Genetics - Chromosome 7: Difference between revisions
mNo edit summary |
|||
| Line 17: | Line 17: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* | * '''Maintenance of Mest imprinted methylation in blastocyst-stage mouse embryos is less stable than other imprinted loci following superovulation or embryo culture'''{{#pmid:29492315|PMID29492315}} "Assisted reproductive technologies are fertility treatments used by subfertile couples to conceive their biological child. Although generally considered safe, these pregnancies have been linked to genomic imprinting disorders, including Beckwith-Wiedemann and Silver-Russell Syndromes. Silver-Russell Syndrome is a growth disorder characterized by pre- and post-natal growth retardation. The Mest imprinted domain is one candidate region on chromosome 7 implicated in Silver-Russell Syndrome. We have previously shown that maintenance of imprinted methylation was disrupted by superovulation or embryo culture during pre-implantation mouse development. For superovulation, this disruption did not originate in oogenesis as a methylation acquisition defect. However, in comparison to other genes, Mest exhibits late methylation acquisition kinetics, possibly making Mest more vulnerable to perturbation by environmental insult. In this study, we present a comprehensive evaluation of the effects of superovulation and in vitro culture on genomic imprinting at the Mest gene. Superovulation resulted in disruption of imprinted methylation at the maternal Mest allele in blastocysts with an equal frequency of embryos having methylation errors following low or high hormone treatment. This disruption was not due to a failure of imprinted methylation acquisition at Mest in oocytes. For cultured embryos, both the Fast and Slow culture groups experienced a significant loss of maternal Mest methylation compared to in vivo-derived controls. This loss of methylation was independent of development rates in culture. These results indicate that Mest is more susceptible to imprinted methylation maintenance errors compared to other imprinted genes." [[Molecular Development - Epigenetics|Epigenetics]] | ||
|} | |} | ||
{| class="wikitable mw-collapsible mw-collapsed" | {| class="wikitable mw-collapsible mw-collapsed" | ||
Revision as of 10:09, 20 March 2018
Introduction
|

|
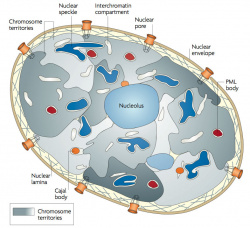
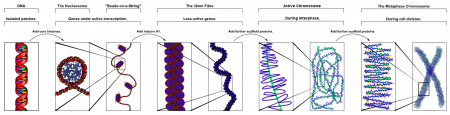
| Chromosome territories (interphase) | Chromosome (Chromatin) structure (mitosis) |
| Human Chromosomes: 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | X | Y |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Chromosome 7 <pubmed limit=5>Chromosome 7</pubmed> |
Development Genes
WNT
| Table - Human Wnt Family | ||||
| Approved Symbol |
Approved Name | Previous Symbols |
Synonyms | Chromosome |
|---|---|---|---|---|
| WNT2 | Wnt family member 2 | INT1L1 | IRP | 7q31.2 |
| WNT16 | Wnt family member 16 | 7q31.31 | ||
| Links: Developmental Signals - Wnt | OMIM Wnt1 | HGNC | Bmp Family | Fgf Family | Pax Family | R-spondin Family | Sox Family | Tbx Family | Wnt Family | ||||
| Human WNT Family | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
| |
| Idiogram Chromosome Banding - The term refers to the light and dark pattern, seen after staining with a dye, of individual chromosomes identified in metaphase. It is only in meiosis and mitosis during metaphase that chromosomes can be easily identified, during the normal cell life (interphase) the chromosomes are unravelled and distributed within the nucleus in chromosome territories. A band is that part of a chromosome which is clearly distinguishable from nearby regions by appearing darker or brighter with one or more banding techniques. | |
| Genetic abnormality locations: 1-4 | 5-8 | 9-12 | 13-16 | 17-20 | 21-XY | sSMC | |
| |
| Links: Genetics | Abnormal Development - Genetic |
Cite this page: Hill, M.A. (2024, April 17) Embryology Genetics - Chromosome 7. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Genetics_-_Chromosome_7
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
Cite this page: Hill, M.A. (2024, April 17) Embryology Genetics - Chromosome 7. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Genetics_-_Chromosome_7
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
