File:Lymph node - high endothelial venule.jpg: Difference between revisions
| Line 14: | Line 14: | ||
** recognize chemokine X presented in the lumen | ** recognize chemokine X presented in the lumen | ||
** signalled to activate their integrins and firmly adhere to HEVs. | ** signalled to activate their integrins and firmly adhere to HEVs. | ||
* '''d''' - Some | * '''d''' - Some adhering lymphocytes | ||
** migrate across HEVs and are stimulated by chemokine Y, which is immobilized on MAC25. | |||
* '''e''' - Farther outside | * '''e''' - Farther outside | ||
** other chemokine-binding molecules, such as collagen IV and fibronectin, can capture a different chemokine (Z). | ** other chemokine-binding molecules, such as collagen IV and fibronectin, can capture a different chemokine (Z). | ||
Revision as of 10:23, 28 February 2012
Lymph Node - High Endothelial Venule
Chemokines and chemokine-binding molecules expressed in and around high endothelial venules (HEVs) in a concentric manner might function coordinately in lymphocyte trafficking across HEVs.
Original authors propose that several chemokine-binding molecules are expressed in a concentric manner in HEVs and their surrounds. The coordinated actions of chemokines and chemokine-binding molecules in and around HEVs are shown in the above cartoon in sequential order.
- a - In the lumen of HEVs
- heparan sulphate proteoglycans (red) can capture and present a lymphoid chemokine (X, green) in situ.
- Duffy antigen receptor for chemokines (DARC; black) — a non-signalling chemokine receptor — captures and scavenges inflammatory chemokines, such as CXC-chemokine ligand 1 (CXCL1; pink), which is constitutively produced by HEVs.
- b - In the basal lamina of HEVs
- another chemokine-binding protein MAC25 (dark blue) can capture a chemokine, such as CC-chemokine ligand 21 (CCL21), CXCL10 or CCL5 (Y, light blue).
- In addition, other components in the basal lamina such as collagen and fibronectin (grey) can capture a different chemokine (Z, yellow) expressed in the HEV area.
- c - Lymphocytes rolling along the HEV endothelium
- recognize chemokine X presented in the lumen
- signalled to activate their integrins and firmly adhere to HEVs.
- d - Some adhering lymphocytes
- migrate across HEVs and are stimulated by chemokine Y, which is immobilized on MAC25.
- e - Farther outside
- other chemokine-binding molecules, such as collagen IV and fibronectin, can capture a different chemokine (Z).
- Some of the extravasating lymphocytes might interact with chemokine Z and be stimulated to move into this area.
- f - By sequentially recognizing the multiple chemokines presented on the tissue matrix
- more and more lymphocytes extravasate and progressively move from the inside to the outside of HEVs.
See also Lymph node cartoon.
- Lymph Node Cartoons: Detailed structure | Cartoon with Histology | Lymphocyte traffic | Simple structure | Simple node anatomy | Wiki node image | Internal structure | Mesenteric lymph node | Histology | Gallery | Lymph Node Development
Reference
<pubmed>15122201</pubmed>| Nat Rev Immunol.
Adapted by permission from Macmillan Publishers Ltd: Nat. Rev. Immunol.: 2004, 4(5);360-70, copyright (2004)
Licensee: Mark A Hill License Number: 2857291045727 Publication: Nature Reviews Immunology
Title: Lymphocyte trafficking across high endothelial venules: dogmas and enigmas Type Of Use: post on the internet
https://s100.copyright.com/CustomerAdmin/PLF.jsp?lID=2012021_1330385293727
File history
Yi efo/eka'e gwa ebo wo le nyangagi wuncin ye kamina wunga tinya nan
| Gwalagizhi | Nyangagi | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 10:12, 28 February 2012 | 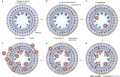 | 959 × 615 (193 KB) | Z8600021 (talk | contribs) | |
| 10:10, 28 February 2012 | 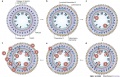 | 959 × 615 (192 KB) | Z8600021 (talk | contribs) | ==Lymph Node - High Endothelial Venule== Chemokines and chemokine-binding molecules expressed in and around high endothelial venules (HEVs) in a concentric manner might function coordinately in lymphocyte trafficking across HEVs. We propose that several |
You cannot overwrite this file.
File usage
The following page uses this file: