Zebrafish Development: Difference between revisions
From Embryology
No edit summary |
|||
| Line 8: | Line 8: | ||
== Some Recent Findings == | == Some Recent Findings == | ||
[[ | [[File:Nipbl heart and organ patterning.png|thumb|Nipbl heart and organ patterning<ref name="PMID22039349"><pubmed>22039349</pubmed></ref>]] | ||
File:Nipbl heart and organ patterning.png]] | |||
{| | {| | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
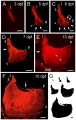
* '''Multifactorial Origins of Heart and Gut Defects in nipbl-Deficient Zebrafish, a Model of Cornelia de Lange Syndrome'''<ref><pubmed>22039349</pubmed></ref> "Cornelia de Lange Syndrome (CdLS) is the founding member of a class of multi-organ system birth defect syndromes termed cohesinopathies, named for the chromatin-associated protein complex cohesin, which mediates sister chromatid cohesion. Most cases of CdLS are caused by haploinsufficiency for Nipped-B-like (Nipbl), a highly conserved protein that facilitates cohesin loading. ... These findings support the view that birth defects in CdLS arise from collective effects of quantitative changes in gene expression. Interestingly, both the phenotypes and gene expression changes in nipbl morphants differed from those in mutants or morphants for genes encoding cohesin subunits, suggesting that the transcriptional functions of Nipbl cannot be ascribed simply to its role in cohesin loading. (OMIM - [http://omim.org/entry/122470 CDLS1] | [http://omim.org/entry/300590 CDLS2] | [http://omim.org/entry/610759 CDLS3]) | * '''Multifactorial Origins of Heart and Gut Defects in nipbl-Deficient Zebrafish, a Model of Cornelia de Lange Syndrome'''<ref name="PMID22039349"><pubmed>22039349</pubmed></ref> "Cornelia de Lange Syndrome (CdLS) is the founding member of a class of multi-organ system birth defect syndromes termed cohesinopathies, named for the chromatin-associated protein complex cohesin, which mediates sister chromatid cohesion. Most cases of CdLS are caused by haploinsufficiency for Nipped-B-like (Nipbl), a highly conserved protein that facilitates cohesin loading. ... These findings support the view that birth defects in CdLS arise from collective effects of quantitative changes in gene expression. Interestingly, both the phenotypes and gene expression changes in nipbl morphants differed from those in mutants or morphants for genes encoding cohesin subunits, suggesting that the transcriptional functions of Nipbl cannot be ascribed simply to its role in cohesin loading. (OMIM - [http://omim.org/entry/122470 CDLS1] | [http://omim.org/entry/300590 CDLS2] | [http://omim.org/entry/610759 CDLS3]) | ||
* '''The zebrafish transcriptome during early development'''<ref><pubmed>21609443</pubmed></ref> "The three earliest developmental stages were similar when comparing highly expressed genes, whereas the 50% epiboly stage differed from the other three stages in the identity of highly expressed genes, number of uniquely expressed genes and enrichment of GO molecular functions. Taken together, these observations indicate a major transition in gene regulation and transcriptional activity taking place between the 512-cell and 50% epiboly stages, in accordance with previous studies." | * '''The zebrafish transcriptome during early development'''<ref><pubmed>21609443</pubmed></ref> "The three earliest developmental stages were similar when comparing highly expressed genes, whereas the 50% epiboly stage differed from the other three stages in the identity of highly expressed genes, number of uniquely expressed genes and enrichment of GO molecular functions. Taken together, these observations indicate a major transition in gene regulation and transcriptional activity taking place between the 512-cell and 50% epiboly stages, in accordance with previous studies." | ||
Revision as of 12:38, 3 November 2011
Introduction
Zebrafish or zebra danio (danio rerio) are seen as the latest "model' for embryological development studies. These embryos have the great advantage that they develop as "see through" embryos, that is, all internal development can be clearly observed from the outside in the living embryo. Much of the early modern work using this embryo model began with the papers of Kimmel.[1][2]
Several large laboratories in the US are now developing large breeding programs to carry out "knockouts" and to find spontaneous mutants of interest.
| Fish Links: Zebrafish Development | Medaka Development | Salmon Development | Movie - Zebrafish Heart | Student Group Project - Zebrafish | Recent References | Category:Zebrafish | Category:Medaka |
Some Recent Findings
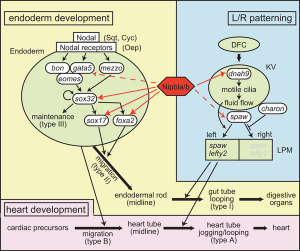
Nipbl heart and organ patterning[3]
|
Timeline and Stages of Embryonic Development
| Duration | Period Name | Image |
| 0 - 0.75 hrs | Zygote Period | 
|
| 0.75 - 2.25 hrs | Cleavage Period | 
|
| 2.25 - 5.25 hrs | Blastula Period | 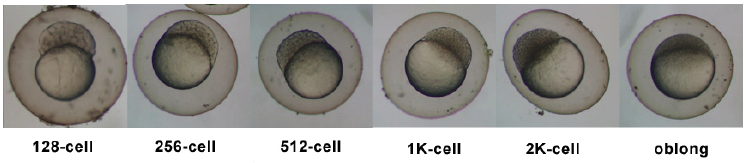
|
| 5.25 - 10.33 hrs | Gastrula Period | 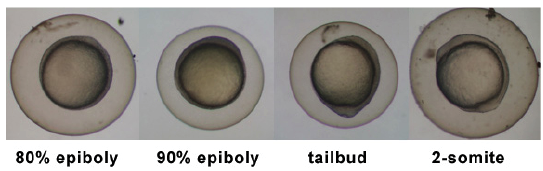
|
| 10.33 - 24 hrs | Segmentation Period | 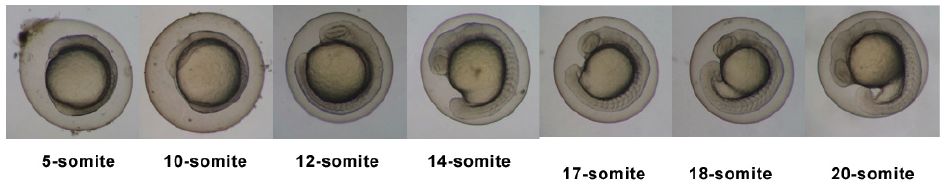
|
| 24 - 48 hrs | Pharyngula Period | 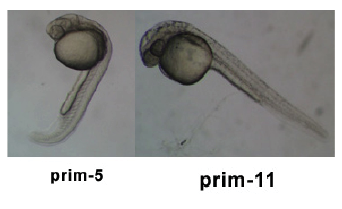
|
| 48-72 hrs | Hatching Period | 
|
| 72 hrs - 30 Days | Larval Period | 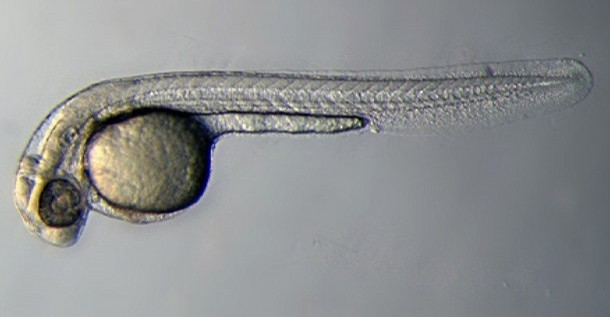
|
Molecular
Fibroblast Growth Factor
- Fgf8 and Fgf3 - regulating the segmentation of the pharyngeal endoderm into pouches. [7]
- Fgf24 and Fgf8 - promotes posterior mesodermal development.[8]
- Sox9 - required for cartilage morphogenesis.[9]
References
Reviews
Articles
Search Pubmed
Search Pubmed: Zebrafish Development
Additional Images
External Links
| Animal Development: axolotl | bat | cat | chicken | cow | dog | dolphin | echidna | fly | frog | goat | grasshopper | guinea pig | hamster | horse | kangaroo | koala | lizard | medaka | mouse | opossum | pig | platypus | rabbit | rat | salamander | sea squirt | sea urchin | sheep | worm | zebrafish | life cycles | development timetable | development models | K12 |
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 20) Embryology Zebrafish Development. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Zebrafish_Development
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G