Molecular Development - microRNA: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 15: | Line 15: | ||
* '''A survey of small RNAs in human sperm'''<ref><pubmed>21989093</pubmed></ref> "Bioinformatic analysis revealed the presence of multiple classes of small RNAs in human spermatozoa. The primary classes resolved included microRNA (miRNAs) (≈7%), Piwi-interacting piRNAs (≈17%), repeat-associated small RNAs (≈65%). A minor subset of short RNAs within the transcription start site/promoter fraction (≈11%) frames the histone promoter-associated regions enriched in genes of early embryonic development. These have been termed quiescent RNAs. CONCLUSIONS A complex population of male derived sncRNAs that are available for delivery upon fertilization was revealed. Sperm miRNA-targeted enrichment in the human oocyte is consistent with their role as modifiers of early post-fertilization. The relative abundance of piRNAs and repeat-associated RNAs suggests that they may assume a role in confrontation and consolidation. This may ensure the compatibility of the genomes at fertilisation." | * '''A survey of small RNAs in human sperm'''<ref><pubmed>21989093</pubmed></ref> "Bioinformatic analysis revealed the presence of multiple classes of small RNAs in human spermatozoa. The primary classes resolved included microRNA (miRNAs) (≈7%), Piwi-interacting piRNAs (≈17%), repeat-associated small RNAs (≈65%). A minor subset of short RNAs within the transcription start site/promoter fraction (≈11%) frames the histone promoter-associated regions enriched in genes of early embryonic development. These have been termed quiescent RNAs. CONCLUSIONS A complex population of male derived sncRNAs that are available for delivery upon fertilization was revealed. Sperm miRNA-targeted enrichment in the human oocyte is consistent with their role as modifiers of early post-fertilization. The relative abundance of piRNAs and repeat-associated RNAs suggests that they may assume a role in confrontation and consolidation. This may ensure the compatibility of the genomes at fertilisation." | ||
|} | |} | ||
{| class="wikitable collapsible collapsed" | {| class="wikitable mw-collapsible mw-collapsed" | ||
! More recent papers | ! More recent papers | ||
|- | |- | ||
| Line 24: | Line 24: | ||
<pubmed limit=5>microRNA Embryology</pubmed> | <pubmed limit=5>microRNA Embryology</pubmed> | ||
|} | |} | ||
==Functional miRNA Grouping== | ==Functional miRNA Grouping== | ||
The following identifies microRNAs that have been identified with specific developmental processes, based upon a commercial collation of basic research data. | The following identifies microRNAs that have been identified with specific developmental processes, based upon a commercial collation of basic research data. | ||
* Pluripotency | |||
* let-7a, let-7b, let-7c, let-7d, let-7e, let-7g, miR-101, miR-106b, miR-125b, miR-130a, miR-133b, miR-141, miR-15a, miR-17, miR-182, miR-183, miR-18a, miR-18b, miR-205, miR-20a, miR-20b, miR-21, miR-214, miR-22, miR-222, miR-23b, miR-24, miR-302a, miR-302c, miR-345, miR-424, miR-498, miR-518b, miR-520g. | ** let-7a, let-7b, let-7c, let-7d, let-7e, let-7g, miR-101, miR-106b, miR-125b, miR-130a, miR-133b, miR-141, miR-15a, miR-17, miR-182, miR-183, miR-18a, miR-18b, miR-205, miR-20a, miR-20b, miR-21, miR-214, miR-22, miR-222, miR-23b, miR-24, miR-302a, miR-302c, miR-345, miR-424, miR-498, miR-518b, miR-520g. | ||
===Early Development=== | ===Early Development=== | ||
* Embryoid Bodies | * Embryoid Bodies | ||
* Definitive Endoderm | ** miR-132, miR-181a, miR-9. | ||
* [[Endoderm|Definitive Endoderm]] | |||
** miR-205, miR-375. | |||
===Ectoderm=== | ===Ectoderm=== | ||
* | * [[Neural System Development|Neural Development]] | ||
* Eye Development | ** let-7b, miR-103a, miR-106b, miR-10b, miR-124, miR-125b, miR-130a, miR-132, miR-134, miR-137, miR-16, miR-181a, miR-182, miR-183, miR-20a, miR-210, miR-219-5p, miR-22, miR-23b, miR-24, miR-26a, miR-302a, miR-302c, miR-7, miR-9, miR-96. | ||
* Epidermal Differentiation | * Eye Development | ||
* Inner Ear Development | ** miR-130a, miR-196a, miR-219-5p, miR-23b, miR-96. | ||
* [[Integumentary System Development|Epidermal Differentiation]] | |||
** let-7b, miR-205, miR-210, miR-23b, miR-26a. | |||
* Inner Ear Development | |||
** miR-182, miR-183, miR-96. | |||
===Mesoderm=== | ===Mesoderm=== | ||
* | * Haematopoiesis | ||
* T Cell Development | ** let-7e, miR-125a-5p, miR-142-3p, miR-223. | ||
* Erythropoiesis | * T Cell Development | ||
* Lymphopoiesis | ** let-7a, let-7f, miR-106b, miR-142-5p, miR-146b-5p, miR-150, miR-15a, miR-15b, miR-16, miR-181a, miR-20a, miR-222, miR-26a. | ||
* Megakaryopoiesis | * Erythropoiesis | ||
* Monocyte Differentiation | ** let-7a, let-7b, let-7c, let-7d, let-7f, let-7g, let-7i, miR-126, miR-128a, miR-137, miR-155, miR-15b, miR-16, miR-17, miR-181a, miR-182, miR-185, miR-206, miR-21, miR-22, miR-222, miR-24, miR-26a, miR-96. | ||
* Myelopoiesis | * Lymphopoiesis | ||
* Angiogenesis | ** let-7b, miR-125b, miR-128a, miR-16, miR-181a, miR-21, miR-24. | ||
* Myogenesis | * Megakaryopoiesis | ||
* Osteogenesis | ** miR-106b, miR-10a, miR-10b, miR-122, miR-126, miR-127-5p, miR-129-5p, miR-134, miR-146a, miR-150, miR-155, miR-17, miR-18a, miR-192, miR-20a, miR-20b, miR-21, miR-22, miR-301a, miR-33a, miR-378, miR-92a, miR-93. | ||
* Adipogenesis | * Monocyte Differentiation | ||
** miR-155, miR-222, miR-424. | |||
* Myelopoiesis | |||
** miR-103a, miR-128a, miR-17, miR-181a, miR-24. | |||
* Angiogenesis | |||
** miR-126, miR-130a, miR-218, miR-222, miR-92a. | |||
* Myogenesis | |||
** miR-1, miR-125b, miR-206, miR-26a. | |||
* Osteogenesis | |||
** miR-141, miR-15b, miR-424. | |||
* Adipogenesis | |||
** let-7b, let-7c, let-7e, miR-100, miR-101, miR-103a, miR-10b, miR-146b-5p, miR-155, miR-182, miR-192, miR-194, miR-196a, miR-21, miR-210, miR-214, miR-22, miR-24, miR-498, miR-96, miR-99a. | |||
* Chondrogenesis: let-7f, miR-1, miR-132, miR-181a, miR-196a, miR-96, miR-99a. | * Chondrogenesis: let-7f, miR-1, miR-132, miR-181a, miR-196a, miR-96, miR-99a. | ||
* Heart Development: miR-1, miR-208, miR-488. | * Heart Development: miR-1, miR-208, miR-488. | ||
===Endoderm=== | ===Endoderm=== | ||
* Liver Development | * Liver Development | ||
* Pancreatic Development | ** let-7a, let-7b, let-7c, miR-10a, miR-122, miR-125b, miR-192, miR-21, miR-22, miR-23b, miR-92a, miR-99a. | ||
* Intestinal Development | * Pancreatic Development | ||
** miR-15a, miR-15b, miR-16, miR-195, miR-214, miR-375, miR-7, miR-9. | |||
* Intestinal Development | |||
** let-7d, let-7e, miR-103a, miR-106b, miR-125b, miR-126, miR-130a, miR-141, miR-146b-5p, miR-17, miR-192, miR-194, miR-21, miR-215, miR-301a, miR-424. | |||
Data: SABiosciences [http://www.sabiosciences.com/mirna_pcr_product/HTML/MIHS-103Z.html Cell Differentiation & Development miRNA PCR Array] | :'''Data:''' SABiosciences [http://www.sabiosciences.com/mirna_pcr_product/HTML/MIHS-103Z.html Cell Differentiation & Development miRNA PCR Array] | ||
==References== | ==References== | ||
Revision as of 09:56, 12 June 2015
| Embryology - 24 Apr 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
Micro RNA (miRNA) are 22 nucleotide non-coding RNAs. These small pieces of RNA have been identified as important negative regulators in both development and adult cell processes involving gene expression.
Some Recent Findings
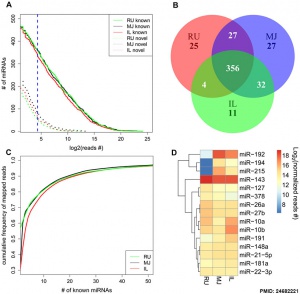
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: microRNA Embryology <pubmed limit=5>microRNA Embryology</pubmed> |
Functional miRNA Grouping
The following identifies microRNAs that have been identified with specific developmental processes, based upon a commercial collation of basic research data.
- Pluripotency
- let-7a, let-7b, let-7c, let-7d, let-7e, let-7g, miR-101, miR-106b, miR-125b, miR-130a, miR-133b, miR-141, miR-15a, miR-17, miR-182, miR-183, miR-18a, miR-18b, miR-205, miR-20a, miR-20b, miR-21, miR-214, miR-22, miR-222, miR-23b, miR-24, miR-302a, miR-302c, miR-345, miR-424, miR-498, miR-518b, miR-520g.
Early Development
- Embryoid Bodies
- miR-132, miR-181a, miR-9.
- Definitive Endoderm
- miR-205, miR-375.
Ectoderm
- Neural Development
- let-7b, miR-103a, miR-106b, miR-10b, miR-124, miR-125b, miR-130a, miR-132, miR-134, miR-137, miR-16, miR-181a, miR-182, miR-183, miR-20a, miR-210, miR-219-5p, miR-22, miR-23b, miR-24, miR-26a, miR-302a, miR-302c, miR-7, miR-9, miR-96.
- Eye Development
- miR-130a, miR-196a, miR-219-5p, miR-23b, miR-96.
- Epidermal Differentiation
- let-7b, miR-205, miR-210, miR-23b, miR-26a.
- Inner Ear Development
- miR-182, miR-183, miR-96.
Mesoderm
- Haematopoiesis
- let-7e, miR-125a-5p, miR-142-3p, miR-223.
- T Cell Development
- let-7a, let-7f, miR-106b, miR-142-5p, miR-146b-5p, miR-150, miR-15a, miR-15b, miR-16, miR-181a, miR-20a, miR-222, miR-26a.
- Erythropoiesis
- let-7a, let-7b, let-7c, let-7d, let-7f, let-7g, let-7i, miR-126, miR-128a, miR-137, miR-155, miR-15b, miR-16, miR-17, miR-181a, miR-182, miR-185, miR-206, miR-21, miR-22, miR-222, miR-24, miR-26a, miR-96.
- Lymphopoiesis
- let-7b, miR-125b, miR-128a, miR-16, miR-181a, miR-21, miR-24.
- Megakaryopoiesis
- miR-106b, miR-10a, miR-10b, miR-122, miR-126, miR-127-5p, miR-129-5p, miR-134, miR-146a, miR-150, miR-155, miR-17, miR-18a, miR-192, miR-20a, miR-20b, miR-21, miR-22, miR-301a, miR-33a, miR-378, miR-92a, miR-93.
- Monocyte Differentiation
- miR-155, miR-222, miR-424.
- Myelopoiesis
- miR-103a, miR-128a, miR-17, miR-181a, miR-24.
- Angiogenesis
- miR-126, miR-130a, miR-218, miR-222, miR-92a.
- Myogenesis
- miR-1, miR-125b, miR-206, miR-26a.
- Osteogenesis
- miR-141, miR-15b, miR-424.
- Adipogenesis
- let-7b, let-7c, let-7e, miR-100, miR-101, miR-103a, miR-10b, miR-146b-5p, miR-155, miR-182, miR-192, miR-194, miR-196a, miR-21, miR-210, miR-214, miR-22, miR-24, miR-498, miR-96, miR-99a.
- Chondrogenesis: let-7f, miR-1, miR-132, miR-181a, miR-196a, miR-96, miR-99a.
- Heart Development: miR-1, miR-208, miR-488.
Endoderm
- Liver Development
- let-7a, let-7b, let-7c, miR-10a, miR-122, miR-125b, miR-192, miR-21, miR-22, miR-23b, miR-92a, miR-99a.
- Pancreatic Development
- miR-15a, miR-15b, miR-16, miR-195, miR-214, miR-375, miR-7, miR-9.
- Intestinal Development
- let-7d, let-7e, miR-103a, miR-106b, miR-125b, miR-126, miR-130a, miR-141, miR-146b-5p, miR-17, miR-192, miR-194, miR-21, miR-215, miR-301a, miR-424.
- Data: SABiosciences Cell Differentiation & Development miRNA PCR Array
References
Reviews
<pubmed>21850044</pubmed> <pubmed>21742789</pubmed> <pubmed>21576351</pubmed> <pubmed>21504869</pubmed> <pubmed>21486922</pubmed> <pubmed>19148191</pubmed>
Articles
<pubmed></pubmed> <pubmed></pubmed> <pubmed>23617334</pubmed> <pubmed>20520778</pubmed> <pubmed>18029362</pubmed>| Nucleic Acids Res. <pubmed></pubmed>
Search Pubmed
Search Pubmed Now: microRNA
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- Memorial Sloan-Kettering Cancer Center - microRNA.org - Targets and Expression
- SABiosciences - Cell Differentiation & Development miRNA PCR Array
- Sigma-Aldrich - miRNA Introduction
- miRNAMap - http://mirnamap.mbc.nctu.edu.tw/
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 24) Embryology Molecular Development - microRNA. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Molecular_Development_-_microRNA
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
