Developmental Signals - TGF-beta: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 10: | Line 10: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* '''Spatio-temporal distribution of Smads and role of Smads/TGF-β/BMP-4 in the regulation of mouse bladder organogenesis'''<ref name ="PMID23620745"><pubmed>23620745</pubmed> "Although Shh, TGF-β and BMP-4 regulate radial patterning of the bladder mesenchyme and smooth muscle differentiation, it is not known what transcription factors, local environmental cues or signaling cascades mediate bladder smooth muscle differentiation. ...Based on the Smad expression patterns, we suggest that individual or combinations of Smads may be necessary during mouse bladder organogenesis and may be critical mediators for bladder smooth muscle differentiation." [[Urinary Bladder Development]] | * '''Spatio-temporal distribution of Smads and role of Smads/TGF-β/BMP-4 in the regulation of mouse bladder organogenesis'''<ref name ="PMID23620745"><pubmed>23620745</pubmed></ref> "Although Shh, TGF-β and BMP-4 regulate radial patterning of the bladder mesenchyme and smooth muscle differentiation, it is not known what transcription factors, local environmental cues or signaling cascades mediate bladder smooth muscle differentiation. ...Based on the Smad expression patterns, we suggest that individual or combinations of Smads may be necessary during mouse bladder organogenesis and may be critical mediators for bladder smooth muscle differentiation." [[Urinary Bladder Development]] | ||
|} | |} | ||
Revision as of 01:01, 1 December 2013
Introduction
Transforming Growth Factor-beta (TGF-β)
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: TGF-β Embryology <pubmed limit=5>TGF-β Embryology</pubmed> |
Structure
The TGF precursor protein has three distinct regions:
- signal peptide - targets it to the endoplasmic reticulum and secretion
- propeptide - or the latency associated peptide
- mature peptide - cleaved from the precursor protein and is actively involved in signalling
- cleaved by Furin - a convertase
- cleaved at a dibasic arginine-X-X-arginine (RXXR) site
Function
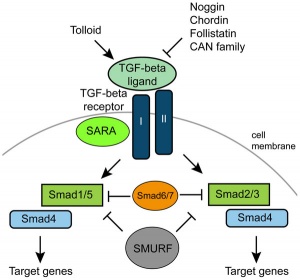
Signaling Pathway

Receptor
- active peptide forms a hetero- or homodimer
- binds to a specific TGF-β Type II receptor
- Type II receptor then recruits a TGF-β Type I receptor
- phosphorylates it via its serine/threonine kinase domain
- phosphorylated Type I receptors then phosphorylate receptor-associated Smad proteins (R-Smads), including Smad1/5 and Smad2/3
- Type II receptor - MlTgfRII
- Type I receptors - MlTgfRIa, MlTgfRIb, and MlTgfRIc
Intracellular Signaling
- R-Smad proteins are composed of two main functional domains
- Mad-homology domains 1 and 2 (MH1 and MH2)
- Smad1/5 - associated with BMP-like signalling.
- Smad2/3 - associated with TGF-β-like signaling.
OMIM
About OMIM "Online Mendelian Inheritance in Man OMIM is a comprehensive, authoritative, and timely compendium of human genes and genetic phenotypes. The full-text, referenced overviews in OMIM contain information on all known mendelian disorders and over 12,000 genes. OMIM focuses on the relationship between phenotype and genotype. It is updated daily, and the entries contain copious links to other genetics resources." OMIM
References
Reviews
Articles
<pubmed>19192293</pubmed>| BMC Evol Biol. <pubmed>17077151</pubmed>
Online Textbooks
- Molecular Biology of the Cell. 4th edition. Alberts B, Johnson A, Lewis J, et al.New York: Garland Science; 2002. Signal Proteins of the TGF-β Superfamily Act Through Receptor Serine/Threonine Kinases and Smads
- Molecular Cell Biology. 4th edition. Lodish H, Berk A, Zipursky SL, et al.New York: W. H. Freeman; 2000. Figure 24-20 TGFβ signaling
- Madame Curie Bioscience Database [Internet]. Austin (TX): Landes Bioscience; 2000. TGFβ-dependent Epithelial-Mesenchymal Transition
Search PubMed
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- Abcam TGF-Beta Antibodies
- SABiosciences TGF-Beta Pathway
- RnD Systems Transforming Growth Factor Beta 1
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 25) Embryology Developmental Signals - TGF-beta. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_TGF-beta
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
