Developmental Signals - Notch: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 3: | Line 3: | ||
The notch proteins were first identified in drosophila development and have since been identified as regulators of cell fate decisions during development. These are a family of cell surface transmembrane receptors that pass once through the plasma membrane. | The notch proteins were first identified in drosophila development and have since been identified as regulators of cell fate decisions during development. These are a family of cell surface transmembrane receptors that pass once through the plasma membrane. | ||
| Line 12: | Line 13: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* '''Notch regulation of myogenic versus endothelial fates of cells that migrate from the somite to the limb'''<ref name=PMID24927569><pubmed>24927569</pubmed></ref> "Multipotent Pax3-positive (Pax3(+)) cells in the somites give rise to skeletal muscle and to cells of the vasculature. We had previously proposed that this cell-fate choice depends on the equilibrium between Pax3 and Foxc2 expression. In this study, we report that the Notch pathway promotes vascular versus skeletal muscle cell fates. ...We now demonstrate that in addition to the inhibitory role of Notch signaling on skeletal muscle cell differentiation, the Notch pathway affects the Pax3:Foxc2 balance and promotes the endothelial versus myogenic cell fate, before migration to the limb, in multipotent Pax3(+) cells in the somite of the mouse embryo." [[Musculoskeletal_System_-_Limb_Development|Limb Development]] | [[Musculoskeletal_System_-_Muscle_Development|Muscle Development]] | |||
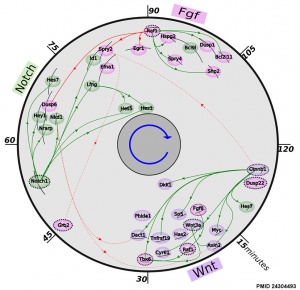
* '''The precise timeline of transcriptional regulation reveals causation in mouse somitogenesis network'''<ref name=PMID24304493><pubmed>24304493</pubmed>| [http://www.biomedcentral.com/1471-213X/13/42 BMC Dev Biol.]</ref> "In vertebrate development, the segmental pattern of the body axis is established as somites, masses of mesoderm distributed along the two sides of the neural tube, are formed sequentially in the anterior-posterior axis. This mechanism depends on waves of gene expression associated with the Notch, Fgf and Wnt pathways." | * '''The precise timeline of transcriptional regulation reveals causation in mouse somitogenesis network'''<ref name=PMID24304493><pubmed>24304493</pubmed>| [http://www.biomedcentral.com/1471-213X/13/42 BMC Dev Biol.]</ref> "In vertebrate development, the segmental pattern of the body axis is established as somites, masses of mesoderm distributed along the two sides of the neural tube, are formed sequentially in the anterior-posterior axis. This mechanism depends on waves of gene expression associated with the Notch, Fgf and Wnt pathways." | ||
* '''Role of p63 and the Notch pathway in cochlea development and sensorineural deafness'''<ref name=PMID20513039><pubmed>20513039</pubmed></ref> "The ectodermal dysplasias are a group of inherited autosomal dominant syndromes associated with heterozygous mutations in the Tumor Protein p63 (TRP63) gene. Here we show that, in addition to their epidermal pathology, a proportion of these patients have distinct levels of deafness. ...these data demonstrate that TAp63, acting via the Notch pathway, is crucial for the development of the organ of Corti, providing a molecular explanation for the sensorineural deafness in ectodermal dysplasia patients with TRP63 mutations." [[Sensory - Hearing and Balance Development]] | * '''Role of p63 and the Notch pathway in cochlea development and sensorineural deafness'''<ref name=PMID20513039><pubmed>20513039</pubmed></ref> "The ectodermal dysplasias are a group of inherited autosomal dominant syndromes associated with heterozygous mutations in the Tumor Protein p63 (TRP63) gene. Here we show that, in addition to their epidermal pathology, a proportion of these patients have distinct levels of deafness. ...these data demonstrate that TAp63, acting via the Notch pathway, is crucial for the development of the organ of Corti, providing a molecular explanation for the sensorineural deafness in ectodermal dysplasia patients with TRP63 mutations." [[Sensory - Hearing and Balance Development]] | ||
| Line 18: | Line 20: | ||
* '''Notch signalling in the paraxial mesoderm is most sensitive to reduced Pofut1 levels during early mouse development'''<ref><pubmed>19161597</pubmed></ref> "Notch-dependent processes apparently differ with respect to their requirement for levels of POFUT1. Normal Lfng expression and anterior-posterior somite patterning is highly sensitive to reduced POFUT1 levels in early mammalian embryos, whereas other early Notch-dependent processes such as establishment of left-right asymmetry or neurogenesis are not. Thus, it appears that in the presomitic mesoderm (PSM) Notch signalling is particularly sensitive to POFUT1 levels. Reduced POFUT1 levels might affect Notch trafficking or overall O-fucosylation. Alternatively, reduced O-fucosylation might preferentially affect sites that are substrates for LFNG and thus important for somite formation and patterning." | * '''Notch signalling in the paraxial mesoderm is most sensitive to reduced Pofut1 levels during early mouse development'''<ref><pubmed>19161597</pubmed></ref> "Notch-dependent processes apparently differ with respect to their requirement for levels of POFUT1. Normal Lfng expression and anterior-posterior somite patterning is highly sensitive to reduced POFUT1 levels in early mammalian embryos, whereas other early Notch-dependent processes such as establishment of left-right asymmetry or neurogenesis are not. Thus, it appears that in the presomitic mesoderm (PSM) Notch signalling is particularly sensitive to POFUT1 levels. Reduced POFUT1 levels might affect Notch trafficking or overall O-fucosylation. Alternatively, reduced O-fucosylation might preferentially affect sites that are substrates for LFNG and thus important for somite formation and patterning." | ||
|} | |} | ||
{| class="wikitable collapsible collapsed" | {| class="wikitable mw-collapsible mw-collapsed" | ||
! More recent papers | ! More recent papers | ||
|- | |- | ||
| Line 84: | Line 86: | ||
* OMIM - [http://www.ncbi.nlm.nih.gov/omim/190198 NOTCH 1] | * OMIM - [http://www.ncbi.nlm.nih.gov/omim/190198 NOTCH 1] | ||
Revision as of 08:56, 2 July 2014
| Embryology - 19 Apr 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
The notch proteins were first identified in drosophila development and have since been identified as regulators of cell fate decisions during development. These are a family of cell surface transmembrane receptors that pass once through the plasma membrane.
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Notch <pubmed limit=5>Notch</pubmed> |
Notch Receptors
NOTCH1
NOTCH2
NOTCH3
NOTCH4
Notch Ligands
- JAG1
- JAG2
- DLL1
- DLL3
- DLL4
Functions
Developmental patterning signal.
Mesoderm Development
Cartilage Development
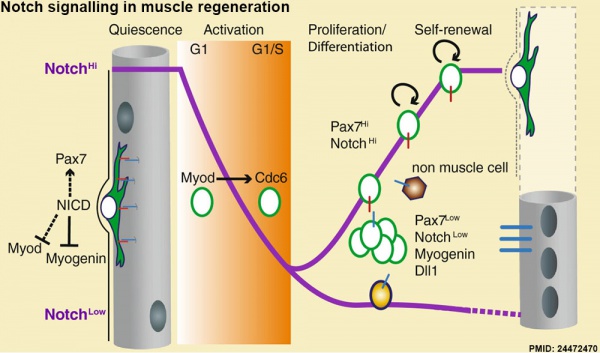
Muscle Regeneration
Notch signalling in muscle regeneration[7]
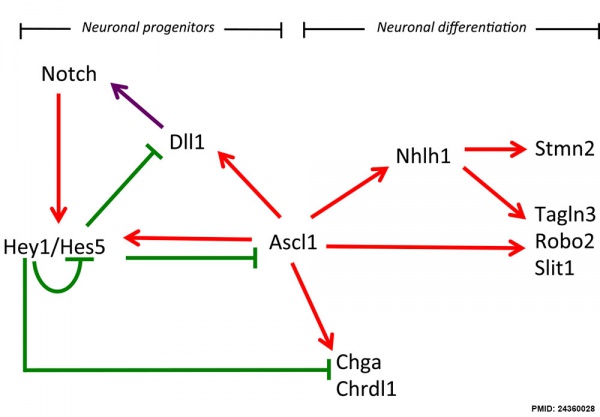
Hypothalamus Development
Hypothalamus Development Gene Interaction Model[8]
- Links: Hypothalamus Development
References
- ↑ 1.0 1.1 <pubmed>24304493</pubmed>| BMC Dev Biol.
- ↑ <pubmed>24927569</pubmed>
- ↑ <pubmed>20513039</pubmed>
- ↑ <pubmed>19590010</pubmed>
- ↑ <pubmed>19590010</pubmed>
- ↑ <pubmed>19161597</pubmed>
- ↑ <pubmed>24472470</pubmed>| BMC Dev Biol.
- ↑ 24360028<pubmed>24360028</pubmed>| Neural Dev.
Reviews
<pubmed></pubmed> <pubmed></pubmed> <pubmed></pubmed> <pubmed>23165243</pubmed> <pubmed> 22399351</pubmed> <pubmed>22397947</pubmed> <pubmed>21828089</pubmed>
Search Pubmed
Search Bookshelf Notch
Search Pubmed Now: Notch Signaling
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- OMIM - NOTCH 1
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 19) Embryology Developmental Signals - Notch. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Notch
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G