Developmental Signals - Sox: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 18: | Line 18: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* '''Long-term expandable SOX9+ chondrogenic ectomesenchymal cells from human pluripotent stem cells'''<ref name=PMID25818812><pubmed>25818812</pubmed></ref> "Here we report the successful generation and long-term expansion of SOX9-expressing CD271(+)PDGFRα(+)CD73(+) chondrogenic ectomesenchymal cells from the PAX3/SOX10/FOXD3-expressing MIXL1(-)CD271(hi)PDGFRα(lo)CD73(-) neural crest-like progeny of human pluripotent stem cells in a chemically defined medium supplemented with Nodal/Activin/transforming growth factorβ (TGFβ) inhibitor and fibroblast growth factor (FGF). When "primed" with TGFβ, such cells efficiently formed translucent cartilage particles, which were completely mineralized in 12 weeks in immunocompromized mice." [[Musculoskeletal System - Cartilage Development|Cartilage Development]] | |||
* '''Just how conserved is vertebrate sex determination?'''<ref name="PMID23390004"><pubmed>23390004</pubmed></ref> "Sex determination in vertebrate embryos has long been equated with gonadal differentiation into testes or ovaries. This view has been challenged over the years by reports of somatic sexual dimorphisms pre-dating gonadal sex differentiation. ...We illustrate these differences by comparing key sex genes in fishes versus birds and mammals, with emphasis on DM domain genes, the SOX-9AMH pathway in the testis and the FOXL2-Aromatase pathway in the ovary. Such comparisons facilitate the identification of ancient versus derived genes involved in gonadal sex determination. The data indicate that vertebrate sex-determining cascades are not as conserved as once thought." | * '''Just how conserved is vertebrate sex determination?'''<ref name="PMID23390004"><pubmed>23390004</pubmed></ref> "Sex determination in vertebrate embryos has long been equated with gonadal differentiation into testes or ovaries. This view has been challenged over the years by reports of somatic sexual dimorphisms pre-dating gonadal sex differentiation. ...We illustrate these differences by comparing key sex genes in fishes versus birds and mammals, with emphasis on DM domain genes, the SOX-9AMH pathway in the testis and the FOXL2-Aromatase pathway in the ovary. Such comparisons facilitate the identification of ancient versus derived genes involved in gonadal sex determination. The data indicate that vertebrate sex-determining cascades are not as conserved as once thought." | ||
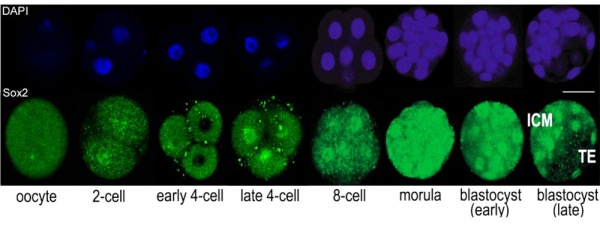
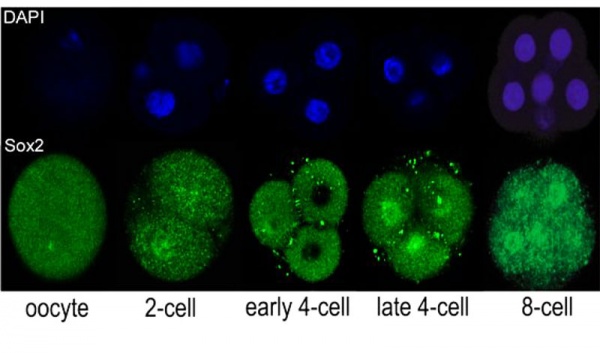
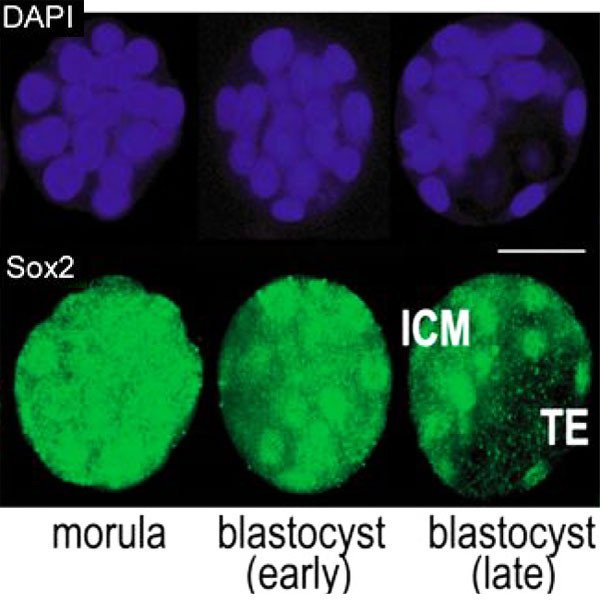
* '''Sox2 is essential for formation of trophectoderm in the preimplantation embryo'''<ref name="PMID21103067"><pubmed>21103067</pubmed>| [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2980489 PMC2980489] | [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0013952 PLoS One.]</ref> "In preimplantation mammalian development the transcription factor Sox2 (SRY-related HMG-box gene 2) forms a complex with Oct4 and functions in maintenance of self-renewal of the pluripotent inner cell mass (ICM). ...We conclude that the first essential function of Sox2 in the preimplantation mouse embryo is to facilitate establishment of the trophectoderm lineage. Our findings provide a novel insight into the first differentiation event within the preimplantation embryo, namely the segregation of the ICM and TE lineages." | * '''Sox2 is essential for formation of trophectoderm in the preimplantation embryo'''<ref name="PMID21103067"><pubmed>21103067</pubmed>| [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2980489 PMC2980489] | [http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0013952 PLoS One.]</ref> "In preimplantation mammalian development the transcription factor Sox2 (SRY-related HMG-box gene 2) forms a complex with Oct4 and functions in maintenance of self-renewal of the pluripotent inner cell mass (ICM). ...We conclude that the first essential function of Sox2 in the preimplantation mouse embryo is to facilitate establishment of the trophectoderm lineage. Our findings provide a novel insight into the first differentiation event within the preimplantation embryo, namely the segregation of the ICM and TE lineages." | ||
| Line 70: | Line 71: | ||
==OMIM== | ==OMIM== | ||
* [http://www.ncbi.nlm.nih.gov/omim/602148 Sox 1] | * [http://www.ncbi.nlm.nih.gov/omim/602148 Sox 1] | ||
* [http://omim.org/entry/608160 Sox 9] | |||
{{About_OMIM}} | |||
==References== | ==References== | ||
Revision as of 10:43, 4 July 2015
| Embryology - 18 Apr 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
The SRY (480000) and SOX proteins share a DNA-binding domain known as the HMG box, defined by a 79-amino acid region. All SOX proteins have a single HMG box and bind linear DNA in a sequence-specific manner, resulting in the bending of DNA through large angles. Bending causes the DNA helix to open for some distance, which may affect binding and interactions of other transcription factors. SOX1, SOX2 (184429), and SOX3 (313430) show the closest homology to SRY. They share maximum homology within the HMG domain and are expressed mainly in the developing nervous system of the mouse (Collignon et al., 1996). These genes share significant homology outside the HMG box also and are highly conserved throughout their evolution.
Sox2 is first expressed in very early (morula, blastocyst) development, and has also been identified as one of the 4 "Yamanaka Factors" required to generate an induced pluripotential stem cell (iPS cell). It also forms a trimeric complex with OCT4, yet another "Yamanaka Factor".
Mammals have 20 different SOX proteins that can be subdivided into 8 groups: A, B1, B2, C, D, E, F, G, H.
- Sox Links: Sox transcription factors cartoon | Image 1 - Preimplantation Mouse | Image 2 - Preimplantation Mouse | Image 3 - Preimplantation Mouse | Sox | Induced Stem Cells | Yamanaka Factors
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Sox Expression <pubmed limit=5>Sox Expression</pubmed> |
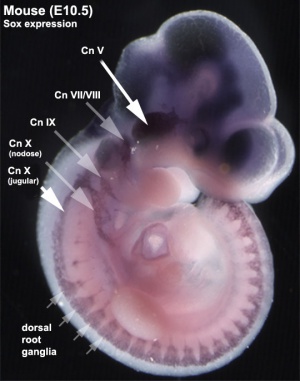
Early Mouse Expression
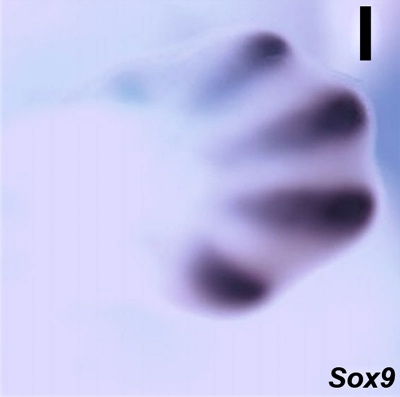
Limb Expression
Sox9 expression in E12.5 wild-type mouse embryonic forelimb.[5]
Function
Respiratory Development
Sox2
- regulates patterning of the anterior foregut into ventral (trachea) and dorsal (esophagus) fates
- endoderm expression during formation of foregut derivatives
- declines in regions undergoing lung bud morphogenesis
- declines in ventral region generating the trachea
- Links: Respiratory System Development | StemBook - Specification and patterning of the respiratory system
Rib Development
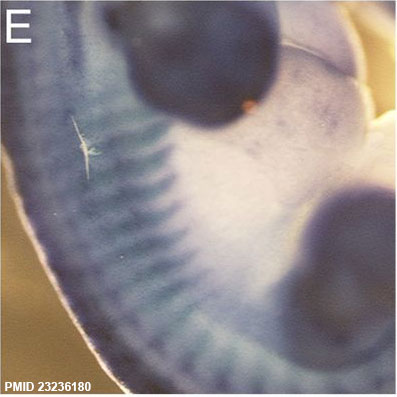
Sox9 expression in the Mouse (E12.5) rib primordial.[6]
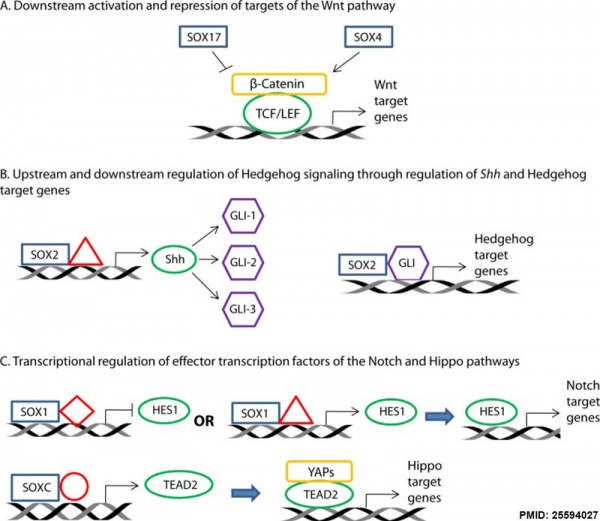
Signaling Pathway
SOX Developmental Signaling Pathways[7]
OMIM
About OMIM "Online Mendelian Inheritance in Man OMIM is a comprehensive, authoritative, and timely compendium of human genes and genetic phenotypes. The full-text, referenced overviews in OMIM contain information on all known mendelian disorders and over 12,000 genes. OMIM focuses on the relationship between phenotype and genotype. It is updated daily, and the entries contain copious links to other genetics resources." OMIM
References
- ↑ <pubmed>20704721</pubmed>| BMC Dev Biol.
- ↑ <pubmed>25818812</pubmed>
- ↑ <pubmed>23390004</pubmed>
- ↑ <pubmed>21103067</pubmed>| PMC2980489 | PLoS One.
- ↑ <pubmed>17194222</pubmed>| PMC1713256 | PLoS Genet.
- ↑ <pubmed>23236180</pubmed>| PMC3535641 | PNAS
- ↑ <pubmed>25594027</pubmed>| Oncoscience
Reviews
<pubmed></pubmed> <pubmed>21309066</pubmed> <pubmed>17584862</pubmed>
Search PubMed: Sox
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 18) Embryology Developmental Signals - Sox. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Sox
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G