File:Mitosis peroxisomes 01.jpg
Original file (1,200 × 394 pixels, file size: 112 KB, MIME type: image/jpeg)
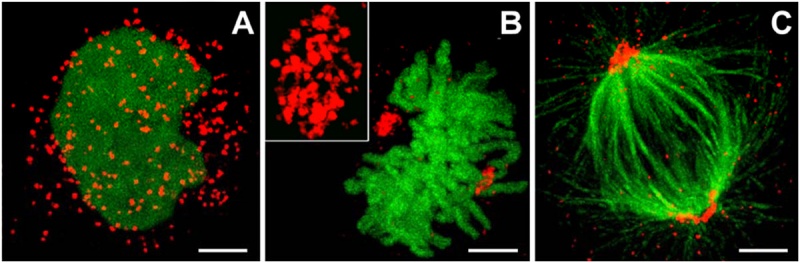
Peroxisome location at Interphase and during Mitosis
Peroxisome clusters (red) in Human Embryonic Kidney 293 (HEK293) cells during:
(A) interphase.
(B) metaphase. The inset in (B) shows a magnification of a peroxisomal cluster.
(C) Tubulin fibers are labelled in green with Alexa Fluor©-488 via an anti-α-tubulin antibody to reveal peroxisomal clustering at the spindle poles. Bars: 2 µm.
Confocal images (Leica TCS 4Pi) of HEK293 cells co-expressing mRuby-PTS and H2B-EGFP
- Links: Image - Interphase and Mitosis | Image - Interphase | Image - Mitosis | Image - Mitosis | Mitosis | ATCC HEK293
Reference
<pubmed>19194514</pubmed>| PMC2633614 | PLoS One.
Citation: Kredel S, Oswald F, Nienhaus K, Deuschle K, Röcker C, et al. (2009) mRuby, a Bright Monomeric Red Fluorescent Protein for Labeling of Subcellular Structures. PLoS ONE 4(2): e4391. doi:10.1371/journal.pone.0004391
Editor: Amy S. Gladfelter, Dartmouth College, United States of America
Received: July 9, 2008; Accepted: December 22, 2008; Published: February 5, 2009
Copyright: © 2009 Kredel et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Journal.pone.0004391.g004.jpg
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 18:44, 9 September 2012 | 1,200 × 394 (112 KB) | Z8600021 (talk | contribs) | ==Peroxisome location at Interphase and Mitosis== Peroxisome clusters in HEK293 cells. (A–C) Confocal images (Leica TCS 4Pi) of HEK293 cells co-expressing mRuby-PTS and H2B-EGFP during (A) interphase (B) metaphase. The inset in (B) shows a magnif |
You cannot overwrite this file.
File usage
The following page uses this file: