|
|
| Line 143: |
Line 143: |
| :OMIM: [http://www.omim.org/entry/600036 OTX1] | [http://www.omim.org/entry/600037 OTX2] | [http://www.omim.org/entry/607410 OTX3] | | :OMIM: [http://www.omim.org/entry/600036 OTX1] | [http://www.omim.org/entry/600037 OTX2] | [http://www.omim.org/entry/607410 OTX3] |
|
| |
|
| :'''Links:''' [[Endocrine - Pineal Development|Pineal Development]] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Otx PubMed - Otx] | | :'''Links:''' [[Endocrine - Pineal Development|Pineal]] | [[Sensory - Vision Development|Vision]] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Otx PubMed - Otx] |
| ==Additional Images== | | ==Additional Images== |
|
| |
| <gallery> | | <gallery> |
| File:Hox deuterostomes phylogenetic tree.jpg|Hox deuterostomes phylogenetic tree PMID 23819519 | | File:Hox deuterostomes phylogenetic tree.jpg|Hox deuterostomes phylogenetic tree PMID 23819519 |
Revision as of 18:01, 21 May 2016
Introduction
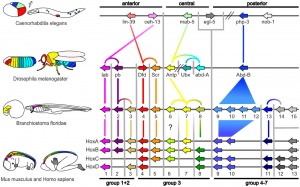
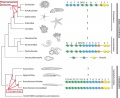
Proposed Hox protein classification
[1]The family of homeobox (Hox) proteins has been a focus of research for over 30 years. This family of genes were also the basis of the embryo patterning studies that led to the Nobel Prize in Medicine 1995. We now know that in addition to whole embryo axes patterning, this family of genes has many roles in establishing pattern throughout the embryo in different tissues and organs.
There has recently been a revival of an earlier theory[2] that Hox expression during vertebrate pattern formation is linked to the process of segmentation of paraxial mesoderm during somitogenesis.[3]
This signalling pathway has also been implicated in many developmental abnormalities and diseases.
|

|
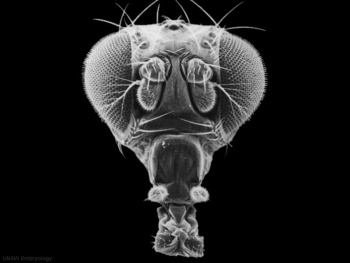
| Fly wild-type head
|
Fly antennapedia head
|
| Category:Hox
Some Recent Findings
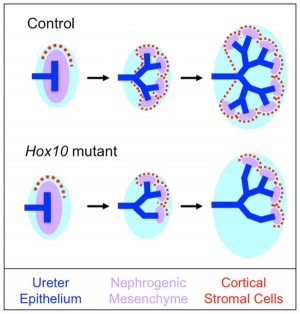
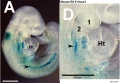
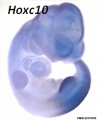
Model Hox10 kidney development
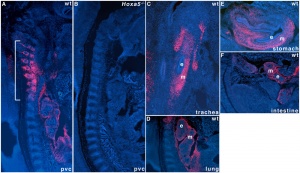
Mouse Hoxa5 expression (E12.5)
[4]
- A single three-dimensional chromatin compartment in amphioxus indicates a stepwise evolution of vertebrate Hox bimodal regulation[5] "The HoxA and HoxD gene clusters of jawed vertebrates are organized into bipartite three-dimensional chromatin structures that separate long-range regulatory inputs coming from the anterior and posterior Hox-neighboring regions. This architecture is instrumental in allowing vertebrate Hox genes to pattern disparate parts of the body, including limbs. ...We find that, in contrast to the architecture in vertebrates, the amphioxus Hox cluster is organized into a single chromatin interaction domain that includes long-range contacts mostly from the anterior side, bringing distant cis-regulatory elements into contact with Hox genes."
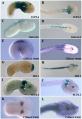
- Homeobox Genes of Caenorhabditis elegans and Spatio-Temporal Expression[6] "We show that, out of 103 homeobox genes, 70 are co-orthologous to human homeobox genes. 14 are highly divergent, lacking an obvious ortholog even in other Caenorhabditis species. One of these homeobox genes encodes 12 homeodomains, while three other highly divergent homeobox genes encode a novel type of double homeodomain, termed HOCHOB. To understand how transcription factors regulate cell fate during development, precise spatio-temporal expression data need to be obtained. Using a new imaging framework that we developed, Endrov, we have generated spatio-temporal expression profiles during embryogenesis of over 60 homeobox genes, as well as a number of other developmental control genes using GFP reporters." Worm Development
- A Hox regulatory network of hindbrain segmentation is conserved to the base of vertebrates[7] "Here, using a novel tool that allows cross-species comparisons of regulatory elements between jawed and jawless vertebrates, we report deep conservation of both upstream regulators and segmental activity of enhancer elements across these distant species. Regulatory regions from diverse gnathostomes drive segmental reporter expression in the lamprey hindbrain and require the same transcriptional inputs (for example, Kreisler (also known as Mafba), Krox20 (also known as Egr2a)) in both lamprey and zebrafish. We find that lamprey hox genes display dynamic segmentally restricted domains of expression; we also isolated a conserved exonic hox2 enhancer from lamprey that drives segmental expression in rhombomeres 2 and 4. Our results show that coupling of Hox gene expression to segmentation of the hindbrain is an ancient trait with origin at the base of vertebrates that probably led to the formation of rhombomeric compartments with an underlying Hox code."
- Hoxb1b controls oriented cell division, cell shape and microtubule dynamics in neural tube morphogenesis[8] "Hox genes are classically ascribed to function in patterning the anterior-posterior axis of bilaterian animals; however, their role in directing molecular mechanisms underlying morphogenesis at the cellular level remains largely unstudied. ...Hoxb1b regulates mitotic spindle rotation during the oriented neural keel symmetric mitoses that are required for normal neural tube lumen formation in the zebrafish."
- Developmental Dynamics special issue January 2014 Hox/Tale transcription factors in development and disease
|
| More recent papers
|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
- This search now requires a manual link as the original PubMed extension has been disabled.
- The displayed list of references do not reflect any editorial selection of material based on content or relevance.
- References also appear on this list based upon the date of the actual page viewing.
References listed on the rest of the content page and the associated discussion page (listed under the publication year sub-headings) do include some editorial selection based upon both relevance and availability.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References
Search term: Hox
<pubmed limit=5>Hox</pubmed>
|
| Older papers
|
- Evolution of anterior Hox regulatory elements among chordates[9] "The Hox family of transcription factors has a fundamental role in segmentation pathways and axial patterning of embryonic development and their clustered organization is linked with the regulatory mechanisms governing their coordinated expression along embryonic axes. Among chordates, of particular interest are the Hox paralogous genes in groups 1-4 since their expression is coupled to the control of regional identity in the anterior nervous system, where the highest structural diversity is observed. ...Together, our results indicate that during chordate evolution, cis-elements dependent upon Hox/Pbx regulatory complexes, are responsible for key aspects of segmental Hox expression in neural tissue and appeared with urochordates after cephalochordate divergence."
- Hox10 Genes Function in Kidney Development in the Differentiation and Integration of the Cortical Stroma [10] "Consistent with loss of cortical stromal cell function, Hox10 mutant kidneys display reduced and aberrant ureter branching, decreased nephrogenesis. These data therefore provide critical novel insights into the cellular and genetic mechanisms governing cortical cell development during kidney organogenesis. These results, combined with previous evidence demonstrating that Hox11 genes are necessary for patterning the metanephric mesenchyme, support a model whereby distinct populations in the nephrogenic cord are regulated by unique Hox codes, and that differential Hox function along the AP axis of the nephrogenic cord is critical for the differentiation and integration of these cell types during kidney organogenesis."
- Proposed Hox protein classification[1]"Our classification scheme offers a higher-resolution classification that is in accordance with phylogenetic as well as experimental data and, thereby, provides a novel basis for experiments, such as comparative and functional analyses of Hox-proteins."
- Homeobox A7 up-regulation of epidermal growth factor receptor expression in human granulosa cells[11]"Our present study reveals a novel mechanistic role for HOXA7 in modulating granulosa cell proliferation via the regulation of EGFR. This finding contributes to the knowledge of the pro-proliferation effect of HOXA7 in granulosa cell growth and differentiation."
- Hoxa5 transcriptional complexity in the mouse embryo[4]Our observation that the Hoxa5 larger transcripts possess a developmentally-regulated expression combined to the increasing sum of data on the role of long noncoding RNAs in transcriptional regulation suggest that the Hoxa5 larger transcripts may participate in the control of Hox gene expression.
|
Classification
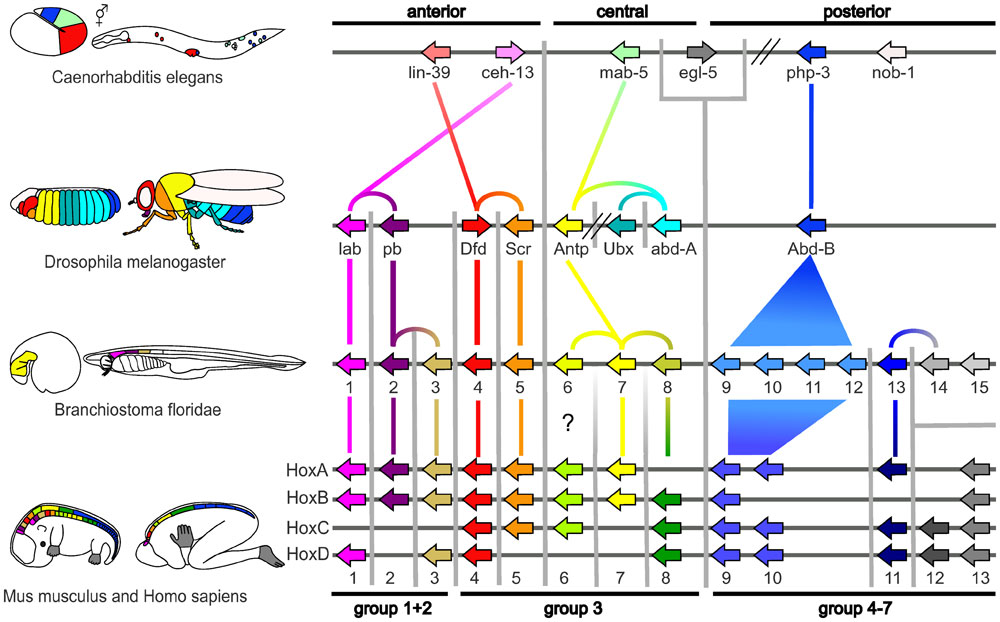
|
| Proposed Hox protein classification[1]
|
Human Hox

Chromosomal Distribution of Human Homeobox Genes[12]
Functions
Developmental patterning signal.
Neural
Segmentation
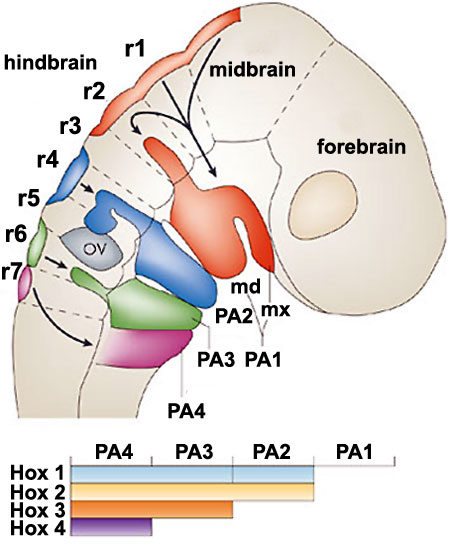
|
Hindbrain neural crest migration and Hox expression pattern[13]
A schematic diagram of a chick head at embryonic day two (Hamburger Hamilton Stages), showing pathways of neural crest migration in the chick and mouse embryo and patterns of Hox gene expression in the pharyngeal arches. Hox genes are expressed in neural crest cells, which emigrate predominantly from even-numbered rhombomeres into the pharyngeal (branchial) arches generating skeletal tissues and cranial ganglia.
Note that the first pharyngeal arch is free of Hox expression.
Legend
- PA - pharyngeal arch
- Md - mandibular part of pharyngeal arch 1
- Mx - maxillary part of pharyngeal arch 1
- OV - otic vesicle
- r - rhombomere
Adapted by permission from Macmillan Publishers Ltd: Nature Reviews Neuroscience (<pubmed>17948031</pubmed>), copyright (2007)
|
- Links: Neural System Development | Neural Crest Development
Phrenic Motor Neurons
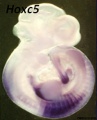
Hox5 (Hoxa5 and Hoxc5) required for phrenic motor column (PMC) development that form the respiratory motor neurons driving the diaphragm for respiration.[14]
- Links: Respiratory System Development
Axial Skeleton
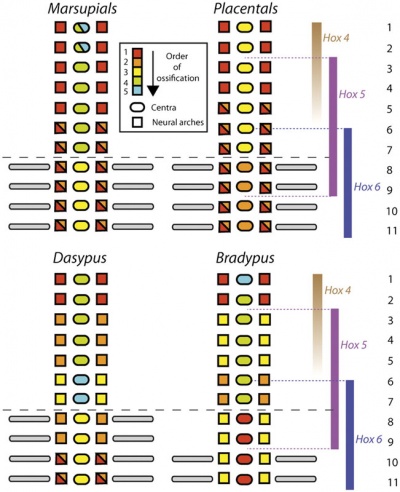
Vertebral element ossification between species.[15]
|
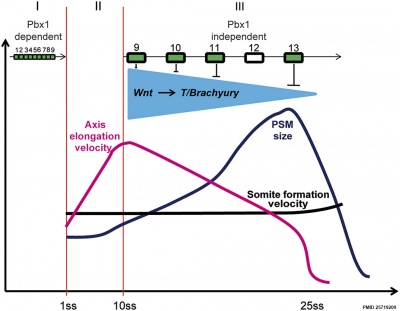
Chicken model of Hox in paraxial mesoderm precursors in the epiblast/tail-bud during axis elongation[16]
|
- Links: Somitogenesis | Axial Skeleton Development
Limb
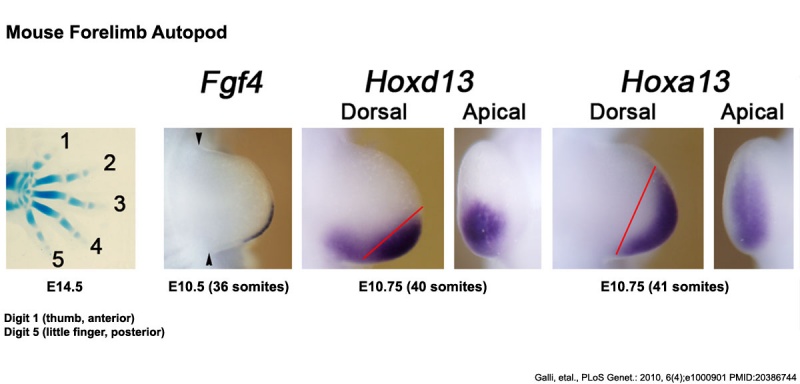
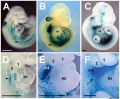
Mouse Limb Patterning Fgf and Hox Expression[17]
Fgf and Hox expression in E10.5 to 10.75 wild-type embryonic forelimb autopod, compared to future E14.5 digit arrangement.
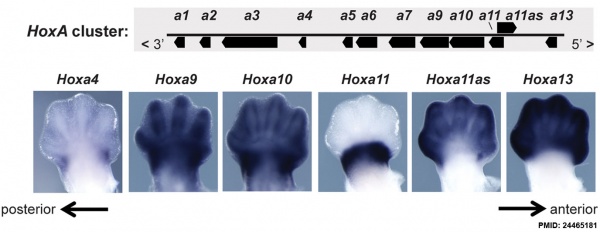
Expression of Hoxa4, Hoxa9, Hoxa10, Hoxa11, Hoxa11 antisense (Hoxa11as), and Hoxa13 in E12.5 limb buds.[18]
- Links: Limb Development
Other
Signaling Pathway
Otx
Otx is a txanscription factor essential for the normal development of the brain, cerebellum, pineal gland, and eye.
OTX is a homeobox family gene related to a gene expressed in the developing Drosophila head termed 'orthodenticle.' OTX transcription factors bind with high affinity to TAATCC/T elements on DNA.
- OMIM: OTX1 | OTX2 | OTX3
- Links: Pineal | Vision | PubMed - Otx
Additional Images
Hox deuterostomes phylogenetic tree PMID 23819519
Mouse forelimb bud Fgf4-Hoxd13-Hoxa13
Mouse spleen Hox11 expression
References
- ↑ 1.0 1.1 1.2 <pubmed>20520839</pubmed>| PLoS One
- ↑ Meinhardt H. Models For Biological Pattern Formation. Academic Press; 1982.
- ↑ <pubmed>25785959</pubmed>
- ↑ 4.0 4.1 <pubmed>20485555</pubmed>| PLoS One
- ↑ <pubmed>26829752</pubmed>
- ↑ <pubmed>26024448</pubmed>| PLoS One.
- ↑ <pubmed>25219855</pubmed>
- ↑ <pubmed>24449840</pubmed>
- ↑ <pubmed>22085760</pubmed>
- ↑ Yallowitz AR, Hrycaj SM, Short KM, Smyth IM, Wellik DM (2011) Hox10 Genes Function in Kidney Development in the Differentiation and Integration of the Cortical Stroma. PLoS ONE 6(8): e23410. PMID 21858105 PloS One
- ↑ <pubmed>20540809</pubmed>| Reprod Biol Endocrinol.
- ↑ <pubmed>17963489</pubmed>| BMC Biol.
- ↑ <pubmed>17948031</pubmed>
- ↑ <pubmed>23103965/pubmed>
- ↑ <pubmed>20956304</pubmed>| Proc Natl Acad Sci U S A.
- ↑ <pubmed>25719209</pubmed>| Elife.
- ↑ <pubmed>20386744</pubmed>| PMC2851570 | PLoS Genet.
- ↑ <pubmed>24465181</pubmed>| PLoS Biol.
Reviews
<pubmed>20435029</pubmed>
<pubmed>19651304</pubmed>
<pubmed>15944185</pubmed>
<pubmed>11604126</pubmed>
Articles
Search Pubmed
Search Bookshelf hox
July 2010 "hox" All (3509) Review (545) Free Full Text (1453)
Search Pubmed Now: Hox Homeobox
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 23) Embryology Developmental Signals - Homeobox. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Homeobox
- What Links Here?
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G