File:Mouse E13.5 gene expression.jpg: Difference between revisions
(==Analysis of Mouse Stage E13.5 Specific Gene Expression== 1140 transcripts were found to be significantly up-regulated exclusively at one developmental time point. A large fraction of these stage specific genes were found to be novel, in particular at E) |
No edit summary |
||
| Line 1: | Line 1: | ||
==Analysis of Mouse Stage E13.5 Specific Gene Expression== | ==Analysis of Mouse Stage E13.5 Specific Gene Expression== | ||
1140 transcripts were found to be significantly up-regulated exclusively at one developmental time point. A large fraction of these stage specific genes were found to be novel, in particular at E13.5 only 5% of the transcripts had confirmed musculoskeletal knockout phenotypes and 6% of the genes have been validated by in situ hybridization. A survey of available in situ expression profiles for genes with peak expression levels at | 1140 transcripts were found to be significantly up-regulated exclusively at one developmental time point. A large fraction of these stage specific genes were found to be novel, in particular at E13.5 only 5% of the transcripts had confirmed musculoskeletal knockout phenotypes and 6% of the genes have been validated by in situ hybridization. A survey of available in situ expression profiles for genes with peak expression levels at E13.5 revealed that they display a wide range of patterns, and include closely related members of gene families such as Irx, Smarc and zinc finger transcription factors. | ||
| Line 8: | Line 8: | ||
Figure 6. doi:10.1371/journal.pone.0028358.g006 | Figure 6. doi:10.1371/journal.pone.0028358.g006 | ||
[[Category:Mouse | [[Category:Mouse E13.5]] | ||
Latest revision as of 00:14, 14 February 2013
Analysis of Mouse Stage E13.5 Specific Gene Expression
1140 transcripts were found to be significantly up-regulated exclusively at one developmental time point. A large fraction of these stage specific genes were found to be novel, in particular at E13.5 only 5% of the transcripts had confirmed musculoskeletal knockout phenotypes and 6% of the genes have been validated by in situ hybridization. A survey of available in situ expression profiles for genes with peak expression levels at E13.5 revealed that they display a wide range of patterns, and include closely related members of gene families such as Irx, Smarc and zinc finger transcription factors.
- Limb Links: Mouse limb skeleton cartoon | Mouse limb tissue development | Limb Development | Mouse Development
Reference
Taher L, Collette NM, Murugesh D, Maxwell E, Ovcharenko I & Loots GG. (2011). Global gene expression analysis of murine limb development. PLoS ONE , 6, e28358. PMID: 22174793 DOI.
Copyright
This is an open-access article, free of all copyright, and may be freely reproduced, distributed, transmitted, modified, built upon, or otherwise used by anyone for any lawful purpose. The work is made available under the Creative Commons CC0 public domain dedication.
Figure 6. doi:10.1371/journal.pone.0028358.g006
File history
Yi efo/eka'e gwa ebo wo le nyangagi wuncin ye kamina wunga tinya nan
| Gwalagizhi | Nyangagi | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 00:12, 14 February 2013 | 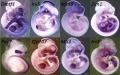 | 1,764 × 1,100 (266 KB) | Z8600021 (talk | contribs) | ==Analysis of Mouse Stage E13.5 Specific Gene Expression== 1140 transcripts were found to be significantly up-regulated exclusively at one developmental time point. A large fraction of these stage specific genes were found to be novel, in particular at E |
You cannot overwrite this file.
File usage
The following page uses this file: