|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
- This search now requires a manual link as the original PubMed extension has been disabled.
- The displayed list of references do not reflect any editorial selection of material based on content or relevance.
- References also appear on this list based upon the date of the actual page viewing.
References listed on the rest of the content page and the associated discussion page (listed under the publication year sub-headings) do include some editorial selection based upon both relevance and availability.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References
Search term: Developmental Genetics | Embryology Genetics | Mitochondrial Genetics | trisomy
| class="wikitable mw-collapsible mw-collapsed"
|
Older papers
|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
|
Diploid Chromosome Number
| Species |
Chromosome Number
|
| Human (Homo sapiens)
|
46
|
| Mouse (Mus musculus) |
40
|
| Fruit fly (Drosophila melanogaster) |
8
|
| Worm (Caenorhabditis elegans) |
12
|
| Frog (Xenopus laevis) |
36
|
| Dog (Canis familiaris) |
78
|
| Chicken (Gallus gallus) |
78
|
| Echidna (Tachyglossus) |
63 male 64 female
|
| Cow (Bos primigenius) |
60
|
| Species Chromosome Number
|
| Species |
Chromosome Number
|
| Human (Homo sapiens)
|
46
|
| Mouse (Mus musculus) |
40
|
| Fruit fly (Drosophila melanogaster) |
8
|
| Worm (Caenorhabditis elegans) |
12
|
| Frog (Xenopus laevis) |
36
|
| Dog (Canis familiaris) |
78
|
| Chicken (Gallus gallus) |
78
|
| Echidna (Tachyglossus) |
63 male 64 female
|
| Cow (Bos primigenius) |
60
|
|
|
|
| Links: Genetics
|
Chromatin Structure
| Cells not undergoing cell division have their DNA dispersed throughout the nucleus in what are know as "chromosomal domains". There is also a peripheral nuclear rim area that does not contain gene-rich regions of DNA, which tend to be located in the core of the nucleus.
|
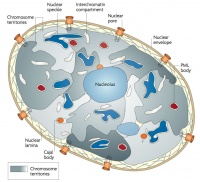 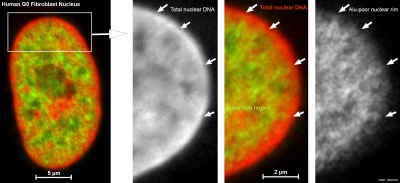
Adult G0 fibroblast DNA and gene localization.[3]
|
Cells undergoing division have their DNA compacted into chromosomes with a short arm (p), a long arm (q), and a mid-section (centromere). The duplicated chromosomes are also joined together as pairs at the centromere. These chromosomal arms are only seen when the chromosome is folded for cell division.
- p used to identify the chromosome short arm (possibly French, petit).
- q used to identify the chromosome long arm (possibly French, tall), the next letter in alphabet after p
These letters prefixed by the chromosome number and followed by the chromosome band number, indicate gene location.
|
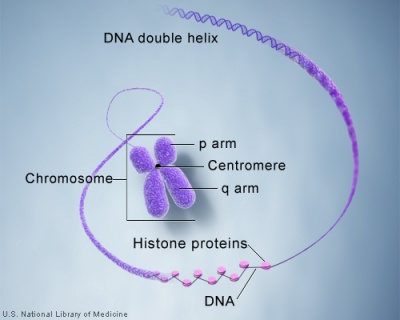
Chromosome pair structure
|
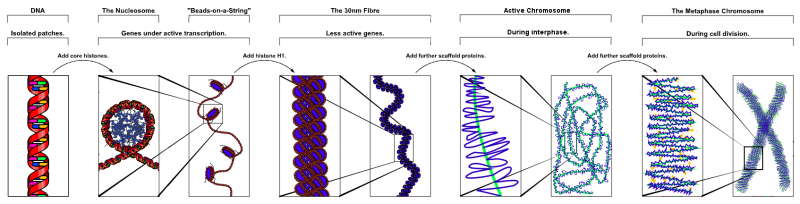 The folding of DNA and associated protein to form a chromosome.
The folding of DNA and associated protein to form a chromosome.
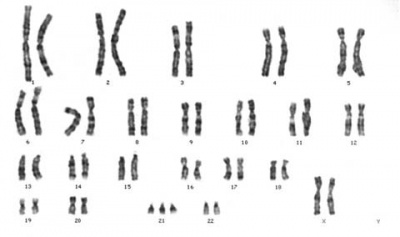
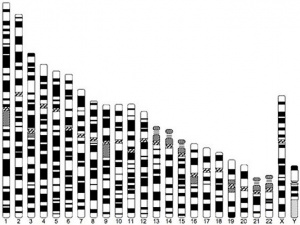
Chromosome Banding
The term refers to the light and dark pattern, seen after staining with a dye, of individual chromosomes identified in metaphase. It is only in meiosis and mitosis during metaphase that chromosomes can be easily identified, during the normal cell life (interphase) the chromosomes are unravelled and distributed within the nucleus in chromosome territories. A band is that part of a chromosome which is clearly distinguishable from nearby regions by appearing darker or brighter with one or more banding techniques.
Depending on the type of stain used a number of different banding patterns can be seen:
- G-banding - banding pattern seen by treating with trypsin and then staining with the dye giemsa.
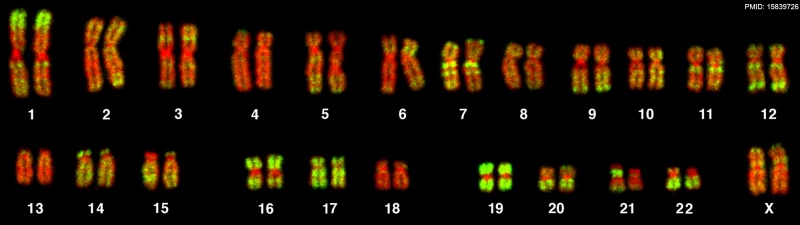
- R-banding - banding pattern seen as a of reverse giemsa chromosome banding, producing bands complementary to G-bands often used to determine whether there are deletions. Can be fluorescent using the dye acridine orange.
- Q-banding - banding pattern seen by treating with a fluorochrome or the fluorescent dye quinacrin.
- C-banding - banding pattern seen for centromeric or constitutive heterochromatin, the centromere appears as a stained band compared to other regions.
Metaphase is a cell division term referring to the third mitotic stage, mitotic spindle kinetochore microtubules align chromosomes in one midpoint plane. Metaphase ends when sister kinetochores separate. Originally based on light microscopy of living cells and electron microscopy of fixed and stained cells. A light microscope analysis called a "metaphase spread" was originally used to detect chromosomal abnormalities in cells.
- Links: Histology Stains
Human Genome
Human Genome Length 3,101,788,170 bp. Mitochondrial Genome 16,568 bp.
- Links: Human Genome Project (HGP) | NCBI Human Genome Resources | History of the Human Genome Project | Project Timeline
Nuclear Genome
Mitochondrial Genome
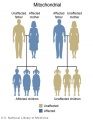
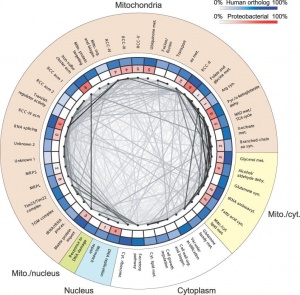 Eukaryotic mitochondrial genomes - In humans this genome is maternally inherited.
- Exists as multiple copies within the matrix of each mitochondrion within the cytoplasm of cells.
- In 1981 the human mitochondrial genome was sequenced.
- The genome is a small circular DNA molecule 16,568 bp in length containing 37 genes.
- 24 genes specify RNA molecules involved in protein synthesis (22 transfer RNAs (tRNA) and 2 ribosomal RNAs (rRNA))
- 13 genes encode proteins required for the biochemical reactions that make up respiration.
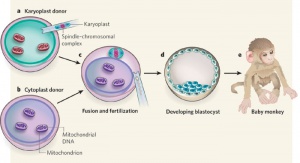
Swapping mitochondrial DNA mammalian oocytes[4] Assisted Reproductive Technology
- Links: Mitochondria | 16569 bp Homo sapiens mitochondrion | Mitochondrial DNA Deletion Syndromes
Chromosome Regions
Chromosome Abnormalities
Ploidy
| ICD-11 LD42 Polyploidies
|
| LD42.0 Triploidy - A disease caused by one additional set of chromosomes, for a total of 69 chromosomes. Triploidy can present with albuminuria, edema, or hypertension in the mother. The fetus may present with microcephaly and a placenta that is enlarged and filled with cysts in the case of extra maternally inherited chromosomes, while extra paternally inherited chromosomes cause severe growth problems, an enlarged head, and a small placenta that does not have cysts. Non-mosaic triploidy is highly lethal, and is rarely observed in live births. Confirmation is through observation of an additional set of chromosomes by karyotyping.
LD42.1 Tetraploidy - A disease caused by two additional sets of chromosomes, for a total of 92 chromosomes. This disease commonly results in spontaneous abortion during the first trimester. Live births of tetraploidy individuals are very rare. These cases are characterized by facial dysmorphism, severely delayed growth and developmental delay. Confirmation is through observation of two additional set of chromosomes by karyotyping.
|
Ploidy refers to the chromosomal genetic content of cells. Euploidy (euploid) is the genetic term used to describe the normal cell genome chromosomal set (n, 2n, 3n) or complement for a species, in humans this is diploid (2n).
The terms used to describe the classes of numerical chromosomal abnormalities include:
- Aneuploidy are chromosome mutations in which chromosome number is abnormal (increased or reduced), nondisjunction in meiosis or mitosis (anaphase of meiosis I, sister chromatids fail to disjoin at either meiosis II or at mitosis) is the cause of most aneuploids.
- Polyploidy includes triploidy, usually due to two sperm fertilizing a single egg.
- Mixoploidy includes mosaicism, where there are two or more genetically different cell lines in an individual.
Trisomy
Can occur in autosomes and sex chromosomes. Presence of one extra chromosome, for a total of three. The most common form is with teh autosome, Trisomy 21.
- Links: Abnormal Development - Genetic | Trisomy 21 | Trisomy 18 | Trisomy 13
Autosome Deletions
| ICD-11 Autosome Deletions
|
|
|
Some Human Disease Gene Locations
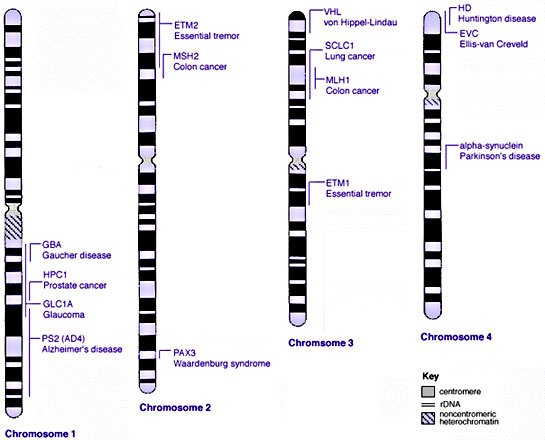
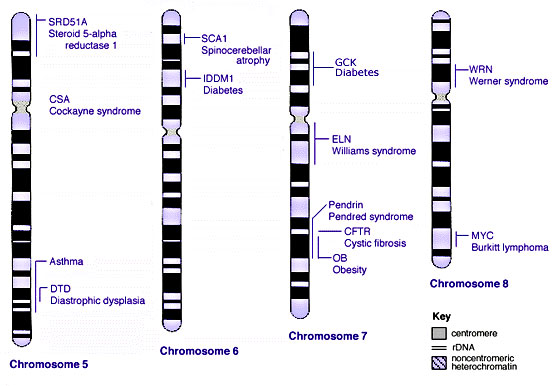
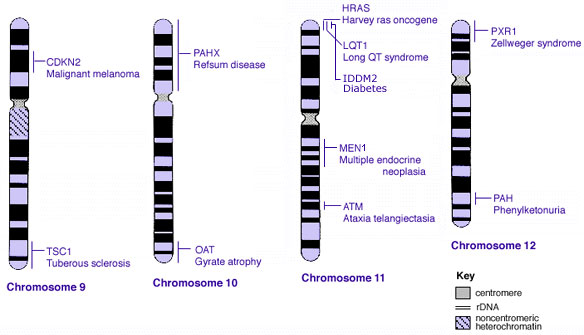
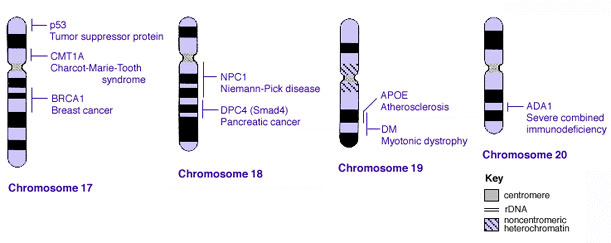
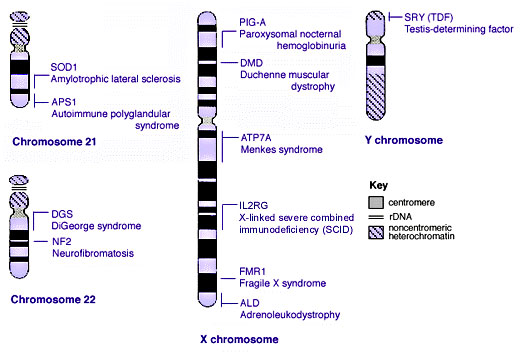
Inheritance Genetics
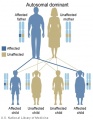
Autosomal dominant inheritance
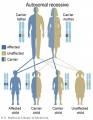
Autosomal recessive inheritance
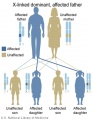
X-Linked dominant (affected father)
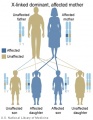
X-Linked dominant (affected mother)
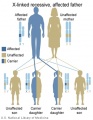
X-Linked recessive (affected father)
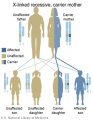
X-Linked recessive (carrier mother)
Mitochondrial genome inheritance
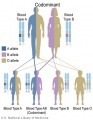
- Inheritance Pattern images: Genetic Abnormalities | autosomal dominant | autosomal recessive | X-linked dominant (affected father) | X-Linked dominant (affected mother) | X-Linked recessive (affected father) | X-Linked recessive (carrier mother) | mitochondrial inheritance | Codominant inheritance | Genogram symbols | Genetics
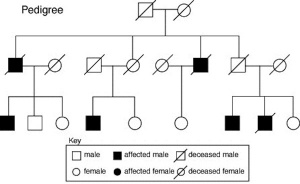
Genogram
A clinical diagram constructed to show an individual's family relationships and medical history, more detailed than a pedigree chart including non-genetic factors such as family emotional and social relationships. Additional colour coded symbols and connectors are used to show these relationships and interactions.
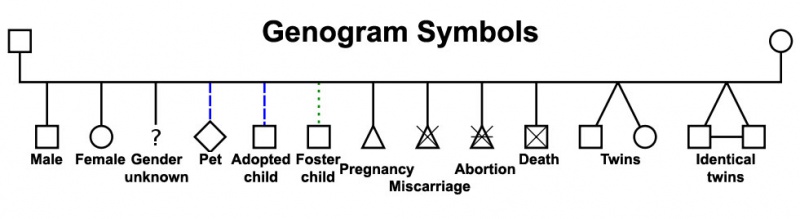
DNA Sequencing
Changes in genes can occur by a number of different methods including mutations, deletions and epigenetic modifications. Specific changes in DNA sequences can also be detected by a range of techniques, including direct sequencing of the DNA.
Recent technological developments have improved how DNA sequencing occurs and we are said to now be up to the "third generation" of sequencing techniques.[5]
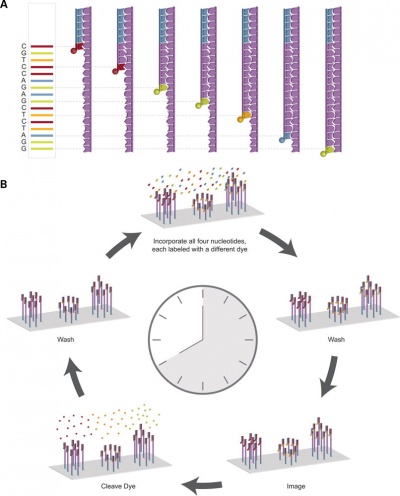
|
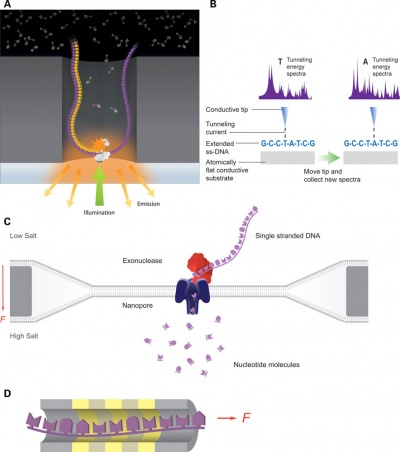
|
| First and second generation sequencing[5]
|
Third generation sequencing[5]
|
Using Genetic Databases
This exercise is an exploration of the available WWW and other database resources that relate to Human genetic diseases. The exercise is to explore the Human genome and its relationship to examples of know human genetic diseases that affect development and impact on health.
To start with, think of a specific Human disease and see what you can find out about:
- Known gene?
- What are the major effects of the disorder?
- Does it have an effect/role in development?
- Known mutations?
- Likely hood of mortality?
- History of the disease?
- Who discovered the cause of the disease?
- Most recent published data relationg to the disease?
- What therapies are being explored?
- Where to next?
Genetic Editing
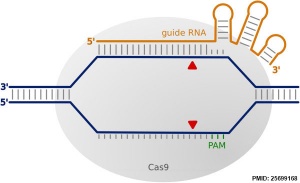 File:CRISPR/Cas9 interaction with target DNA Originally a more general approach was used to study the fly model of development, where random genome mutation (random mutagenesis) was carried out to identify a specific fly phenotype and then researchers would go back and find the gene that had been altered.
More recently in vertebrate models of development, mainly in mice, genetic editing was carried to to "knock out" (KO mice) a specific gene and then to look for a specific phenotype. Generally targeting known genetic disorder genes, but later a range of genes involved in signalling, proliferation, migration, cell cytoskeleton. This technology has developed to the stage where we can now not only "knock out" but also "knock in" as well as "transiently knock out" (at a specific stage) a specific gene of interest. This KO technology was complex and required long term projects to generate these knock out animal models.
CRISPR
More recently a new technology called CRISPR-Cas9 (clustered regularly interspaced short palindromic repeats/CRISPR-associated nuclease 9) has allowed more accurate and faster genetic editing. There has been concern in the scientific community that this technique may be applied to the human genome, and lead to "germ line" changes in the human genome. See also review.[6]
| CRISPR History
|
| Like any great discovery, some contention as to who writes the history...
|
- Links: Nature News 12 March 2015
Other Gene Editing Mechanisms
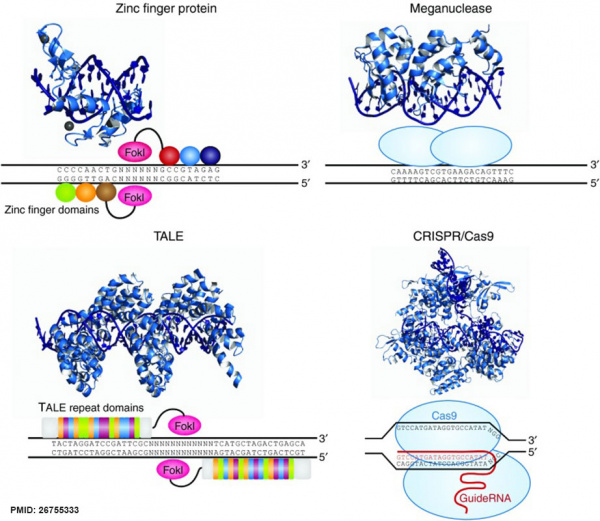
DNA Targeting Platforms for Genome Editing[9]
| Zinc finger nucleases
|
Meganucleases
|
| Zinc finger (ZF) proteins are the most abundant class of transcription factors and the Cys2-His2 zinc finger domain is one of the most common DNA-binding domains encoded in the human genome. The crystal structure of Zif268 has served as the basis for understanding DNA recognition by zinc fingers. In the presence of a zinc atom, the zinc finger domain forms a compact ββα structure with the α-helical portion of each finger making contact with 3 or 4 bp in the major groove of the DNA. Tandem fingers in a zinc finger array wrap around the DNA to bind extended target sequences such that a three-finger protein binds a 9 bp target site.
|
Meganuclease technology involves re-engineering the DNA-binding specificity of naturally occurring homing endonucleases. The largest class of homing endonucleases is the LAGLIDADG family, which includes the well-characterized and commonly used I-CreI and I-SceI enzymes.140 Through a combination of rational design and selection, these homing endonucleases can be re-engineered to target novel sequences.
|
| TALENs
|
CRISPR/Cas nucleases
|
| The discovery of a simple one-to-one code dictating the DNA-binding specificity of TALE proteins from the plant pathogen Xanthomonas again raised the exciting possibility for modular design of novel DNA-binding proteins.114,115 Highly conserved 33–35 amino acid TALE repeats each bind a single base pair of DNA with specificity dictated by two hypervariable residues. Crystal structures of TALEs bound to DNA revealed that each repeat forms a two-helix structure connected by a loop which presents the hypervariable residue into the major groove as the protein wraps around the DNA in a superhelical structure. These modular TALE repeats can be linked together to build long arrays with custom DNA-binding specificities.
|
CRISPR-Cas RNA-guided nucleases are derived from an adaptive immune system that evolved in bacteria to defend against invading plasmids and viruses. Decades of work investigating CRISPR systems in various microbial species has elucidated a mechanism by which short sequences of invading nucleic acids are incorporated into CRISPR loci. They are then transcribed and processed into CRISPR RNAs (crRNAs) which, together with a trans-activating crRNAs (tracrRNAs), complex with CRISPR-associated (Cas) proteins to dictate specificity of DNA cleavage by Cas nucleases through Watson-Crick base pairing between nucleic acids. Building off of two studies showing that the three components required for the type II CRISPR nuclease system are the Cas9 protein, the mature crRNA and the tracrRNA, Doudna, Charpentier and colleagues showed through in vitro DNA cleavage experiments that this system could be reduced to two components by fusion of the crRNA and tracrRNA into a single guide RNA (gRNA). Furthermore, they showed that re-targeting of the Cas9/gRNA complex to new sites could be accomplished by altering the sequence of a short portion of the gRNA. Thereafter, a series of publications demonstrated that the CRISPR/Cas9 system could be engineered for efficient genetic modification in mammalian cells. Collectively these studies have propelled the CRISPR/Cas9 technology into the spotlight of the genome-editing field.
|
(text extract above from original article[9]
References
- ↑ Tang L, Zeng Y, Du H, Gong M, Peng J, Zhang B, Lei M, Zhao F, Wang W, Li X & Liu J. (2017). CRISPR/Cas9-mediated gene editing in human zygotes using Cas9 protein. Mol. Genet. Genomics , 292, 525-533. PMID: 28251317 DOI.
- ↑ Liang P, Xu Y, Zhang X, Ding C, Huang R, Zhang Z, Lv J, Xie X, Chen Y, Li Y, Sun Y, Bai Y, Songyang Z, Ma W, Zhou C & Huang J. (2015). CRISPR/Cas9-mediated gene editing in human tripronuclear zygotes. Protein Cell , 6, 363-372. PMID: 25894090 DOI.
- ↑ Bolzer A, Kreth G, Solovei I, Koehler D, Saracoglu K, Fauth C, Müller S, Eils R, Cremer C, Speicher MR & Cremer T. (2005). Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLoS Biol. , 3, e157. PMID: 15839726 DOI.
- ↑ Shoubridge EA. (2009). Developmental biology: Asexual healing. Nature , 461, 354-5. PMID: 19759608 DOI.
- ↑ 5.0 5.1 5.2 Schadt EE, Turner S & Kasarskis A. (2010). A window into third-generation sequencing. Hum. Mol. Genet. , 19, R227-40. PMID: 20858600 DOI.
- ↑ Sandve GK, Gundersen S, Johansen M, Glad IK, Gunathasan K, Holden L, Holden M, Liestøl K, Nygård S, Nygaard V, Paulsen J, Rydbeck H, Trengereid K, Clancy T, Drabløs F, Ferkingstad E, Kalas M, Lien T, Rye MB, Frigessi A & Hovig E. (2013). The Genomic HyperBrowser: an analysis web server for genome-scale data. Nucleic Acids Res. , 41, W133-41. PMID: 23632163 DOI.
- ↑ Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA & Charpentier E. (2012). A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science , 337, 816-21. PMID: 22745249 DOI.
- ↑ Lander ES. (2016). The Heroes of CRISPR. Cell , 164, 18-28. PMID: 26771483 DOI.
- ↑ 9.0 9.1 Maeder ML & Gersbach CA. (2016). Genome-editing Technologies for Gene and Cell Therapy. Mol. Ther. , 24, 430-46. PMID: 26755333 DOI.
Online Textbooks
- Genomes (2nd edition) Brown, T.A. New York and London: Garland Science ; c2002 PMID20821850
- Introduction to Genetic Analysis Griffiths, Anthony J.F.; Miller, Jeffrey H.; Suzuki, David T.; Lewontin, Richard C.; Gelbart, William M. New York: W. H. Freeman & Co.; c1999
- Modern Genetic Analysis Griffiths, Anthony J.F.; Gelbart, William M.; Miller, Jeffrey H.; Lewontin, Richard C. New York: W. H. Freeman & Co.; c1999
- Human Molecular Genetics 2 Strachan, Tom and Read, Andrew P. New York and London: Garland Science; c1999
- Genetics for Surgeons Morrison, Patrick J.; Spence, Roy A.J., authors Hatchwell, Eli, series editor London: Remedica; c2000
- Sequence - Evolution - Function: Computational Approaches in Comparative Genomics Koonin, Eugene V; Galperin, Michael Y. Norwell (MA): Kluwer Academic Publishers ; c2003
- The Genetic Landscape of Diabetes [Internet] Dean, Laura; McEntyre, J.R. Bethesda (MD): National Library of Medicine (US), NCBI; 2004 Jun
- Madame Curie Bioscience Database Chapters taken from the Madame Curie Bioscience Database (formerly, Eurekah Bioscience Database) Eurekah.com and Landes Bioscience and Springer Science+Business Media; c2009
Reviews
Salsman J & Dellaire G. (2017). Precision genome editing in the CRISPR era. Biochem. Cell Biol. , 95, 187-201. PMID: 28177771 DOI.
Harrison MM, Jenkins BV, O'Connor-Giles KM & Wildonger J. (2014). A CRISPR view of development. Genes Dev. , 28, 1859-72. PMID: 25184674 DOI.
Götz M & Huttner WB. (2005). The cell biology of neurogenesis. Nat. Rev. Mol. Cell Biol. , 6, 777-88. PMID: 16314867 DOI.
Articles
Tang L, Zeng Y, Du H, Gong M, Peng J, Zhang B, Lei M, Zhao F, Wang W, Li X & Liu J. (2017). CRISPR/Cas9-mediated gene editing in human zygotes using Cas9 protein. Mol. Genet. Genomics , 292, 525-533. PMID: 28251317 DOI.
Liang P, Xu Y, Zhang X, Ding C, Huang R, Zhang Z, Lv J, Xie X, Chen Y, Li Y, Sun Y, Bai Y, Songyang Z, Ma W, Zhou C & Huang J. (2015). CRISPR/Cas9-mediated gene editing in human tripronuclear zygotes. Protein Cell , 6, 363-372. PMID: 25894090 DOI.
Search PubMed
Not easy to generate a good search term for this topic.
Search "Genetic Development" All (151839) Review (31389) Free Full Text (54818)
Search Pubmed: Genetic Development | Genome
Genetic Terms
| Genetic Terms (expand to view)
|
genetic abnormalities | Molecular Development | meiosis | mitosis
- Alpha-Fetoprotein test (APF test) A prenatal test to measure the amount of a fetal protein in the mother's blood (or amniotic fluid). Abnormal amounts of the protein may indicate genetic or developmental problems in the fetus. Serum alpha-fetoprotein (AFP) is a fetal glycoprotein produced by the yolk sac and fetal liver. Low levels of AFP normally occur in the blood of a pregnant woman, high levels may indicate neural tube defects (spina bifida, anencephaly). (More? Alpha-Fetoprotein)
- anaphase B - Cell division term referring to the part of anaphase during which the poles of the mitotic spindle move apart. (More? mitosis)
- antisense - a sequence of DNA that is complementary usually to coding sequence of DNA or mRNA. Has been used experimentally to perturb or block gene expression. Also a mechanism that has been found to occur naturally as a regulatory mechanism.
- aneuploidy - Genetic term used to describe an abnormal number of chromosomes mainly (90%) due to chromosome malsegregation mechanisms in maternal meiosis I.
- autosomal inheritance - some hereditary diseases are described as autosomal which means that the disease is due to a DNA error in one of the 22 pairs that are not sex chromosomes. Both boys and girls can then inherit this error. If the error is in a sex chromosome, the inheritance is said to be sex-linked.
- base - another term for nucleotide (usually a t c g).
- base pair - Double stranded DNA has nucleotides A-T, C-G, paired by hydrogen bonds (2 for AT, 3 for GC). Note this means that GC is harder to separate that AT.
- cis-acting elements - DNA sequences that through transcription factors or other trans-acting elements or factors, regulate the expression of genes on the same chromosome.
- cohesin - a multi-protein subunit complex required to keep the sister chromatids together until their separation at anaphase (both in mitosis and meiosis), can also form rings that connect two DNA segments.
- copy number variation - (copy number variants,CNVs) a DNA segment of one kilobase (kb) or larger that is present at a variable copy number in comparison with a reference genome. (More? Nature)
- disomy - Genetic term referring to the presence of two chromosomes of a homologous pair in a cell, as in diploid. See chromosomal number genetic disorders uniparental disomy and aneuploidy. Humans have pairs usually formed by one chromosome from each parent.
- DNMT - DNA methyltransferase.
- DNA - DeoxyriboNucleic Acid. The genetic material found in mammalian chromosomes and mitochondria. Consisting of 4 nucleic acids (ATCG) that combine in a triptych (3 nucleotide codon) code for protein amino acids (3nt = 1aa).
- DNA duplex - double stranded base-paired DNA forming a helix.
- dominant inheritance - With autosomal dominant inheritance, there is an error in one of the 22 chromosome pairs. But the damaged gene dominates over the normal gene received from the other parent. If one of the parents has a disease caused by an autosomal dominant gene, all the children will have a 50 per cent risk of inheriting the dominant gene and a 50 per cent chance of not inheriting it. The children who do not inherit the damaged dominant gene will not themselves suffer from the disease, nor will they be able to pass the gene on to future children. This type of inheritance is present for example in Huntington's disease.
- Down Syndrome - The historic name used for Trisomy 21, named after the original identifier Down, J.L.H. in a 1866 paper (cited above).
- enhancer - A cis-regulatory sequence that can regulate levels of transcription from an adjacent promoter. Many tissue-specific enhancers can determine spatial patterns of gene expression in higher eukaryotes. Enhancers can act on promoters over many tens of kilobases of DNA and can be 5' or 3' to the promoter they regulate.
- exon - a block of protein encoding sequence of DNA in a gene. Many proteins are made of several exons "stitched" or spliced together by editing out non-coding (intron) sequences.
- fasta - a format for listing DNA sequence, where the first line has descritive information followed on the next line by the sequence without numbering.
- GC repeat - a string of GC sequence repeated several times. Also associated with GC expansion, a mutational process that may lead eventually to serious gene expression effects.
- gene - a sequence of DNA that encodes an individual protein.
- genetic code - the 3 nucleotide sequence that forms a codon for a single amino acid or stop. See the gene code.
- genome - the complete genetic information in the form of DNA available to a specific species.
- hairpin loop - a folding of RNA generated by base pairing making a "===()" structure, the end loop and or stem of this structure can then interact with proteins or other RNA.
- heteroplasmy - within a cell when more than one type of mitochondrial DNA (mtDNA) genome exists within the mitochondrion, or between mitochondria. Heteroplasmy can generate a pathogenic mutation, for example transfer RNA leucine (tRNALeu(UUR)) 3243A > G mutant can result in diabetes.
- HyperD - hypermethylated domain has a role in regulating gene expression and epigenomics. The early embryo has large alterations in methylation patterns and DNA modification. (More? PMID 1943996)
- HypoD - hypomethylated domain has a role in regulating gene expression and epigenomics. The early embryo has large alterations in methylation patterns and DNA modification. (More? PMID 1943996)
- igDMR - imprinted germline differentially methylated regions
- intron - a block of DNA within a gene not encoding a protein. Edited, spliced, out during transcription into mRNA. Originally thought not to contain any information, but more and more this appears not to be the case. Some intron sequences have been shown to regulate gene expression during development (eg c elegans, Lin 14)
- karyotype - [Greek, karyon = kernel or nucleus + typos= stamp] The chromosomal makeup of a cell. Karyotyping describes the clinical genetic test.
- meiosis I (MI) The first part of meiosis resulting in separation of homologous chromosomes, in humans producing two haploid cells (N chromosomes, 23), a reductional division.
- Meiosis I: Prophase I - Metaphase I - Anaphase I - Telophase I
- meiosis II - (MII) The second part of meiosis. In male human spermatogenesis, producing of four haploid cells (23 chromosomes, 1N) from the two haploid cells (23 chromosomes, 1N), each of the chromosomes consisting of two sister chromatids produced in meiosis I. In female human oogenesis, only a single haploid cell (23 chromosomes, 1N) is produced. Meiosis II: Prophase II - Metaphase II - Anaphase II - Telophase II
- mosaic autosome monosomy - genetic abnormality caused by embryonic fusion or loss of an autosome early in embryonic development, resulting in a subset of cells in the body having only one of a pair of autosomes. ICD-11 LD43.1 Mosaic monosomy of autosome
- mRNA - messenger, transcribed from DNA in the nucleus and in mitochondria. Is translated by the ribosome in the cytoplasm (or mitochondrial matrix). Intermediate step in gene expression. (DNA-> mRNA-> protein).
- mutation - any process which results in the alteration of the DNA sequence. Some conservative mutations may have no effect on the final amino acid encoded.
- nuchal translucency - (fetal nuchal-translucency thickness) An initial Trisomy 21 diagnostic ultrasound measurement in the fetal neck region carried out by trans-abdominal ultrasound at gestational age GA 10–14 weeks. Fetal sagittal section scan at a magnification that the fetus occupied at least 75% of the image. Measured is the maximum thickness of the subcutaneous translucency between the skin and the soft tissue overlying the cervical spine.
- Philadelphia chromosome - (Philadelphia translocation) Genetic term referring to a chromosomal abnormality resulting from a reciprocal translocation between chromosome 9 and 22 (t(9;22)(q34;q11)). This is associated with the disease chronic myelogenous leukemia (CML).
- ploidy - refers to the chromosomal genetic content of cells.
- promoter - A regulatory region a short distance upstream from the 5' end of a transcription start site that acts as the binding site for RNA polymerase II. A region of DNA to which RNA polymerase IIbinds in order to initiate transcription.
- point mutation - a change in a single nucleotide.
- recessive inheritance - With autosomal recessive inheritance, the diseased individual has inherited the same gene damage from both father and mother. The damage is found on both chromosomes in the pair. But as this is not ´dominant gene damageª, neither father nor mother show any sign of disease, they are healthy carriers of the gene. We are all carriers of about five recessive genes of this type, but as spouses are seldom carriers of exactly the same damaged gene(s), all will probably go well in the next generation.
- regulatory sequence - (regulatory region, regulatory area) is a segment of DNA where regulatory proteins such as transcription factors bind preferentially.
- ribosome - complex of rRNA and ribosomal proteins, bind mRNA and translate it into protein.
- RNA - RiboNucleic Acid. The intermediate nucleic acid involved in gene expression. It comes in 3 forms: tRNA, mRNA, rRNA.
- rRNA - ribosomal, translates mRNA into protein. rRNA provides the "scaffolding" on which many ribosomal proteins are assembled as 2 subunits that themselves assemble to form a ribosome. rRNA genes are localized to the nucleolus in the nucleus, a sometimes visible region of DNA usually constantly being transcribed.
- segmental aneuploidies - generated when a small piece of a chromosome is gained or lost during cell division, resulting in subchromosomal copy number (CN) changes.
- single umbilical artery - (SUA) Placental cord with only a single placental artery (normally paired). This abnormality can be detected by ultrasound (colour flow imaging of the fetal pelvis) and is used as an indicator for further prenatal diagnostic testing for chromosomal abnormalities and other systemic defects. (More? Prenatal Diagnosis | Ultrasound)
- telophase - Cell division term referring to the fifth mitotic stage, where the vesicles of the nuclear envelope reform around the daughter cells, the nucleoli reappear and the chromosomes unfold to allow gene expression to begin. This phase overlaps with cytokinesis, the division of the cell cytoplasm.
- telomere - regions at the end of chromosomes. Shortening of the telomeres is thought to be associated with cellular aging. The enzyme that maintains the telomere is called telomerase. Introducing this gene into a cell can extend the cells lifespan.
- topologically associating domain - (TAD) a self-interacting genomic region, DNA sequences within a TAD physically interact with each other more frequently than with sequences outside the TAD.
- transcription factor - a protein which binds to DNA activating (usually) gene expression. There are many different ways and forms that this activation can take place, but most transcription factors fall into specific classes (eg zinc fingers, helix loop helix).
- triple markers - alpha-fetoprotein, human chorionic gonadotropin, and unconjugated estriol. ))trisomy 21}}
- triploidy - genetic abnormality caused by one additional set of chromosomes, for a total of 69 chromosomes. Maternally present with albuminuria, edema, or hypertension. Extra maternally inherited chromosomes, microcephaly and an enlarged placenta that is enlarged and filled with cysts. Extra paternally inherited chromosomes, severe growth problems, enlarged head, and a small placenta. Non-mosaic triploidy is highly lethal, and is rarely observed in live births. ICD-11 LD42.0 Triploidy
- trisomy mosaicism - a rare chromosome disorder characterized by having an extra copy of a chromosome in a proportion, but not all, of a person’s cells.
- tRNA - transfer, binds single amino acids acts as a "donor' for protein synthesis.
- uniparental disomy - Genetic term referring to cells containing both copies of a homologous pair of chromosomes from one parent and none from the other parent.
|
|
|
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 18) Embryology Molecular Development - Genetics. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Molecular_Development_-_Genetics
- What Links Here?
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
|