User:Z5020373
| Student Information (expand to read) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Individual Assessments | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Please leave this template on top of your student page as I will add your assessment items here. Beginning your online work - Working Online in this course
Click here to email Dr Mark Hill | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 1 Assessment - Researching a Topic | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
In the lab I showed you how to find the PubMed reference database and search it using a topic word. Lab 1 assessment will be for you to use this to find a research reference on "fertilization" and write a brief summary of the main finding of the paper.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 2 Assessment - Uploading an Image | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
OK you are now in a group
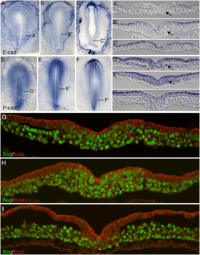
Initially the topic can be as specific or as broad as you want. Chicken embryo E-cad and P-cad gastrulation[1] References
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 4 Assessment - GIT Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
ANAT2341 Quiz Example | Category:Quiz | ANAT2341 Student 2015 Quiz Questions | Design 4 quiz questions based upon gastrointestinal tract. Add the quiz to your own page under Lab 4 assessment and provide a sub-sub-heading on the topic of the quiz. An example is shown below (open this page in view code or edit mode). Note that it is not just how you ask the question, but also how you explain the correct answer. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 5 Assessment - Course Review | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Complete the course review questionnaire and add the fact you have completed to your student page. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 6 Assessment - Cleft Lip and Palate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 7 Assessment - Muscular Dystrophy | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 8 Assessment - Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| A brief quiz was held in the practical class on urogenital development. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 9 Assessment - Peer Assessment | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 10 Assessment - Stem Cells | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
As part of the assessment for this course, you will give a 15 minutes journal club presentation in Lab 10. For this you will in your current student group discuss a recent (published after 2011) original research article (not a review!) on stem cell biology or technology.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 11 Assessment - Heart Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Read the following recent review article on heart repair and from the reference list identify a cited research article and write a brief summary of the paper's main findings. Then describe how the original research result was used in the review article.
<pubmed>26932668</pubmed>Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Team role: Resource investigator
After reading about the various team roles, I have concluded that I am probably the Resource Investigator in any team situation. I am usually very enthusiastic about new projects, and I like to search about the internet for articles or research papers. By the end of it I find myself hoarding all these articles into a folder. I do like to build on ideas rather than create them which is something I identified with in the description. I always have a positive attitude and I like connecting people because it makes me happy to see everyone working together.
Fertilisation
The most interesting thing I found about this lecture would probably be learning about the polar bodies. Up until this lecture I never really gave it much thought as to where the extra DNA from meiosis goes, so finding about the existence of not 2, but 3 polar bodies was really interesting. I also thought that the Zona Pellucida protein (ZP2) is a very fascinating but smart mechanism to make human spermatozoa species specific.
| Mark Hill 4 August 2016 - Thank you for adding this content before the lab. I would suggest that rather than using a template that you simply paste on this current page with separate subheadings for each lab/assessment item. Also please no names, just your student number. |
Lab 1
Lab Attendance
Z5020373 (talk) 14:36, 5 August 2016 (AEST)
Lab Demonstrations
External Link
Internal Link
https://embryology.med.unsw.edu.au/embryology/index.php/2011_Lab_1
Referencing
PMID 27486280
Lab 1 Assessment
<pubmed>27123200</pubmed>
This research article explores whether choosing to a conduct embryo transfer (ET) on a weekend or weekday will affect the success of clinical pregnancy in IVF procedures. In the study, ET transfers were performed on either weekdays or weekends in patients with similar clinical characteristics, such as age, body-mass index and duration of infertility. Clinical pregnancy was determined using blood pregnancy tests and ultrasound examination and was defined as the “presence of a gestational sac with a foetal heart beat.” After the ETs, the authors found that there was an overall 42.8% success rate of clinical pregnancy in patients from both groups, with a 14.6% increase in the pregnancy rate when weekend ETs where compared to weekday ETs. The study however, did not examine any possible reasons to explain this increase in implantation rate although a few potential factors were discussed from previous findings in this area of research. These included endometrial receptivity which occurs 5 days after the post-ovulatory progesterone surge. The article mentioned that uterine receptivity and implantation could be affected by the junctional zone. The extent of junctional zone contractility differs throughout the ovarian cycle and an increased contractility just before ET has been previously shown to significantly decrease the likelihood of successful implantation. Since weekends are more relaxing than weekdays they suggest a possible correlation between that and reduced junctional zone contractions leading to easier ETs. Therefore from this study, it was concluded that ETs performed during the weekends are more successful than those performed during the weekdays identifying a potential factor that can improve ETs in IVF situations.
| Mark Hill (talk) 14:24, 15 August 2016 (AEST) - This is a good summary of a paper that looks at potential environmental/endocrine effects on reproductive fertility. You needed to put the reference at the top rather than just the PMID number, fix this and you can get this full mark for the exercise.
Mark Hill 18 August 2016 - You did not fix, so I have adjusted the final mark. |
Assessment 4/5 |
Lab 2
Lab Attendance
Z5020373 (talk) 14:41, 12 August 2016 (AEST)
Lab 2 Assessment
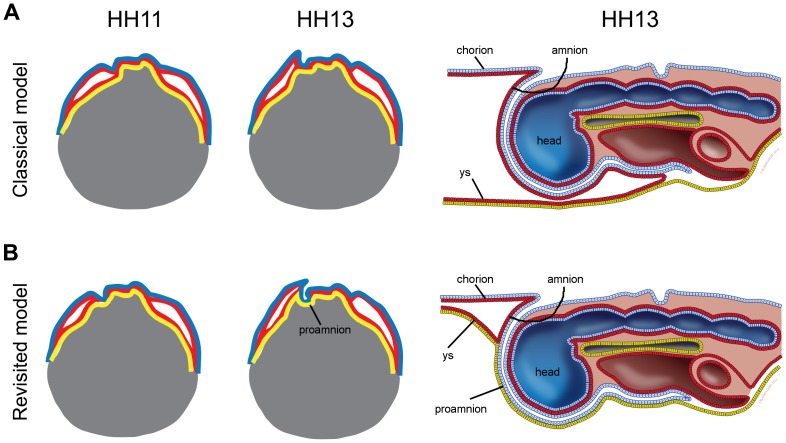
Chicken embryo amion fold development[1]
| Mark Hill 14:24, 15 August 2016 (AEST) - Very good, the image relates to early development and contains the reference, copyright and student template. (5/5) |
Lab 3
Lab Attendance
Z5020373 (talk) 13:53, 19 August 2016 (AEST)
| Mark Hill 31 August 2016 - Lab 3 Assessment Quiz - Mesoderm and Ectoderm development. | Assessment 2.5/5 |
Lab 4
Lab Attendance
Z5020373 (talk) 13:02, 26 August 2016 (AEST)
Lab 4 Assessment
Take the Quiz
Make your selection for all questions before clicking submit.
| Mark Hill 13 October 2016 - These seem well designed GIT quiz questions that test topic understanding. | Assessment 5/5 |
Lab 5
Lab Attendance
Z5020373 (talk) 13:11, 2 September 2016 (AEST)
Lab 5 Assessment
Questionaire completed and submitted Z5020373 (talk) 13:17, 9 September 2016 (AEST)
| Mark Hill 11 October 2016 - Questionnaire on course structure. | Assessment 5/5 |
Lab 6
Lab Attendance
Z5020373 (talk) 13:05, 9 September 2016 (AEST)
Lab 6 Assessment
<pubmed>25382630</pubmed> Cleft palate arises when the bilateral palatal shelves fail to fuse. Many genetic and environmental factors have been identified to contribute to this deformity. One genetic mutation associated with cleft palate is the loss transforming growth factor-beta receptor (TGF-βR). A recent article by Hill et al. demonstrated that loss of TGF-βR3 reduced the expression of several ligands and receptors in the TGF-β/BMP family, including three TGF-β ligands and BMP2. During embryonic development, these molecules are involved in cell growth and differentiation.
A loss of TGF-β/BMP signaling, by receptor loss was found to be associated with cleft palate formation due to aberrant cell cycle progression and altered gene expression. Furthermore changes to TGF-β/BMP signaling also interrupted vascular development and remodeling as well as osteogenic differentiation during palate formation. TGF-βR3 is therefore essential for maintaining the expression of TGF-β and BMP molecules and without this receptor processes of palatal shelf elongation, elevation and fusion are disrupted leading to cleft palate.
| Mark Hill 13 October 2016 - Good reference and explanation. | Assessment 5/5 |
Lab 7
Lab Attendance
Z5020373 (talk) 13:05, 16 September 2016 (AEST)
Lab 7 Assessment
1. What is/are the dystrophin mutation(s)?
- The dystrophin gene is located on the short arm of the X chromosome at position 21.2. [2]
- It is the largest gene found in nature spanning 1.5% of the X-chromosome which is about 2.5 Mb of genomic sequence. [3]
- Its size could explain why it is so susceptible to spontaneous mutations.
- Encodes for Dystrophin protein.
- Mutations such as large deletions (60-70% of DMD cases), large duplications (10% of DMD cases) and point mutations (15-30% of DMD cases) can occur resulting in gene inactivation (therefore loss of function).[4]
2. What is the function of dystrophin?
- Dystrophin is part of a protein complex that work together to strengthen muscle fibers and protect them from injury as muscles contract and relax.
- Dystrophin complex acts as an anchor, connecting each muscle cell's structural framework (cytoskeleton) with the lattice of proteins and other molecules outside the cell (extracellular matrix).
- May also play a role in cell signaling by interacting with proteins that send and receive chemical signals e.g. in alpha-syntrophin.[5]
3. What other tissues/organs are affected by this disorder?
- Muscle (skeletal, cardiac and smooth): fatigue, difficulty with motor skills
- Respiratory system: pneumonia
- Cardiac system: cardiac myopathy
- Central Nervous system (CNS): neurobehavioural disorders e.g. ADHD and dyslexia [6]
4. What therapies exist for DMD?
- There is no cure available for DMD, and the current interventions are based on preventing further muscle wasting and management of symptoms and complications
- Treatments include:
- Physical therapy
- Orthopedic appliances
- Medication:
- Two corticosteroids mainly used in DMD treatment are Prednisone/Prednisolone and Deflazacort, an oxazoline derivative of prednisolone, administered by two common regimens: daily and intermittent [7]
- Prednisone and prednisolone show an anti-inflammatory effect
- Deflazacort acts on muscle regeneration and differentiation
- Future could include:
- Cell-based therapies using stem cells to replace dystrophin gene (a potential cure).
- Gene therapies to deliver a therapeutic gene to skeletal and cardiac muscle, in order to restore the dystrophin protein PMID [8]
- A study by Nelson et al. in 2016, showed that in vivo CRISPR-Cas9–mediated dystrophin restoration in mdx mouse model of DMD removed the mutated exon 23 from the dystrophin gene and improved muscle structure and function.[9]
5. What animal models are available for muscular dystrophy?
- The most widely used animal model for DMD is the mdx mouse, which has a spontaneous point mutation in exon 23 that causes the absence of the dystrophin protein in the muscle.
- However, other animal models inlcude:[10]
- GRMD dog - Dystrophin-deficient dog
- HFMD cat - Dystrophin-deficient cat (clinically a poor model)
- This animal model allows testing ans screening of potential treatments and without these animal models, and without these animal models we would not have any known therapies today
- For example, mdx in vivo studies have led to U.S. FDA approval of Exondys 51 (eteplirsen) injection, the first drug approved to treat patients with Duchenne muscular dystrophy (DMD). Exondys 51 is specifically indicated for patients who have a confirmed mutation of the dystrophin gene amenable to exon 51 skipping, which affects about 13 percent of the population with DMD. [11]
| Mark Hill 13 October 2016 - Muscular Dystrophy questions have been comprehensively answered and you have cited your sources. | Assessment 5/5 |
Lab 8 Assessment
Completed the quiz during the lab.
| Mark Hill 27 October 2016 - Well done. Q3 gonads determine the other genital development. Q5 theca and interstitial. | Assessment 6/8 |
Lab Attendance
Z5020373 (talk) 13:07, 23 September 2016 (AEST)
Lab 8 Assessment
These are very good reviews of the project pages, with some specific examples. They include a balanced critical assessment, given the existing status of some of these pages. 8/10
Group 1: Wnt signalling pathway
Firstly, a proper introduction to the Wnt pathway and its various roles in the developing embryo should be added to the page. Even though the page is focusing on Wnt signalling in fetal skin development I still think the other contributions of Wnt signalling need to be at least mentioned in the intro. Mark also suggested adding a table to show the origins of the pathway and how our knowledge about Wnt signaling has evolved.
The page contains appropriate headings and subheadings to address the different mechanisms of Wnt signalling (canonical, non-canonical and non-canonical Wnt/Ca2+ pathway). The main points for each of these pathways are up on the page which is good. Clearly, these signalling mechanisms are quite complex and adding images could help the reader to visualise how the key molecules in this pathway interact with other molecules to induce downstream effects. But otherwise, the detail in the canonical pathway is adequate for a student to understand.
The references included in each section of the page is evidence of research done to support the content on the page. The page also includes up to date information to reflect current research in this area. This information should be integrated into some sort of discussion to show how this contributes to knowledge about the signalling pathway affects developmental events. Also, all the references need to be cited correctly which I am sure you guys are aware of and will do.
Don’t forget the topic is ‘Signalling in Development’, therefore the focus should be on Wnt signalling in the developing embryo.
Overall, Group 1 has made good progress. Keep it up!
Group 2: Notch signalling pathway
The page has a proper introduction stating the critical functions of Notch signalling, examples of diseases associated with Notch mutations as well as a brief description of the mechanism of Notch signalling. There is also a timeline on the page which none of the other groups have managed to do so good job!
Some headings are missing text but those with content (e.g. canonical pathway section) are covered in extensive detail. The subheadings and headings chosen for the page indicate the group’s in depth understanding of the key components of the Notch signalling pathway. Furthermore, the page contains correct referencing and citation of text and images.
The page also discusses Notch signalling in animal models as well. A recent study was included for Drosophila. What about the other two?
Overall, Group 2 has made excellent progress. Well done!
Group 3: FGFR signalling pathway
The headings and subheadings on the page is used very effectively to aid the progression of information. Through the sequence of the headings, it allows the reader to build their understanding about FGFR signalling. The FGFR page definitely address the topic of this assessment - signalling in development, and links FGF signalling to a number of developmental events. This reflects the large contributions of FGFR in development which the page successfully portrays.
In the overview section, it states: “As shown in the image, an acidic box…”. Make it clear which image you are referring to because I can’t find it.
The table for the subtypes of FGFR has been acknowledged that it is incomplete but it gives a good snapshot to function and associated abnormalities of the different FGFR subtypes.
The page includes a student drawn image which summarises the FGFR signalling pathway. None of the other groups have included a student drawn image so good job! The images uses colours to distinguish particular molecules and shows the downstream signalling events to affect gene transcription in the cell. To me the image is a bit blurry on the page, so maybe change the pixels of the image to make it larger and easier to see?
The page includes a quiz which is clever and will definitely make the page stand out from the other groups.
Overall, Group 3 has made good progress. Good job!
Group 4: Hedgehog signalling pathway
Firstly, the page is missing an introduction to the signalling pathway. There is also text missing under the first few subheadings. Since the hedgehog pathway research began as early as the 1970s, a table including the key events in the Hedgehog research would be interesting to add.
The page includes a nice overview of the Hedgehog pathway captured in the image however, it needs a reference to acknowledge the original source of the image. Consider relocating the image to the mechanism of signalling section. This may help the reader understand the processes better if they have that image there.
Mammals have 3 Hedgehog homologues (DHH, IHH and SHH). I think that is an important point to mention.
Good discussion of animal models since it is one of the key regulators of animal development. Despite having headings without text. Group 3 has made good progress so far. Keep it up!
Group 6: TGF-β signalling pathway
The page contains a good introduction to the TGF-β superfamily, including examples of other signaling proteins. The images help the reader visualise how TGF-β ligand bring the receptors together in a heterotetrameric complex in which the type II receptors phosphorylate and activate the type I receptors. To be pedantic, the second image still needs to include details of the original source.
Remember that the project is meant to focus on ‘Signalling in Development’ and whilst the page addresses the signalling component it does not discuss TGF-β signalling in the development of the embryo. The second image addresses its role in proliferation, migration, growth arrest and apoptosis. This pubmed article: PMID 19289080 discusses TGF-β signalling in early development, axis formation, and patterning of the embryo.
The page does not use the correct referencing or in-text citations.
Overall, Group 6 has made a good start but more research needs to be done. Keep pushing :)
Lab 9
Lab Attendance
Z5020373 (talk) 13:06, 7 October 2016 (AEDT)
Lab 11
Lab Attendance
Z5020373 (talk) 14:02, 21 October 2016 (AEDT)
Lab 11 Assessment
5/5
<pubmed>21350179</pubmed>
Transient Regenerative Potential of the Neonatal Mouse Heart primary literature findings: Based on the fact that urodele amphibians and teleost fish are able to retain the capacity for cardiac regeneration throughout their life, the authors questioned whether the potential for cardiac regeneration mammals is absent, or whether it exists but merely switched off early after birth. They investigated this through observing how the hearts of one day old neonatal mice heart develops. To perform further tests they surgically resected the left ventricular apex to see how the neonatal hearts respond to injury. They were able to see that the the ventricular apex in the mice indeed stimulated a regenerative response that appears to restore the damaged heart to its normal anatomy and function. This regenerative response was marked by cardiomyocyte proliferation with little evidence of hypertrophy or fibrosis, and therefore not a result repair processes. The origin of these cardiomyocytes in the regenerated tissue were determined to be from preexisting cardiomyocytes. The regenerated ventricular apex also had normal systolic function indicating a restoration of normal contractile function. A similar approach was done to seven day old mice and unlike the one day old mice, they displayed extensive fibrosis. Therefore the window for cardiac regeneration is brief in mammals, but there is a capacity to regenerate.
The purpose of the review article is to: "review our current understanding of how cardiomyocyte proliferation is regulated during heart development and regeneration". The primary article clearly relates to the topic of cardiomyocyte proliferation in the context of regeneration, which is why it has been used in the review article. The original research was used to provide evidence to support the statement: "cardiomyocytes in various contexts do harbor endogenous proliferative capabilities, albeit to different degrees". Correlations between the original research and the claims made by the review article include: "various contexts" - which in this case refers to the type of organism in which mice applies; and "endogenous proliferative capabilities" which from the first paragraph, the original research does support.
Lab 12
Lab Attendance
Z5020373 (talk) 13:08, 28 October 2016 (AEDT)
References
| Lab 10 - Stem Cell Presentations 2016 | |
|---|---|
| Group Mark | Assessor General Comments |
|
Group 1: 15/20 Group 2: 19/20 Group 3: 20/20 Group 4: 19/20 Group 5: 16/20 Group 6: 16/20 |
The students put great effort in their presentation and we heard a nice variety of studies in stem cell biology and regenerative medicine today. The interaction after the presentation was great.
As general feedback I would like to advise students to:
|
- ↑ <pubmed>24647352</pubmed>
- ↑ Converse, P.J. (2016) MUSCULAR DYSTROPHY, DUCHENNE TYPE; DMD OMIM http://www.omim.org/entry/310200
- ↑ <pubmed>11917091</pubmed>
- ↑ https://www.duchenneconnect.org/understanding-genetic-testing/types-of-mutations-in-the-dystrophin-gene.html
- ↑ <pubmed>12082140</pubmed>
- ↑ <pubmed>13947981</pubmed>
- ↑ <pubmed>26457695</pubmed>
- ↑ <pubmed>7683332</pubmed>
- ↑ <pubmed>25123483</pubmed>
- ↑ <pubmed>11917091</pubmed>
- ↑ http://www.fda.gov/NewsEvents/Newsroom/PressAnnouncements/ucm521263.htm