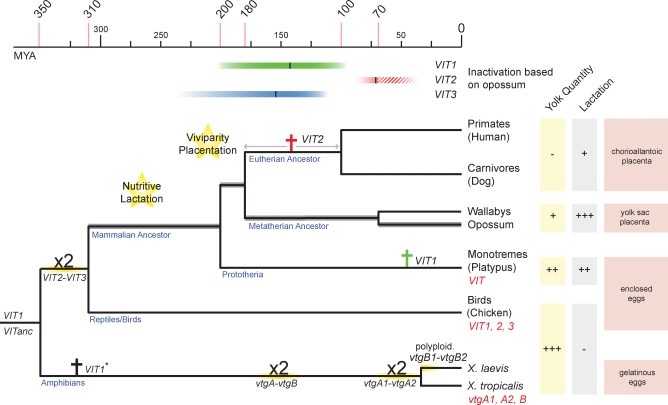
VIT Gene Evolution in Tetrapods
The topology and divergence times of the tree are based on previous studies [19,24,25,41]. Latin crosses indicate VIT inactivation events in eutherians and monotremes. Inactivation estimates (including approximated 95% prediction intervals) based on opossum VIT sequences are indicated by colored bars at the top (see also Figure 3). Duplications (“x2”) are indicated. VITanc is the likely ancestor of both the amphibian vtgA1/vtgA2 and VIT2/VIT3 genes in birds. Functional VIT genes in extant species are indicated in red. The inactivation time of VIT1* on the amphibian branch could not be estimated because of its absence in Xenopus tropicalis.
Pbio.0060063.g001.jpg
Loss of Egg Yolk Genes in Mammals and the Origin of Lactation and Placentation David Brawand, Walter Wahli, and Henrik Kaessmann PLoS Biol. 2008 March; 6(3): e63. Published online 2008 March 18. doi: 10.1371/journal.pbio.0060063. PMCID: PMC2267819
PLoS Biol. 2008 March; 6(3): e63. Published online 2008 March 18. doi: 10.1371/journal.pbio.0060063.
Copyright : © 2008 Brawand et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 09:14, 16 August 2009 |  | 668 × 405 (76 KB) | S8600021 (talk | contribs) | VIT Gene Evolution in Tetrapods The topology and divergence times of the tree are based on previous studies [19,24,25,41]. Latin crosses indicate VIT inactivation events in eutherians and monotremes. Inactivation estimates (including approximated 95% pre |
You cannot overwrite this file.
File usage
There are no pages that use this file.