Genetics - Chromosome 12: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 17: | Line 17: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* ''' | * '''Endometriosis'''{{#pmid:29506960|PMID29506960}} "Developments in high-throughput genotyping technology have driven discovery of genomic regions associated with an increased risk of endometriosis. In all, 16 genomic regions have been associated with risk of endometriosis in one or more populations. The latest meta-analysis including 17,045 endometriosis cases identified 14 genomic regions supported by results from multiple studies. No independent associations were identified from direct genotyping of common and low-frequency protein-coding variants. This suggests that the most common genetic factors that contribute to endometriosis risk are located in regulatory DNA sequences and alter the regulation of gene transcription. Evidence from different methods is essential to identify the target genes and transcripts that contribute to altered disease risk. Potential target genes in three chromosome regions showing altered gene regulation include LINC00339 and CDC42 on chromosome {{Chr1}}, CDKN2A-AS1 on chromosome 9, and VEZT on chromosome {{Chr12}}. | ||
* '''Spinocerebellar Ataxia Type 2'''{{#pmid:29422848|PMID29422848}} "Spinocerebellar ataxia type 2 (SCA2) is an autosomal dominant spinocerebellar degeneration, associated with extended repeats of the trinucleotide CAG in the ATXN2 gene on the long arm of chromosome {{Chr12}}. Magnetic resonance imaging (MRI) of SCA2 showed significant atrophies of the brainstem, middle cerebellar peduncles, and cerebellum. We report two genetically proven SCA2 patients who showed hypertrophy of the inferior olivary nuclei on proton density- and T2-weighted MRI. This pattern has never been reported in patients with SCA1, SCA3, or SCA6, and may make it possible to differentiate SCA2 from other hereditary spinocerebellar ataxias." [[Magnetic Resonance Imaging]] | |||
|} | |} | ||
{| class="wikitable mw-collapsible mw-collapsed" | {| class="wikitable mw-collapsible mw-collapsed" | ||
Latest revision as of 10:28, 20 March 2018
Introduction
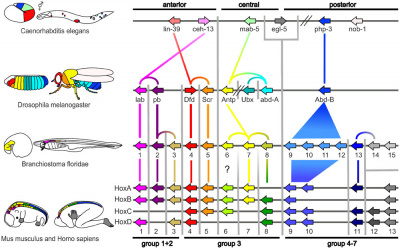
The one set (or four) of the HOX genes are an important set of developmental patterning genes located on chromosome 12.
|

|
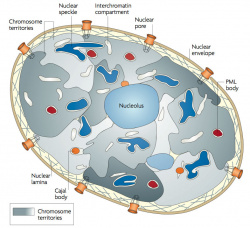
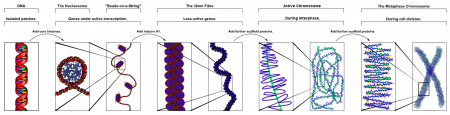
| Chromosome territories (interphase) | Chromosome (Chromatin) structure (mitosis) |
| Human Chromosomes: 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | X | Y |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Chromosome 12 <pubmed limit=5>Chromosome 12</pubmed> |
Development Genes
HOX
Proposed Hox Protein Classification[3]
WNT
| Table - Human Wnt Family | ||||
| Approved Symbol |
Approved Name | Previous Symbols |
Synonyms | Chromosome |
|---|---|---|---|---|
| WNT1 | Wnt family member 1 | INT1 | 12q13.12 | |
| WNT5B | Wnt family member 5B | 12p13.33 | ||
| WNT10B | Wnt family member 10B | "WNT-12, SHFM6" | 12q13.12 | |
| Links: Developmental Signals - Wnt | OMIM Wnt1 | HGNC | Bmp Family | Fgf Family | Pax Family | R-spondin Family | Sox Family | Tbx Family | Wnt Family | ||||
| Human WNT Family | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
| |
| Idiogram Chromosome Banding - The term refers to the light and dark pattern, seen after staining with a dye, of individual chromosomes identified in metaphase. It is only in meiosis and mitosis during metaphase that chromosomes can be easily identified, during the normal cell life (interphase) the chromosomes are unravelled and distributed within the nucleus in chromosome territories. A band is that part of a chromosome which is clearly distinguishable from nearby regions by appearing darker or brighter with one or more banding techniques. | |
| Genetic abnormality locations: 1-4 | 5-8 | 9-12 | 13-16 | 17-20 | 21-XY | sSMC | |
| |
| Links: Genetics | Abnormal Development - Genetic |
Cite this page: Hill, M.A. (2024, April 25) Embryology Genetics - Chromosome 12. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Genetics_-_Chromosome_12
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
Cite this page: Hill, M.A. (2024, April 25) Embryology Genetics - Chromosome 12. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Genetics_-_Chromosome_12
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
- ↑ Fung JN & Montgomery GW. (2018). Genetics of endometriosis: State of the art on genetic risk factors for endometriosis. Best Pract Res Clin Obstet Gynaecol , , . PMID: 29506960 DOI.
- ↑ Yoshii F, Tomiyasu H, Watanabe R & Ryo M. (2017). MRI Signal Abnormalities of the Inferior Olivary Nuclei in Spinocerebellar Ataxia Type 2. Case Rep Neurol , 9, 267-271. PMID: 29422848 DOI.
- ↑ Hueber SD, Weiller GF, Djordjevic MA & Frickey T. (2010). Improving Hox protein classification across the major model organisms. PLoS ONE , 5, e10820. PMID: 20520839 DOI.