2016 Group Project 3: Difference between revisions
m (→History) |
|||
| Line 52: | Line 52: | ||
[[File:FGFR receptor subtype.jpeg|thumb|none|300px|Simplistic illustration of the FGFR receptors adapted from review article [http://www.ncbi.nlm.nih.gov/pubmed/16216232 Functions and regulations of fibroblast growth factor signaling during embryonic development]]] | [[File:FGFR receptor subtype.jpeg|thumb|none|300px|Simplistic illustration of the FGFR receptors adapted from review article [http://www.ncbi.nlm.nih.gov/pubmed/16216232 Functions and regulations of fibroblast growth factor signaling during embryonic development]]] | ||
===Subtypes of FGFR=== | |||
{| class="pretty table" | |||
|-bgcolor="DDCEF2" | |||
| '''FGFR Subtype''' || '''Function''' || '''Abnormalities''' | |||
|-bgcolor="FAF5FF" | |||
| FGFR1 || | |||
*Involved in morphogenesis as well as orchestrating the patterning of the mesodermal germ layer at gastrulation<ref><pubmed>16207751</pubmed></ref> <br> | |||
*Involved in formation of the organ of corti and auditory sensory epithelium <ref><pubmed>12194867</pubmed></ref> <br> | |||
*Expressed in early limb bud <ref><pubmed>1321062</pubmed></ref> <br><br> | |||
*Expressed at epiphyseal growth plate as well as in the perichondrium, prehypertrophic and hypertrophic chondrocytes<ref><pubmed>17169623 </pubmed></ref> <br> | |||
*Is a negative regulator of bone growth<ref><pubmed>16815385</pubmed></ref> <br> | |||
|| | |||
*Pfeiffer Syndrome (Type 1) | |||
*Kallmann syndrome | |||
*Osteoglophonic dysplasia | |||
*8p11 myeloproliferative syndrome | |||
|- | |||
| FGFR2 || | |||
*Activated prior to gastrulation with the purpose of repressing cellular movements in the presumptive anterior neural plate and preventing normal retinal progenitor cells from adopting retinal fates<ref><pubmed>14723847</pubmed></ref> <br> | |||
*Acts as a marker of prechondrogenic condensations<ref><pubmed>9784595</pubmed></ref> <br> | |||
*Expressed in condensing mesenchyme of the early limb bud<ref><pubmed>1315677</pubmed></ref> <br> | |||
*Plays a key role in skeleton development as it is expressed in osteoprogenitor cells and differentiating osteoblasts<ref><pubmed>20489451 </pubmed></ref> <br> | |||
*Is involved in cranial cell replication or differentiation in both humans and mice.<ref><pubmed>15863034 </pubmed></ref> <br> | |||
|| | |||
*Pfeiffer Syndrome (Type 1-3) | |||
*Apert Syndrome | |||
*Crouzon Syndrome | |||
*Beare-Stevenson cutis gryata syndrome<ref><pubmed>17552943 </pubmed></ref> <br> | |||
|-bgcolor="FAF5FF" | |||
| FGFR3 || | |||
*Induces complete growth arrest of cells<ref><pubmed>11779141 </pubmed></ref> <br> | |||
*Is required to promote differentiation of prechondrogenic mesenchymal cells to cartilage-producing chondrocytes <ref><pubmed>8432397 </pubmed></ref> <br> | |||
*Is expressed in chondrocytes, differentiated initially from the core of the mesenchyme condensation<ref><pubmed>8630492 </pubmed></ref> <br> | |||
*Is expressed in reserve and proliferating chondrocytes as the epiphyseal growth plate is formed<ref name="PMID 12080084"><pubmed>12080084</pubmed></ref> <br> | |||
|| | |||
*Achondroplasia (can be severe, with developmental delay and acanthuses) | |||
*Thanatophoric Dysplasia | |||
*Hypochondroplasia | |||
|- | |||
| FGFR4 || | |||
*Involved in proliferation of the blastocyst inner cell mass, differentiation of the presomitic mesoderm and limb bud development<ref name ="PMID 10662638"><pubmed>10662638</pubmed></ref> <br> | |||
*Regulates cholesterol metabolism, bile acid synthesis and liver mineral homeostasis | |||
*It will provide mitogenic and morphogenic signals to regulate normal limb development<ref name="PMID 12080084"/><br> | |||
*Promotes intramembranous ossification and participates in the development of calvarial bone<ref name ="PMID 10662638"/> <br> | |||
|| | |||
*Chondrodysplasia | |||
|- | |||
|} | |||
==References== | ==References== | ||
<references/> | <references/> | ||
Revision as of 16:22, 15 November 2016
| 2016 Student Projects | ||||
|---|---|---|---|---|
| Signalling: 1 Wnt | 2 Notch | 3 FGF Receptor | 4 Hedgehog | 5 T-box | 6 TGF-Beta | ||||
| 2016 Group Project Topic - Signaling in Development
OK you are now in a group, add a topic with your student signature to the group page. | ||||
| This page is an undergraduate science embryology student project and may contain inaccuracies in either descriptions or acknowledgements. | ||||
| Group Assessment Criteria |
|---|
 Science Student Projects Science Student Projects
|
| More Information on Assessment Criteria | Science Student Projects |
Fibroblast Growth Factor Receptor (FGFR) Pathway
Introduction
The Fibroblast Growth Factor (FGF) signalling pathway is critical for regulating progenitor cell proliferation, differentiation, survival and patterning. It is involved in the regulation and development of the early embryo, and is considered to be critical for normal vascular, skeletal and organ development. Furthermore, this pathway is involved in maintaining adult tissues through the regulation of metabolic functions and tissue repair (which is often through the reactivation of the same signalling pathways involved in early development.) [1]
This page will outline the FGFR signalling pathway, the history of scientific discoveries relevant to this pathway, receptor sub-types and a description of signal transduction. It will also describe its various roles in embryonic development including its influence on the patterning of the embryonic axis, as well as limb bud, bone, kidney, external genitalia and inner ear development. There is also a discussion of relevant animals models, such as those of the chick embryo, as well as abnormalities in this pathway relevant to embryonic development, including Achondroplasia, Pfeiffer syndrome and Apert syndrome. A short informative quiz accompanied with feedback is offered for readers to determine how much they have learnt from the information provided. A glossary at the bottom of the page explains specific terms mentioned throughout, along with links to relevant information from UNSW embryology lectures.
History
Ranging from its discovery in 1939 till the present, much has been learned about the nature of Fibroblast growth factor (FGF) in embryonic development. Researchers had noticed the growth stimulating effects that these isolated factors had, in that they induced fibroblast proliferation. Due to their ability to stimulate fibroblast proliferation they were termed "FGFs". Today, a variety of subtypes of FGFs have been discovered and categorised into a large family that exist in organisms including humans as well as nematodes. In addition, it was soon discovered that not all FGFs can stimulate fibroblasts.
The table below outlines some of the significant scientific discoveries regarding the FGFR signalling pathway over the years, as outlined in a review article. [2]
| Year | Scientific Discovery Regarding FGF/FGFR Signalling |
| 1939 | The first paper on FGFs was published through experiments that measured the mitogenic activity of saline extracts of different tissues from the chick. Early work also investigated the idea that uncontrolled proliferation is a hallmark of cancers and the involvement of growth factors such as FGF. |
| 1974 | FGF growth factor activity was shown to stimulate the growth of a fibroblast cell line in partially purified extracts from bovine pituitary. This lead to the term "fibroblast growth factor" to be derived. |
| 1987 | The interaction with heparin that FGFs have was translated into work regarding the interaction of FGFs with the glycosaminoglycan heparan sulfate within the pericellular and extracellular matrix. |
| 1989 | FGF1 and FGF2 were isolated from brain tissue. |
| 1990 | FGFR tyrosine kinases were identified for the first time |
| 1991 | FGFs were also shown to display growth factor activities on fibroblasts. In addition, the dependence of the growth factor activity of FGFs on heparan sulfate was discovered. |
| 2005 | A further set of FGF proteins termed the FGF homology factors were found to be wholly intracellular such that they do not interact with any of the extracellular receptors and partners of FGFs. |
| 2013 | A small group of FGFs were found to not bind heparan sulfate, but instead to interact with a protein co-receptor named Klotho. |
Overview Of The FGFR Pathway
23 protein families have been identified from the FGF signalling pathway, 18 of which are secreted signalling proteins (FGF1-10, and FGF16-23) that interact with 4 tyrosine kinase FGF Receptors (FGFR1-4), whilst 4 are intracellular non-signalling proteins (iFGFs; FGF11-14). [1]
As illustrated in the image below, FGFRs are comprised of 3 immunoglobulin domains (IgI, IgII, IgIII), with IgIII being the closest to the transmembrane and IgI being the furthest away. Some notable features of this receptor include an acidic box (AD) located in-between IgI and IgII, a heparin-binding domain (HBD) within IgII which is important in signal transduction, and the transmembrane (TM) structure of IgIII which has both kinase and interkinase domains (KD and IKD) within the intracellular space. FGF ligands linked to heparin sulfate proteoglycan (HSPG) bind to both the IgII and IgIII domain of the receptor (with the heparin component specifically binding to IgII) resulting in dimerisation of the receptors and activation of signal transduction pathways through the phosphorylation of tyrosine residues, as discussed in more detail under the subheading signal transduction. [3]
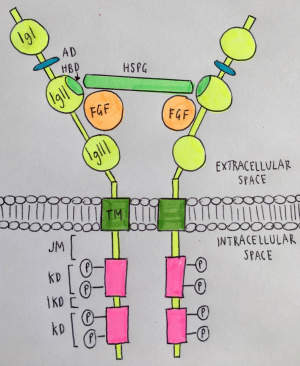
Subtypes of FGFR
| FGFR Subtype | Function | Abnormalities |
| FGFR1 |
|
|
| FGFR2 |
|
|
| FGFR3 |
|
|
| FGFR4 |
|
|
References
- ↑ 1.0 1.1 <pubmed>25772309</pubmed>
- ↑ <pubmed>26793421</pubmed>
- ↑ <pubmed>16216232</pubmed>[1]
- ↑ <pubmed>16207751</pubmed>
- ↑ <pubmed>12194867</pubmed>
- ↑ <pubmed>1321062</pubmed>
- ↑ <pubmed>17169623 </pubmed>
- ↑ <pubmed>16815385</pubmed>
- ↑ <pubmed>14723847</pubmed>
- ↑ <pubmed>9784595</pubmed>
- ↑ <pubmed>1315677</pubmed>
- ↑ <pubmed>20489451 </pubmed>
- ↑ <pubmed>15863034 </pubmed>
- ↑ <pubmed>17552943 </pubmed>
- ↑ <pubmed>11779141 </pubmed>
- ↑ <pubmed>8432397 </pubmed>
- ↑ <pubmed>8630492 </pubmed>
- ↑ 18.0 18.1 <pubmed>12080084</pubmed>
- ↑ 19.0 19.1 <pubmed>10662638</pubmed>