2016 Group Project 2: Difference between revisions
No edit summary |
|||
| Line 28: | Line 28: | ||
Four Notch proteins are involved in the canonical pathway. NOTCH1 to NOTCH4 are single transmembrane receptors and can interact with a variety of ligands, including NOTCH ligands (e.g. Delta ligands) and Serrate ligands. There are three Delta ligands (Dll1, Dll3, and Dll4) and two Serrate ligands (Jagged1 and Jagged2) present in mammals.<ref name="Introduction to Notch Signalling"/> The binding between the Notch receptor and the ligand on the adjacent cell induces the release of the Notch intracellular domain (NICD) via a sequence of proteolytic reactions.<ref name="The Developing Human"/> Cell-cell interaction is therefore critical in the process of triggering Notch signalling. | Four Notch proteins are involved in the canonical pathway. NOTCH1 to NOTCH4 are single transmembrane receptors and can interact with a variety of ligands, including NOTCH ligands (e.g. Delta ligands) and Serrate ligands. There are three Delta ligands (Dll1, Dll3, and Dll4) and two Serrate ligands (Jagged1 and Jagged2) present in mammals.<ref name="Introduction to Notch Signalling"/> The binding between the Notch receptor and the ligand on the adjacent cell induces the release of the Notch intracellular domain (NICD) via a sequence of proteolytic reactions.<ref name="The Developing Human"/> Cell-cell interaction is therefore critical in the process of triggering Notch signalling. | ||
<br><br> | <br><br> | ||
The interaction on the cell surface between Notch and its ligand on an adjacent cell causes the extracellular metalloprotease site (S2 site) to be exposed. The S2 site is then cleaved by transmembrane proteases belonging to the a disintegrin and metalloproteinase/tumour necrosis factor α converting enzyme (ADAM/TACE) family. The remaining Notch fragment subsequently undergoes two more intramembranous cleavages at the S3/S4 sites by the γ-secretase complex. Finally, the NICD is released and enters the nucleus | The interaction on the cell surface between Notch and its ligand on an adjacent cell causes the extracellular metalloprotease site (S2 site) to be exposed. The S2 site is then cleaved by transmembrane proteases belonging to the a disintegrin and metalloproteinase/tumour necrosis factor α converting enzyme (ADAM/TACE) family. The remaining Notch fragment subsequently undergoes two more intramembranous cleavages at the S3/S4 sites by the γ-secretase complex. Finally, the NICD is released and enters the nucleus to interacting with CSL (CBF1, Suppressor of Hairless, Lag-1). CSL is a DNA-binding protein that acts as a transcription factor by forming repressor or activator complexes that interact with Notch target genes.<ref name=PMID10075488/> To stop signalling, the NICD is phosphorylated by kinases and ubiquitinated by E3 ubiquitin ligases, which results in proteasome mediated degradation and subsequent termination of the signal.<ref name="Introduction to Notch Signalling"/> | ||
<br><br> | <br><br> | ||
====Non-canonical pathway==== | |||
<br> | |||
====Transcriptional regulation of Notch signalling==== | ====Transcriptional regulation of Notch signalling==== | ||
<br> | |||
===Roles in Embryonic Development=== | ===Roles in Embryonic Development=== | ||
Revision as of 22:01, 22 September 2016
| 2016 Student Projects | ||||
|---|---|---|---|---|
| Signalling: 1 Wnt | 2 Notch | 3 FGF Receptor | 4 Hedgehog | 5 T-box | 6 TGF-Beta | ||||
| 2016 Group Project Topic - Signaling in Development
OK you are now in a group, add a topic with your student signature to the group page. | ||||
| This page is an undergraduate science embryology student project and may contain inaccuracies in either descriptions or acknowledgements. | ||||
| Group Assessment Criteria |
|---|
 Science Student Projects Science Student Projects
|
| More Information on Assessment Criteria | Science Student Projects |
Notch signalling pathway
Introduction
The Notch signalling pathway is critical for cell differentiation, proliferation, and apoptosis. It is involved in embryonic organ development through the regulation of cell-cell signalling; specifically lateral inhibition, formation of boundaries, and cell lineage assignation.[1][2]
History
| 1914 | First description of a "notch" defect (loss of tissue in the wing) in Drosophila by John S. Dexter, giving the gene its name |
| 1917 | First allele of Notch is identified by Thomas Hunt Morgan |
| 1930s | Donald F. Poulson conducts research into the involvement of Notch in development |
Overview of Molecular Mechanisms
Mammals possess a total of four Notch genes, with five genes for encoding the associated ligands, Delta-like and Jagged. The Notch genes each code for a single transmembrane receptor. Extracellularly, it contains epidermal growth factor (EGF)-like repeats for ligand interaction and Lin-12-Notch (LN) repeats for regulating the interactions between the extracellular and intracellular regions. Intracellularly, Notch has seven ankyrin (ANK) repeats and a transactivation domain (TAD), as well as a proline, glutamine, serine, threonine-rich (PEST) domain for degradation of Notch.
Canonical pathway
The canonical Notch pathway is unique in that it involves direct interaction between adjacent cells, as opposed to paracrine signalling, because both the Notch ligands and Notch receptors are transmembrane proteins found in the cell membrane. Furthermore, the lack of a secondary messenger or amplification process means that the Notch pathway has limited opportunities for regulation and must therefore be tightly controlled. Depending on its developmental and cellular context, activation or inhibition of the pathway can result in a variety of cellular responses, including cell death, proliferation, or differentiation.[3]
Four Notch proteins are involved in the canonical pathway. NOTCH1 to NOTCH4 are single transmembrane receptors and can interact with a variety of ligands, including NOTCH ligands (e.g. Delta ligands) and Serrate ligands. There are three Delta ligands (Dll1, Dll3, and Dll4) and two Serrate ligands (Jagged1 and Jagged2) present in mammals.[3] The binding between the Notch receptor and the ligand on the adjacent cell induces the release of the Notch intracellular domain (NICD) via a sequence of proteolytic reactions.[1] Cell-cell interaction is therefore critical in the process of triggering Notch signalling.
The interaction on the cell surface between Notch and its ligand on an adjacent cell causes the extracellular metalloprotease site (S2 site) to be exposed. The S2 site is then cleaved by transmembrane proteases belonging to the a disintegrin and metalloproteinase/tumour necrosis factor α converting enzyme (ADAM/TACE) family. The remaining Notch fragment subsequently undergoes two more intramembranous cleavages at the S3/S4 sites by the γ-secretase complex. Finally, the NICD is released and enters the nucleus to interacting with CSL (CBF1, Suppressor of Hairless, Lag-1). CSL is a DNA-binding protein that acts as a transcription factor by forming repressor or activator complexes that interact with Notch target genes.[2] To stop signalling, the NICD is phosphorylated by kinases and ubiquitinated by E3 ubiquitin ligases, which results in proteasome mediated degradation and subsequent termination of the signal.[3]
Non-canonical pathway
Transcriptional regulation of Notch signalling
Roles in Embryonic Development
The phylogenetically conserved Notch signalling pathway plays a crucial role in the development of multiple organ systems, and is a major regulator of stem cell fate. It is responsible for the regulation of the transcription of a number of signalling molecules, such as MyoD, Mash1 and GATA2, which are genes controlling the fate of myogenic, neurogenic and haematopoietic stem cells, respectively. [4] The following subsections will further elucidate the vital roles of Notch signalling during normal embryonic development.
Cardiovascular
Cardiomyocyte Specification and Differentiation
Expression during the appropriate window of the timeline of embryogenesis of Notch receptors, ligands and downstream effector molecules elucidates a role for the Notch pathway in the earliest stages of cardiac development. It has been found to restrict the expression of specific cardiogenic genes in a spatiotemporal manner and regulate cardiac field specification as early as during gastrulation. [5] Interestingly, Notch has been found to play both suppressive and promoting roles in cardiogenesis.
For example, it has been shown that Notch suppresses cardiomyocyte cell fate specification during early cardiogenesis. This has been demonstrated through studies such as that carried out by Rones and colleagues (2000), which used activation and inhibition of Notch signaling in Xenopus. [6]
On the other hand, studies such as that by Boni and colleagues (2008) have found that Notch signalling may also promote myogenesis from cardiac progenitor cells. [4] Cardiogenesis has also been promoted by downregulating Notch-1 activity in stem cells of embryos (Nemir et al., 2006).[7]
Despite the understanding that Notch signalling is crucial to embryonic myogenesis, the exact molecular mechanism remains elusive. Research by Buas and colleagues (2010) has explored such mechanisms by studying the Notch target, Hey1, which is known to suppress myogenic differentiation. They concluded that this inhibitory function of Hey1 is primarily mediated through binding near to myogenin and Mef2C promoters, which leads to cessation of target gene expression. [8]
Development of the Atrioventricular Canal
A study by Rutenberg and colleagues (2006) and another by Kokubo and colleagues (2007) implicate a role for Notch signaling in the region between the atria and the ventricles of the heart (the atrioventricular canal, or AVC).[9] [10] They used chicken and mouse models, respectively, to show that other signalling factors, Bmp2 and Tbx2, are restricted to the AVC region by Notch signalling during development.
Furthermore, Watanabe and colleagues (2006) showed that deletion of Notch targets increases Bmp2 expression and expansion of the AVC tissue, however other, non-Notch restrictive factors involved in AVC development are likely to exist.[11]
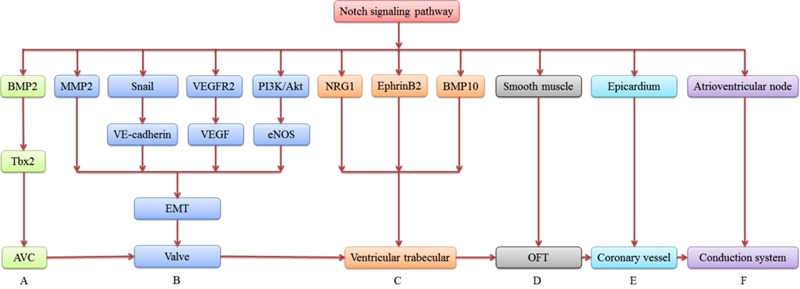
See the summary diagram below for a schematic representation of this molecular interaction during AVC development.
Heart Valve Development
In order for the heart valve to properly form in the embryo, endocardial to mesenchymal transformation (EMT) must occur. The Notch pathway, alongside Wnt and Bmp, has been found to regulate the process of EMT, defects in which can lead to congenital heart valve disease. Interestingly, Timmerman and colleagues (2004) demonstrated that this role of Notch may also promote oncogenic transformation. This team also showed that embryos exhibited abortive endocardial EMT if they were deficient in Notch signalling components in vivo and in vitro. [12]
A more recent paper by Wang and colleagues (2013) further explored the underlying signalling processes and interrelationships of molecules that impact EMT. They found that Jagged1-Notch1 signalling in cells of the endocardium potentiates expression of Wnt4, which in turn carries out paracrine action on adjacent AVC tissue to upregulate Bmp2 expression and thus signal EMT. [13]
Trabeculation
Trabeculation is the initial process of ventricular chamber development that forms a series of cardiomyocyte projections within the lumens of the ventricles of the heart (called trabeculae). Grego-Bessa and colleagues (2007) addressed the roles Notch plays in trabeculation through RBPJk (the gene product of which interacts with Notch) and Notch1 gene manipulation. Mutants of these two genes showed perturbed expression and signalling of EphrinB2, NRG1 and BMP10, alongside reduced proliferation of myocardiocytes. This research ultimately suggested that EphrinB2 is a direct target of Notch in the endocardium that simultaneously requires the Notch-dependent action of BMP10 and NRG1 in order for ventricular myocardium proliferation and differentiation to occur normally. [14] Furthermore, the important process of trabeculation has recently been shown to be controlled by sequential Notch activation by an investigation by D’Amato and colleagues (2015). [15]
Development of the Outflow Tract
TBC
Central Nervous System
Early Neural Differentiation
Notch plays a major role in promoting neural commitment of cells. Lowell and colleagues (2011) used genetic manipulation to discover that the phenotype of stem cells is not affected by constitutively activated Notch in mouse embryonic stem cells (mESCs), however, interfering with Notch signalling -for example by pharmacological means- did impede neural fate determination. This role required Notch signalling via fibroblast growth factor (FGF) receptors. Furthermore, the conservation of the Notch signalling pathway within pluripotent stem cells is implied due to the existence of Notch ligands in stromal cells in human embryonic stem cells (hESCs) that induce neural differentiation. [17]
Das and colleagues (2010) manipulated Notch protein levels during specific stages of neural differentiation and found that if Notch signalling pathways were activated during day 3 of neural development for 6 hours, cell proliferation was dramatically enhanced. This was attributed to the induction by Notch of cyclin D1 expression. Without Notch signalling during neural development, there was reduced cyclin D1 levels. Manipulation of mESCs to express a dominant negative form of cyclin D1 resulted in abrogation of cell proliferation stimulated by Notch. Overall these results imply a temporally-specific role for Notch in CNS development, and that it requires cyclin D1 as a signalling molecule. [18]
Other Systems
Roles in Animal Development
Drosophila melanogaster
Unlike the four Notch paralogs in humans, only one Notch homolog is present in Drosophila.[3]
Research has shown that Notch is crucial for the formation of longitudinal connections in the Drosophila CNS. Kuzina, Song, and Giniger (2011) created temperature-sensitive mutations of Notch genes that prevented the development of mature longitudinal axon tracts. They also found that the Notch phenotype appears at the earliest stages of the development of longitudinal connections in the CNS by observing early stage 13 embryos.[19]
Caenorhabditis elegans
Danio rerio
Abnormalities in Notch signalling
Alagille syndrome
Alagille syndrome (AGS) is an autosomal dominant, multisystem disorder that mainly affects the liver, heart, and kidney. Diagnostic characteristics of the disease include liver disease, cardiac disease, vertebral defects, eye conditions, and facial features, as well as renal and vascular abnormalities.[20] In 94% of clinically diagnosed cases, a mutation in the gene encoding the Notch ligand JAG1 has been identified as a contributing factor. In combination with this, a mutation in the NOTCH2 gene has also been implicated in the manifestation of AGS in some patients.[21]
Cerebral Autosomal-Dominant Arteriopathy with Subcortical Infarcts and Leukoencephalopathy (CADASIL)
CADASIL is an autosomal-dominant disease of the small to medium-sized arteries, mainly in the brain, that leads to dementia and disability in mid-life. The symptoms, age of onset, and prognosis are variable. Distinguishing symptoms include subcortical ischemic events (60-80% of cases), cognitive impairment (60% of late stage disease), migraines (30-40%), mood disturbances (30%), and apathy (40%).[22][23] Approximately 5-10% of CADASIL patients also experience seizures. The mean age of onset is 45 years of age and the disease duration is between 10–40 years. More than 95% of CADASIL cases present with pathogenic mutations in NOTCH3 (located on chromosome 19p13), specifically the epidermal growth factor-like repeat domain.[23][24] The NOTCH3 gene is involved in the normal development of blood vessels in both fetal and adult brains. In adults, NOTCH3 is expressed in the smooth muscle cells of arteries.[22] There is currently no effective treatment available for CADASIL.
Congenital Heart Defects
Current Areas of Research
Glossary
References
- ↑ 1.0 1.1 Moore, K.L., Persaud, T.V.N. & Torchia, M.G. (2015). The developing human: clinically oriented embryology (10th ed.). Philadelphia: Saunders.
- ↑ 2.0 2.1 <pubmed>10075488</pubmed>
- ↑ 3.0 3.1 3.2 3.3 Yamamoto, S., Schulze, K.L. & Bellen, H.J. (2014). Introduction to Notch Signalling. Notch Signaling: Methods and Protocols. Methods in Molecular Biology: 1187
- ↑ 4.0 4.1 <pubmed>18832173</pubmed>
- ↑ <pubmed>19580804</pubmed>
- ↑ <pubmed>10934030</pubmed>
- ↑ <pubmed>16690879</pubmed>
- ↑ <pubmed>19917614</pubmed>
- ↑ <pubmed>17021042</pubmed>
- ↑ <pubmed>17259303</pubmed>
- ↑ <pubmed>16554359</pubmed>
- ↑ <pubmed>14701881</pubmed>
- ↑ <pubmed>23560082</pubmed>
- ↑ <pubmed>17336907</pubmed>
- ↑ <pubmed>26641715</pubmed>
- ↑ <pubmed>24345875</pubmed>
- ↑ <pubmed>16594731</pubmed>
- ↑ <pubmed>20887720</pubmed>
- ↑ <pubmed>21447553</pubmed>
- ↑ <pubmed>26548814</pubmed>
- ↑ <pubmed>16773578</pubmed>
- ↑ 22.0 22.1 <pubmed>21045164</pubmed>
- ↑ 23.0 23.1 <pubmed>20301673</pubmed>
- ↑ <pubmed>24579972</pubmed>