Molecular Development - Ribonucleic acid
| Embryology - 23 Apr 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
The original "RNA family" consisted of just 3 main members; transfer RNA (tRNA), ribosomal RNA (rRNA) and messenger RNAs (mRNA). Involved in gene expression through protein translation (synthesis).
In recent years this RNA family, shown in the table below, has been expanded to include newly identified members: small nuclear RNA (snRNA) , small nucleolar RNA (snoRNA), and short regulatory RNA (piwi-associated RNA (piRNA), endogenous short-interfering RNA (endo-siRNA) and microRNA (miRNA) and now long non-coding RNA (lncRNA). Each of these new family members has a range of potential roles in development and differentiation.
Ribonucleic acid (RNA) typically consists of just 4 nucleotides: Adenine (A), Cytosine (C), Guanine (G) and Uracil (U). Note that Uracil (U) replaces Thymine (T) present in DNA. RNA is also less stable than DNA and exists as a single-stranded molecule, with secondary structure established by internal base pairing. The secondary structures often described as forming "hair-pins", "loops" and "stem-loop" structures, have a close relationship with many of the functions of these molecules.
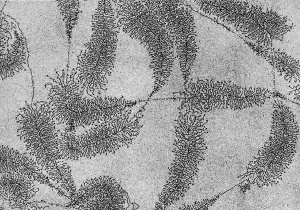
In some species, for example the fly and frog[2], it has been shown that oocyte maternal RNA and its specific localisation has an important role in early development.
- Links: microRNA | Fly Development | Frog Development
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Ribonucleic acid Embryology <pubmed limit=5>Ribonucleic acid Embryology</pubmed> |
RNA Classes
| RNA Class | Acronym | Roles | Examples | Reviews |
|---|---|---|---|---|
| messenger RNA | mRNA | transcribed from DNA in cell nucleus and relocate to cytoplasm for translation on the ribosome | Example | |
| ribosomal RNA | rRNA | structural RNA that allow the assembly of ribosomal proteins in the cytoplasm into ribosomal subunits required for protein translation | Example | |
| transfer RNA | tRNA | carry amino acid in cell cytoplasm to ribosome | PMID 24966867 | |
| small nuclear RNA | snRNA | involved in cell nucleus RNA splicing | assemble with proteins to form small nuclear ribonucleic particles (snRNPs) | PMID 23980890 |
| small nucleolar RNA | snoRNA | modify other small RNAs (rRNAs and tRNAs) | box C/D snoRNAs and the box H/ACA | PMID 21664409 PMID 19446021 |
| microRNA | miRNA | post-transcriptional regulator of gene expression | PMID 25128264 | |
| endogenous short-interfering RNA | endo-siRNA | short regulatory RNA | mainly characterised in plants | PMID 22578318 |
| piwi-associated RNA | piRNA | short regulatory RNA | PMID 18032451 PMID 22103557 | |
| long non-coding RNA | ncRNA | non-coding RNA greater than 200bp in length may have different roles in signalling, protein processing and differentiation | PMID 24829860 |
messenger RNA
(mRNA) RNA family of varying lengths transcribed from DNA in cell nucleus and relocate to cytoplasm for translation on the ribosome.
- The initially transcribed mRNA can be further processed (spliced) with the nucleus to remove introns (non-coding sequence) from between exons (the coding sequence).
- intron-exon-intron-exon -> exon-exon
- Additional non-coding regions are found at either ends of the messenger RNA as untranslated regions (UTRs).
- 5' UTR - encoded region - 3' UTR
ribosomal RNA
(rRNA) Structural RNA that allow the assembly of ribosomal proteins in the cytoplasm into ribosomal subunits required for protein translation.
transfer RNA
(tRNA) RNA family involved with specifically carrying and transferring an amino acid in the cell cytoplasm to ribosome, may have other regulatory roles.
PMID 24966867
small nuclear RNA
(snRNA) RNA family involved in cell nucleus RNA splicing assemble and along with proteins form small nuclear ribonucleic particles (snRNPs).
PMID 23980890
small nucleolar RNA
(snoRNA) This family of RNAs modify other small RNAs (rRNAs and tRNAs) and fall into 2 main classes; box C/D snoRNAs and the box H/ACA.
PMID 21664409 PMID 19446021
microRNA
(miRNA ) Large family of short RNAs involved in post-transcriptional regulators of gene expression.
- short hairpin RNA (shRNA) has a role in RNA interference (RNAi)
- RNA interference (RNAi) is post-transcriptional gene silencing by means of sequence-specific mRNA degradation.
- Note that RNAi has now been developed as a research technique to specifically carry out gene silencing in cells, tissues and embryos.
PMID 25128264
endogenous short-interfering RNA
(endo-siRNA) A family of short regulatory RNA mainly characterised in plants.
- see also short hairpin RNA (shRNA).
PMID 22578318 ===piwi-associated RNA (piRNA) short regulatory RNA PMID 18032451 PMID 22103557
long non-coding RNA
(ncRNA) Family of non-coding RNA greater than 200bp in length may have different roles in signalling, protein processing and differentiation.
PMID 24829860
- Links: Molecular Development
Epigenetics
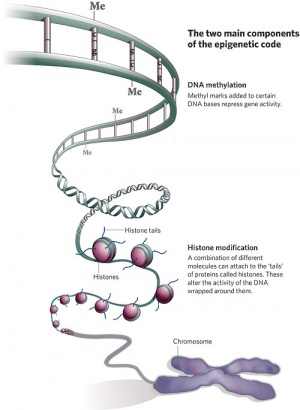
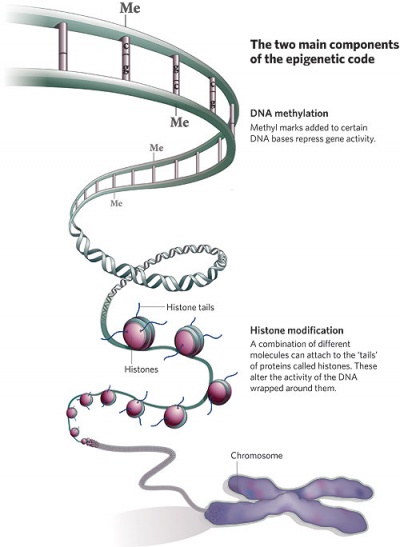
| Epigenetics as the name implies, is the inheritance mechanisms that lie outside the DNA sequence of our genes.
One of the initial discoveries was the effects of DNA methylation upon gene expression and then modifications of nucleosomal histones. This DNA methylation, usually associated with 5-methylcytosine (m5C), leads to transcriptional silencing in vertebrates.
|
|
References
Images
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- NHGR- The ENCODE Project ENCyclopedia Of DNA Elements
- Foldit - Amino Acids
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 23) Embryology Molecular Development - Ribonucleic acid. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Molecular_Development_-_Ribonucleic_acid
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G