File:Neuropore cell shape changes.png: Difference between revisions
(Z5165679 uploaded a new version of File:Neuropore cell shape changes.png) |
mNo edit summary |
||
| Line 2: | Line 2: | ||
Cell shape changes in the midline after mid-hindbrain neuropore (MHNP) closure in vivo. a The relationship between the closing of MHNP and the somite stage in ICR strain mouse. b-f Dorsal views of embryos stained with E-cadherin (Ecad). Non-neural ectodermal cells change their shapes dramatically from bipolar to polygonal after the MHNP closure. g, h Higher manigification views of embryos (c, e) that were co-stained with Ecad, apoptotic marker active-Caspase 3 (aCasp 3), and nuclear staining (DNA). Dorsal views, left is rostral side. Scale bars: (b-f) 100 μm in upper panels, and 25 μm in (g, h) middle and (b-f) lower panels | Cell shape changes in the midline after mid-hindbrain neuropore (MHNP) closure in vivo. a The relationship between the closing of MHNP and the somite stage in ICR strain mouse. b-f Dorsal views of embryos stained with E-cadherin (Ecad). Non-neural ectodermal cells change their shapes dramatically from bipolar to polygonal after the MHNP closure. g, h Higher manigification views of embryos (c, e) that were co-stained with Ecad, apoptotic marker active-Caspase 3 (aCasp 3), and nuclear staining (DNA). Dorsal views, left is rostral side. Scale bars: (b-f) 100 μm in upper panels, and 25 μm in (g, h) middle and (b-f) lower panels | ||
===Reference=== | |||
{{#pmid:30064364}} | |||
Revision as of 11:49, 7 August 2018
Fig. 3
Cell shape changes in the midline after mid-hindbrain neuropore (MHNP) closure in vivo. a The relationship between the closing of MHNP and the somite stage in ICR strain mouse. b-f Dorsal views of embryos stained with E-cadherin (Ecad). Non-neural ectodermal cells change their shapes dramatically from bipolar to polygonal after the MHNP closure. g, h Higher manigification views of embryos (c, e) that were co-stained with Ecad, apoptotic marker active-Caspase 3 (aCasp 3), and nuclear staining (DNA). Dorsal views, left is rostral side. Scale bars: (b-f) 100 μm in upper panels, and 25 μm in (g, h) middle and (b-f) lower panels
Reference
Shinotsuka N, Yamaguchi Y, Nakazato K, Matsumoto Y, Mochizuki A & Miura M. (2018). Caspases and matrix metalloproteases facilitate collective behavior of non-neural ectoderm after hindbrain neuropore closure. BMC Dev. Biol. , 18, 17. PMID: 30064364 DOI.
File history
Yi efo/eka'e gwa ebo wo le nyangagi wuncin ye kamina wunga tinya nan
| Gwalagizhi | Nyangagi | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 11:47, 7 August 2018 | 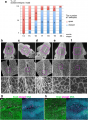 | 1,418 × 1,940 (1.01 MB) | Z5165679 (talk | contribs) | Fig. 3 Cell shape changes in the midline after mid-hindbrain neuropore (MHNP) closure in vivo. a The relationship between the closing of MHNP and the somite stage in ICR strain mouse. b-f Dorsal views of embryos stained with E-cadherin (Ecad). Non-neur... |
| 11:46, 7 August 2018 | 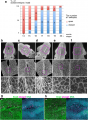 | 1,418 × 1,940 (1.01 MB) | Z5164572 (talk | contribs) | Fig. 3 Cell shape changes in the midline after mid-hindbrain neuropore (MHNP) closure in vivo. a The relationship between the closing of MHNP and the somite stage in ICR strain mouse. b-f Dorsal views of embryos stained with E-cadherin (Ecad). Non-neur... | |
| 11:46, 7 August 2018 | 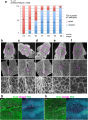 | 1,418 × 1,940 (1.01 MB) | Z5164785 (talk | contribs) | Fig. 3 Cell shape changes in the midline after mid-hindbrain neuropore (MHNP) closure in vivo. a The relationship between the closing of MHNP and the somite stage in ICR strain mouse. b-f Dorsal views of embryos stained with E-cadherin (Ecad). Non-neu... | |
| 11:46, 7 August 2018 | 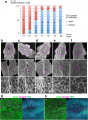 | 1,418 × 1,940 (1.01 MB) | Z8600021 (talk | contribs) | Fig. 3 Cell shape changes in the midline after mid-hindbrain neuropore (MHNP) closure in vivo. a The relationship between the closing of MHNP and the somite stage in ICR strain mouse. b-f Dorsal views of embryos stained with E-cadherin (Ecad). Non-neu... |
You cannot overwrite this file.
File usage
The following 7 files are duplicates of this file (more details):
The following 14 pages use this file: