User:Z5019880
| Student Information (expand to read) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Individual Assessments | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Please leave this template on top of your student page as I will add your assessment items here. Beginning your online work - Working Online in this course
Click here to email Dr Mark Hill | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 1 Assessment - Researching a Topic | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
In the lab I showed you how to find the PubMed reference database and search it using a topic word. Lab 1 assessment will be for you to use this to find a research reference on "fertilization" and write a brief summary of the main finding of the paper.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 2 Assessment - Uploading an Image | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
OK you are now in a group
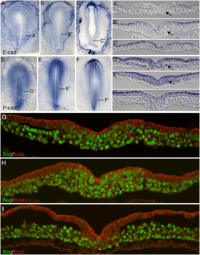
Initially the topic can be as specific or as broad as you want. Chicken embryo E-cad and P-cad gastrulation[1] References
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 4 Assessment - GIT Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
ANAT2341 Quiz Example | Category:Quiz | ANAT2341 Student 2015 Quiz Questions | Design 4 quiz questions based upon gastrointestinal tract. Add the quiz to your own page under Lab 4 assessment and provide a sub-sub-heading on the topic of the quiz. An example is shown below (open this page in view code or edit mode). Note that it is not just how you ask the question, but also how you explain the correct answer. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 5 Assessment - Course Review | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Complete the course review questionnaire and add the fact you have completed to your student page. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 6 Assessment - Cleft Lip and Palate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 7 Assessment - Muscular Dystrophy | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 8 Assessment - Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| A brief quiz was held in the practical class on urogenital development. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 9 Assessment - Peer Assessment | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 10 Assessment - Stem Cells | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
As part of the assessment for this course, you will give a 15 minutes journal club presentation in Lab 10. For this you will in your current student group discuss a recent (published after 2011) original research article (not a review!) on stem cell biology or technology.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 11 Assessment - Heart Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Read the following recent review article on heart repair and from the reference list identify a cited research article and write a brief summary of the paper's main findings. Then describe how the original research result was used in the review article.
<pubmed>26932668</pubmed>Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Lab Attendance
Z5019880 (talk) 14:34, 5 August 2016 (AEST)
Z5019880 (talk) 14:41, 12 August 2016 (AEST)
Z5019880 (talk) 14:01, 19 August 2016 (AEST)
Z5019880 (talk) 14:40, 26 August 2016 (AEST)
Z5019880 (talk) 14:27, 2 September 2016 (AEST)
Z5019880 (talk) 20:07, 11 September 2016 (AEST) (Forgot to input on Friday)
Z5019880 (talk) 13:14, 16 September 2016 (AEST)
Z5019880 (talk) 14:56, 23 September 2016 (AEST)
Z5019880 (talk) 15:11, 7 October 2016 (AEDT)
Z5019880 (talk) 00:04, 16 October 2016 (AEDT) (Lab 10 Speech, forgot to input on Friday)
Z5019880 (talk) 13:31, 21 October 2016 (AEDT)
Z5019880 (talk) 13:35, 28 October 2016 (AEDT)
Lab 1 Tasks
Type in group
I feel that the description of team worker best describes me with regards to trying to ensure that tasks ran as a team are ran smoothly. Particulalrly when it comes to differing ideas presented by group members, I try to ensure that I do my best to work in such away that is agreeable with all other group members. The fact that I find myself being indecisive when it comes to a choice between how an assessment should be done by group members or when appraising my own work and that of others, really makes me feel that I fit into the category of teamworker based on its description.
Lecture - Fertilization (Interesting points)
What I found most interesting in the lecture on fertilization was the removal of the extra DNA of the egg in female gametogenesis during meiosis 1 and 2. I find that the concept of division during these meiosis stages that lead to polar bodies which are asymmetrical to the egg (much smaller) to release extra DNA to ensure the egg at the end is haploid, rather than normal symmetrical division of cells is fascinating. From this interest I further researched as to how this asymmetrical division is determined by the cell, which is potentially as a result of formin - 2, a protein that can cause the placement of the meiotic spindles to be eccentric as it induces formation of actin filaments which pull chromosomes to its location [1]. It is this eccentric placement of the meiotic spindles that thus leads to an asymmetrical division.
Lab 1 Assessment
<pubmed>PMC4554382</pubmed> [2] Vitrification of an oocyte entails it rapidly being cooled for preservation, where it can later be used in in-vitro fertilization (IVF). This process relies cryoproctective agents (CPAs) to increase viscosity and decrease the freezing point of the liquid within an oocyte, which in turn prevents the formation of a solid via crystallisation which could potentially kill the cell. CPAs exert osmotic stress and a level of toxicity to the oocyte, which both can be modulated by the temperature of the fluid within an oocyte. It is known that protocols surrounding the procedure of vitrification have not been consistent regarding the temperature at which the oocytes are subjected to during the procedure. In Shanshan et al., varying temperatures during the dilution step of cryopreservation by vitrification were tested to optimize such a procedure with respect to survival rates of the preserved oocytes, and the success of the proceeding IVF.
In Shanshan et al. patients that were selected and sorted into groups based on whether the oocyte to be used in the IVF procedure was from a donor or non donor (autologus). All oocytes used were subjected to the same vitrification protocols up until warming of the cryopreserved oocyte, where such as step was split into two treatments groups, those that warmed at room temperature (20-22°C) or at 37°C. At this point survival rates of the oocytes were taken, and viable oocytes were inseminated and fertilisation was evaluated shortly after. The quality of the resulting embryos were graded, which formed the basis of which embryos were implanted, where successful implantation and progression to clinical pregnancy was measured. It was found from the result of Shanshan et al. that when comparing the treatments groups, warming the oocyte after vitrification at 37°C significantly increased the chances of the oocyte surviving when thawed only for the autologous or non-donor patient group. This was thought to be due to higher temperatures increasing the permeability of CPAs, which reduced the exposure of it to the oocyte during rehydration. The differences in treatment groups appeared to have no significant effect on all the other parameters measured for both patient groups. It is such that based on the findings of Shanshan et al., vitrified oocytes should be warmed during rehydration at 37°C to increase survival rates of the oocyte for non donors.
| Mark Hill 18 August 2016 - You have added the citation correctly and written a brief summary of the article findings. Well, the oocyte would normally be functioning in an in vivo environment of 37°C, does it surprise you that this turns out to be optimal rather than room temperature? Many biological processes would be expected to be optimal at this temperature. | Assessment 5/5 |
Lab 2 Assessment
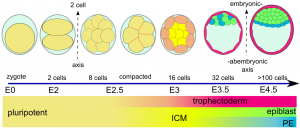
Morphological differences in early mouse embryonic development.
<pubmed>PMC3088645</pubmed>
| Mark Hill 29 August 2016 - All information Reference, Copyright and Student Image template correctly included with the file and referenced on your page here. Note though to display the reference citation correctly with the legend. You need to include the ref name for a citation, as shown below. I have also added <references/> code for refs to display on your page.
Code: <ref name="PMID21573197"><pubmed>21573197</pubmed></ref> Morphological differences in early mouse embryonic development.[3] |
Assessment 5/5 |
Lab 4 Assessment
Gastrointestinal development Quiz
| Mark Hill 17 October 2016 - GIT Quiz covers several topics. Question 1 is a little easy not testing knowledge. Question 2 and 3 are better. Question 4 could have a clearer question structure. Your answer explanations are also useful. | Assessment 5/5 |
Lab 5 Assessment
Completed the survey regarding ANAT2341
| Mark Hill 11 October 2016 - Questionnaire on course structure. | Assessment 5/5 |
Lab 6 Assessment
Interferon regulatory factor 6 (IRF6) gene mutations and its association with cleft lip/palate
<pubmed>15317890</pubmed>
Mutations in the gene encoding the transcription factor IRF6, leading to the gene becoming nonfunctional have been implicated to be associated with non-symptomatic cleft lip and/or palate[4]. The function of IRF6 signaling and its relation to the fusion event between the maxillary and frontonasal prominences but findings has not been well established as of yet, but early findings have shown that a loss of IRF6 in mice lead to a hyper-proliferation, improper terminal differentiation of the epidermal region in the oral cavity, and an absence of keratinizing epithelium. A combination of these factors cause oral adhesions, which block the oral cavity, leading to a disruption in the fusion of the tissues within the oral facial region, causing cleft lip and/or palate[5].
| Mark Hill 17 October 2016 - IRF6 is a factor identified by several students and you summary is useful. It would have been good to also include the signaling pathway involved. | Assessment 5/5 |
Lab 7 Assessment
Duchenne muscular dystrophy
Duchenne muscular dsytrophy (DMD), is an X-linked genetic disorder, where the gene encoding dystrophin is non functioning leading to a lack of or deficient production of dystrophin. Dystrophin is a structural protein which adds to the structural integrity of the skeletal muscle[6]. Without dystrophin the muscle undergoes repetitive cycles of damage and regeneration leading to a loss of function in the muscle due to the tissue being replaced by fibrous tissue. DMD has been estimated to occur in 1 in 3500 male births[7].
What is/are the dystrophin mutation(s)?
The main mutation regarding dystrophin occurs as deletions of the dystrophin gene on the Xp21 locus on the X chromosome, which leads to an inability to produce the protein dystrophin[8].
What is the function of dystrophin?
Dystrophin is a structural protein that anchors structures witihin the muscle fibers to the basement membrane of the endomysium, which overall provides structural integrity and stiffness of the muscle. [9]
What other tissues/organs are affected by this disorder?
DMD affects all tissues that contain and rely on dystrophin for structural integrity, which mainly includes skeletal[10] and cardiac muscle[11].
What therapies exist for DMD?
DMD affects the cardiac muscle leading to cardiacmyopathy, which is a major cause of death in DMD. It is such that research has been centered on prolonging the onset of cardiac myopathy, where studies have shown that the drug tadalafil, a phosphodiesterase type 5 inhibitor, when used prophylacticly can delay the onset of cardiac myopathy in mdx mouse and golden retriever muscular dystrophy models[12].
Lab 8 Assessment
| Mark Hill 27 October 2016 | Assessment 5/8 |
What animal models are available for muscular dystrophy?
With regards to how the disease is studied, animal models are employed which include the mdx mouse[13], which although emulates the disease, is not incredibly useful when it comes to gene therapy testing due to the small amount of muscle mass present relative to humans. It is thus that golden retriever muscular dystrophy models have been used as of late to study the disease and treatments[14].
| Mark Hill 27 October 2016 - Well done. | Assessment 6/8 |
Lab Assessment 9
These are good reviews of the project pages, with some specific examples. They include a balanced critical assessment, given the existing status of these pages. 8/10
Peer assessment
Group 1 Peer Assessment
The start you have made on your project appears to be quite decent. There seems to be a clear overview and scaffold of how your page will look and what it will discuss in the end. For the most part the usage of dot points has made understanding your points with regards to the signalling pathways (Canonical pathway section) a lot easier as opposed to having a wall of text. I would recommend possibly adopting dot points when explaining the pathway regarding the Wnt-Calcium Ion pathway to make it easier to digest. That being said though, there are areas within your wiki page that would most likely benefit from having complete paragraphs such as your sub sections labelled under the non-canonical pathway. It appears that each individual point in the sub section role appears to represent individual points that could be substantially elaborated on. In way I feel that it would make the ideas in the section less disjoint and more clear, given that writing in a paragraph format would be suitable for longer passages. Also for the part where there are there are research articles linked, and descriptions of such articles, it might be better to try integrate such ideas into other main components of your wiki page, because they seem quite out of context and out of nowhere. That being said you could also just put this under a current research heading and talk about it with respect to the current findings of the Wnt pathway.
Another main aspect that should be corrected is that in some sections, there is the assumption in your wiki page that the reader fully understands all your abbreviations. I know it sounds silly but it is probably best that your group coordinates or finds where you first use an abbreviation such as CaMKII in your non canonical pathway section and change it to the unabbreviated name, with the abbreviated name in brackets, where from there you can just use the abbreviated name. Also maybe just providing a glossary of the abbreviated terms and their unabbreviated terms at the end of your page will do as well. Also its good to keep in mind that you may have already done this for some terms, so look out for that as well.
With regards to your referencing, I see that it is quite extensive, but there seems to be a lack of in text citations. As a result, its quite hard for those who read your page to quickly find the appropriate citation with regards to the sentences or dot point being read. For the sections such as “Canonical Pathway: How it works” this isn’t too bad, as there is only one reference, but for the “Non-Canonical Pathway section” there are way too many for it to be easy to tell where the citations are associated to. So overall for this I recommend your group to use in-text citations. Also I’ve noticed that you have used a review to cite your whole “Canonical pathway: How it works” section, which for the most part most likely contains all your information you have stated, but doesn’t give credit to the specific or individual authors included in the review and also requires the reader to go and find the specific sections in the review that you have used to cite your text. It is such that it would be better to use research articles to site your individual points, maybe extracting such research articles from the review article itself.
Overall the start made on your project is appearing to take shape, where I see that there are many subheadings yet to be filled below the “Wnt-Calcium Ion pathway” section. I’m sure if your groups keep up the quality of the work, your page should turn out fine with the addition of incorporating the feedback I have provided.
Group 2 Peer Assessment
Your project is quite good and seems to be on the right track. All your references have been done in-text and have made it really easy to make one’s way to the research article to read more about certain points. Not only that, you have appropriately abbreviated your terms by using the full name initially, and I can see that you have a glossary section which should be beneficial in the future when more terms are added. Your history section is well presented but, it might be important to add references to the papers of the main points of discovery in your history section as to allow people to easily access and find the full article regarding the discovery.
The fact that you have added pictures is quite handy when it comes to using it as an aid to accompanying passage. With regards to the image legend, maybe add more information to it or transfer the description of the image present when clicking into the image onto the legend as to better represent what the image is about while having the passage right next to it. Furthermore, maybe it would be beneficial to add other forms of media such as videos to compliment the passages as well, and help better engage the reader in the topic.
With regards to your section on the canonical pathway, I’ve noticed that the specific genes that are targeted by Notch have been left out and I feel that it is important to mention those genes targets explicitly there as well. That being said, they are mentioned in the proceeding section so it isn’t imperative that you do this. Maybe also try seeing if there is any literature on how the NOTCH receptors come about, such as what genes transcribe it and how the protein is processed and expressed before signalling in the pathway can occur.
I think for the most part there is very little to improve with your wiki page given the quality of it albeit a few minor corrections that I have mentioned above. It is very concise and at no times do I feel that I am reading a wall of text that is disengaging. Thus I feel that as long as such quality is maintained then your wiki page will be quite good when finished.
Group 3 Peer Assessment
With regards to your project I have noticed there are many forms of educational tools employed or being planned other than text, which to me is a big plus with regards to your project. The usage of the table to summarises the different FGFR sub-types is really easy to read and understand, and presents the information in a better way than you could’ve with just a wall of text. Your planned multiple choice section seems like it would be a nice addition to your page where it should help solidify the knowledge of the reader, allowing to check what they know. When doing the quiz section not only would it be good if you added explanations for the correct answers, but maybe also if possible explanations of why the other answers are wrong. There seems to be no issues with your citations given that all of them are in-text and multiple. Also the link between signal transduction, embryonic development and abnormalities is quite smooth and within context of their respective preceding parts, making the page read very well.
With regards to your usage of images, it seems mostly good and compliments the passages well, but I feel that it would benefit with adding more information to the legend, possibly by moving some of the description when clicking into the image into the legend. Also since your first image contains mainly abbreviations, maybe it would be good to collate all abbreviations and add it to the glossary such that the reader can easily refer to what the abbreviations mean.
With respect to your signal transduction section, all the components of the pathway seem to have been included, but for the most part how each factor interacts with one another has been left out. Elaborating on how each factor interacts and activates one another such as how FRS2 recruits GRB2 and SHP2, and how those events actually promote activation of RAS. I feel adding this will really improve the depth of this section, and make it less about a bunch of different components and more about how the work together in the context of their individual functions. Also I feel that the history section could be expanded on, maybe to include more time points or critical areas of discovery for the FGFR pathway.
Overall I think your project is shaping up quite well, and that with the addition of the suggestions made above, would make your project quite good. Having used many images, a table, and including the quiz has really made your page quite interactive and engaging which has really benefited your page. Also your subheadings and included passages have appeared to cover most important topics within your signalling pathway.
Group 5 Peer Assessment
It is quite clear that what has been provided in your wiki page is extensive and well researched. The inclusion of tables summarizing the different T-box genes although extensive, is very concise and easy to read. I feel that this table really links all the elements of your page together, where you have included its function and related it to embryological development and abnormalities which you go on later to elaborate in other sections. I feel this really complements the introduction and gives a good feel for what’s to come in the rest of the page. The addition of what the term T-box means also is a nice touch, giving context and some history regarding the name.
Your origins section of T-box is quite well outlined, but as mentioned in your page, having a timeline with critical points of discovery with regards to the genes would probably be more beneficial as it would be a lot easier to read a see the time points as a whole. That being said, having the timeline alongside your outline would probably work well, as your outline can serve to elaborate on the timeline. With regards to your subheadings, it seems to they are quite extensive and cover practically all the key components of the T-box genes, and it is also good to see that there is a glossary subheading in place. Content wise there seems to be limited to no issues, but with regards to abbreviations, I have found that the usage hasn’t always been after the fact of providing the full name first. For example, bone morphogenic protein’s abbreviation is used consistently throughout the first part of the wiki page, but it is only described by its full name and then abbreviation later on. This is something you should check out and fix by either adding the full name the first time the abbreviation is used, or adding all these terms to the glossary.
With regards to the pictures they all seem to compliment the sections well and are quite plentiful. That being said though the picture in the “Marsupial forelimb development” does not appear to have the copyright information regarding to its usage, and referencing does not appear to be in full. This is also the same for the picture under the subheading “Organisms used in animal models for T-box”. Other than that the referencing is perfectly fine within the text.
Overall this project is really good and without any major flaws when it comes to the content. A few touch ups here and there with regards to my suggestion above, and your project should be good to go along as the quality is kept at this level.
Group 6 Peer Assessment
In this project page a good start has been made with the inclusion of images to compliment the signalling pathway description. Appropriate abbreviations appear to be used, where the full name is used first. Also the additions of a glossary and further reading subheading is a nice touch, which should allow for better understanding of the topic should the reader want more information. In terms of the diversity of the subheadings though, it seems that a lot more could be added, such as animal models used to research the signalling pathway and also possibly abnormalities that may arise from the errors or mutations in the pathway. It is probably wise to also add a section regarding embryological development and what role TGF beta signalling pathway has in it, which should help provide context to the abnormalities section when added.
The usage of pictures is appropriate for the section it has been put in, and compliments the signal transduction pathway description well, but the picture labelled “Process of TGF-beta signalling pathway” does not appear to be referenced or have the appropriate copyright under it. There is also no legend for this picture to briefly describe it. Also for most of the page there are limited to no references, where for the signalling pathway section, it appears your groups has used websites rather than peer reviewed articles as a source. The websites are probably good starting points to get a general idea of the pathway, but it is probably better if you find peer reviewed papers to cite, which potentially the websites you have used have cited. Also when citing it is best to use in text citation such that the reader can easily see which paper you are referring to when describing certain facts.
Also in your groups signalling section, it is mentioned that SMAD when activated, recruits various transcriptional regulators that control expression of numerous genes. This is quite vague and is probably a good idea to mention some of these factors, and also what genes they regulate and the importance of such genes. Doing this should also provide your group with a good link to the embryological development section with regards to TGF-beta signalling.
It still seems that overall there is a lot of work to be done on your groups page, but a good start and effort has been made to include various images and also subheadings. I feel that if your group incorporates some of the suggested subheadings described above, and other feedback mentioned, the page should be greatly improved.
Lab Assessment 11
Assessment - 5/5
Paper cited in review article
<pubmed>26256209</pubmed>
In this research article, the authors looked at how innervation of the heart in mice and zebra fish models affected the regeneration and proliferation of the cardiac myocytes after damage. More specifically they looked at how reduction of the innervation affected the regeneration whether it was via direct mechanical denervation or though the inhibition of cholinergic nerve function. In this case of cholinergic nerve inhibition, transgenic zebra fish were used, which over expressed semaphorin3aa in the myocardium. This lead to reduced ventricular innervation, which in turn upon immunohistological examination, where makers for Mef2 (staining cardiomyocyte nuclei) and PCNA (marker for proliferation) were used, showed a significant reduction in cardiomyocyte proliferation over zebra fish without over expression of semaphorin3aa. Further studies performed used pharmacological drugs such as atropine and methoctramine, which were both cholinergic inhibitors on neonatal mice and zebrafish hearts, were similar staining from the above experiement was used which identfied reduced proliferation in the treatment group over the control. Beta adrenergic receptors were also blocked with the adrenergic inhibitor, propanolol, which showed an increase in proliferation of cardiomyocytes over the control, suggesting that cholirnergic nerve transmission specifically played a role in the regenerative potential of the heart.
Mechanical denervation in neonatal mice was performed via cutting the left vagus nerve, which removed cholinergic innervation to the right side of the heart predominately. A myocardial infarction was then induced in the neonatal mice, and the heart was removed for immunohistological examination. via staining of phosphorylated H3, which is a marker for cardiac myocyte proliferation. No significant results were found when comparing the treatments to the control in this subset of the experiments. Furthermore experiments on mechanical denervation were continued with trying to increase proliferation by adding nerve growth factors to the tissue such as neurgrelin 1 (NRG1) and nerve growth factor (NGF). Measurments were taken as the level of DNA synthesis in the cells yousing H-thymidine incorporation, and showed that the usage of NRG1 increased cardiomyocyte DNA synthesis, unlike NGF. Finally results looking at the immune function in resected hearts of zebra fish showed that resection of hearts that had undergone vagotomy had a blunted immune response (based on looking at immune factors such as Cxcl5 and IL1b)as compared to ones that had not undergone this removal of the vagus nerve, implying a immune a possible immune component to heart repair[15].
Context in review article
5/5
<pubmed>26932668</pubmed>
Te review article is touches on the research that had been done previously regarding our understanding in the regulation of cardiomyocyte proliferation in response to injury, in the context of the fact that adult cardiomyocytes have poor proliferative capabilities when compared to those in the developing fetus and neonates. As a result the review has looked at many factors that may contribute to such regulation of regeneration, including that of non muscle type cells in aiding proliferation. One of these type happened to be nerve cells innervating the heart which given the results of the paper mentioned above showed it played a role in increasing the proliferative capabilities of the heart. These result further fit into the paper as they used neonatal mice as one of their models of study, which is in line with the review article context when it came to neonates having greater proliferatieve capability. Overall this paper added context regarding nerve cell innervation of the heart to proliferation of the cardiomyocytes to the review alongside a potential reason of why neonates have possible better proliferative capabilities (greater innervation of the hear possibly). This overall built upon the overarching topic regarding regeneration of cardiomyocytes and presented consideration of nerve innervation in aiding such proliferation, and also used the article to show the breadth of animal models used in this research area[16].
References
- ↑ <pubmed>12461532</pubmed>
- ↑ <pubmed>25956261</pubmed>
- ↑ <pubmed>21573197</pubmed>
- ↑ <pubmed>15317890</pubmed>
- ↑ <pubmed>17041603</pubmed>
- ↑ <pubmed>3042151</pubmed>
- ↑ <pubmed>1822774</pubmed>
- ↑ <pubmed>3607877</pubmed>
- ↑ <pubmed>3042151</pubmed>
- ↑ <pubmed>2693617</pubmed>
- ↑ <pubmed>27506543</pubmed>
- ↑ <pubmed>27506543</pubmed>
- ↑ <pubmed>344791</pubmed>
- ↑ <pubmed>3290691</pubmed>
- ↑ <pubmed>26256209</pubmed>
- ↑ <pubmed>26932668</pubmed>
| Mark Hill (talk) 12:36, 5 August 2016 (AEST) Very good. maybe a little odd in page formatting this can get messy. I will discuss in today's lab online formatting etc. |
Lab 3 Assessment
| Mark Hill 31 August 2016 - Lab 3 Assessment Quiz - Mesoderm and Ectoderm development. | Assessment 3.5/5 |
| Lab 10 - Stem Cell Presentations 2016 | |
|---|---|
| Group Mark | Assessor General Comments |
|
Group 1: 15/20 Group 2: 19/20 Group 3: 20/20 Group 4: 19/20 Group 5: 16/20 Group 6: 16/20 |
The students put great effort in their presentation and we heard a nice variety of studies in stem cell biology and regenerative medicine today. The interaction after the presentation was great.
As general feedback I would like to advise students to:
|