Developmental Signals - Hippo: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 26: | Line 26: | ||
| | | | ||
* '''Oocyte-expressed yes-associated protein is a key activator of the early zygotic genome in mouse'''<ref name="PMID26902285"><pubmed>26902285</pubmed></ref> "In early mammalian embryos, the genome is transcriptionally quiescent until the zygotic genome activation (ZGA) which occurs 2-3 days after fertilization. Despite a long-standing effort, maternal transcription factors regulating this crucial developmental event remain largely elusive. Here, using maternal and paternal mouse models of Yap1 deletion, we show that maternally accumulated yes-associated protein (YAP) in oocyte is essential for ZGA. Maternal Yap1-knockout embryos exhibit a prolonged two-cell stage and develop into the four-cell stage at a much slower pace than the wild-type controls. Transcriptome analyses identify YAP target genes in early blastomeres; two of which, Rpl13 and Rrm2, are required to mediate maternal YAP's effect in conferring developmental competence on preimplantation embryos. Furthermore, the physiological YAP activator, lysophosphatidic acid, can substantially improve early development of wild-type, but not maternal Yap1-knockout embryos in both oviduct and culture." | * '''Oocyte-expressed yes-associated protein is a key activator of the early zygotic genome in mouse'''<ref name="PMID26902285"><pubmed>26902285</pubmed></ref> "In early mammalian embryos, the genome is transcriptionally quiescent until the zygotic genome activation (ZGA) which occurs 2-3 days after fertilization. Despite a long-standing effort, maternal transcription factors regulating this crucial developmental event remain largely elusive. Here, using maternal and paternal mouse models of Yap1 deletion, we show that maternally accumulated yes-associated protein (YAP) in oocyte is essential for ZGA. Maternal Yap1-knockout embryos exhibit a prolonged two-cell stage and develop into the four-cell stage at a much slower pace than the wild-type controls. Transcriptome analyses identify YAP target genes in early blastomeres; two of which, Rpl13 and Rrm2, are required to mediate maternal YAP's effect in conferring developmental competence on preimplantation embryos. Furthermore, the physiological YAP activator, lysophosphatidic acid, can substantially improve early development of wild-type, but not maternal Yap1-knockout embryos in both oviduct and culture." | ||
* '''Review - Position- and polarity-dependent Hippo signaling regulates cell fates in preimplantation mouse embryos'''<ref name="PMID25986053"><pubmed>25986053</pubmed></ref> "During the preimplantation stage, mouse embryos establish two cell lineages by the time of early blastocyst formation: the trophectoderm (TE) and the inner cell mass (ICM). Historical models have proposed that the establishment of these two lineages depends on the cell position within the embryo (e.g., the positional model) or cell polarization along the apicobasal axis (e.g., the polarity model). Recent findings have revealed that the Hippo signaling pathway plays a central role in the cell fate-specification process: active and inactive Hippo signaling in the inner and outer cells promote ICM and TE fates, respectively. Intercellular adhesion activates, while apicobasal polarization suppresses Hippo signaling, and a combination of these processes determines the spatially regulated activation of the Hippo pathway in 32-cell-stage embryos. Therefore, there is experimental evidence in favor of both positional and polarity models. At the molecular level, phosphorylation of the Hippo-pathway component angiomotin at adherens junctions (AJs) in the inner (apolar) cells activates the Lats protein kinase and triggers Hippo signaling. In the outer cells, however, cell polarization sequesters Amot from basolateral AJs and suppresses activation of the Hippo pathway. Other mechanisms, including asymmetric cell division and Notch signaling, also play important roles in the regulation of embryonic development. In this review, I discuss how these mechanisms cooperate with the Hippo signaling pathway during cell fate-specification processes." [[Developmental Signals - Notch]] | [[Mouse Development]] | * '''Review - Position- and polarity-dependent Hippo signaling regulates cell fates in preimplantation mouse embryos'''<ref name="PMID25986053"><pubmed>25986053</pubmed></ref> "During the preimplantation stage, mouse embryos establish two cell lineages by the time of early blastocyst formation: the trophectoderm (TE) and the inner cell mass (ICM). Historical models have proposed that the establishment of these two lineages depends on the cell position within the embryo (e.g., the positional model) or cell polarization along the apicobasal axis (e.g., the polarity model). Recent findings have revealed that the Hippo signaling pathway plays a central role in the cell fate-specification process: active and inactive Hippo signaling in the inner and outer cells promote ICM and TE fates, respectively. Intercellular adhesion activates, while apicobasal polarization suppresses Hippo signaling, and a combination of these processes determines the spatially regulated activation of the Hippo pathway in 32-cell-stage embryos. Therefore, there is experimental evidence in favor of both positional and polarity models. At the molecular level, phosphorylation of the Hippo-pathway component angiomotin at adherens junctions (AJs) in the inner (apolar) cells activates the Lats protein kinase and triggers Hippo signaling. In the outer cells, however, cell polarization sequesters Amot from basolateral AJs and suppresses activation of the Hippo pathway. Other mechanisms, including asymmetric cell division and Notch signaling, also play important roles in the regulation of embryonic development. In this review, I discuss how these mechanisms cooperate with the Hippo signaling pathway during cell fate-specification processes." [[Developmental Signals - Notch]] | [[Mouse Development]] | ||
* '''YAP is essential for tissue tension to ensure vertebrate 3D body shape'''<ref name="PMID25778702"><pubmed>25778702</pubmed></ref> "Vertebrates have a unique 3D body shape in which correct tissue and organ shape and alignment are essential for function. For example, vision requires the lens to be centred in the eye cup which must in turn be correctly positioned in the head. Tissue morphogenesis depends on force generation, force transmission through the tissue, and response of tissues and extracellular matrix to force. Although a century ago D'Arcy Thompson postulated that terrestrial animal body shapes are conditioned by gravity, there has been no animal model directly demonstrating how the aforementioned mechano-morphogenetic processes are coordinated to generate a body shape that withstands gravity. Here we report a unique medaka fish (Oryzias latipes) mutant, hirame (hir), which is sensitive to deformation by gravity. hir embryos display a markedly flattened body caused by mutation of YAP, a nuclear executor of Hippo signalling that regulates organ size. We show that actomyosin-mediated tissue tension is reduced in hir embryos, leading to tissue flattening and tissue misalignment, both of which contribute to body flattening. By analysing YAP function in 3D spheroids of human cells, we identify the Rho GTPase activating protein ARHGAP18 as an effector of YAP in controlling tissue tension. Together, these findings reveal a previously unrecognised function of YAP in regulating tissue shape and alignment required for proper 3D body shape." [[Medaka Development]] | * '''YAP is essential for tissue tension to ensure vertebrate 3D body shape'''<ref name="PMID25778702"><pubmed>25778702</pubmed></ref> "Vertebrates have a unique 3D body shape in which correct tissue and organ shape and alignment are essential for function. For example, vision requires the lens to be centred in the eye cup which must in turn be correctly positioned in the head. Tissue morphogenesis depends on force generation, force transmission through the tissue, and response of tissues and extracellular matrix to force. Although a century ago D'Arcy Thompson postulated that terrestrial animal body shapes are conditioned by gravity, there has been no animal model directly demonstrating how the aforementioned mechano-morphogenetic processes are coordinated to generate a body shape that withstands gravity. Here we report a unique medaka fish (Oryzias latipes) mutant, hirame (hir), which is sensitive to deformation by gravity. hir embryos display a markedly flattened body caused by mutation of YAP, a nuclear executor of Hippo signalling that regulates organ size. We show that actomyosin-mediated tissue tension is reduced in hir embryos, leading to tissue flattening and tissue misalignment, both of which contribute to body flattening. By analysing YAP function in 3D spheroids of human cells, we identify the Rho GTPase activating protein ARHGAP18 as an effector of YAP in controlling tissue tension. Together, these findings reveal a previously unrecognised function of YAP in regulating tissue shape and alignment required for proper 3D body shape." [[Medaka Development]] | ||
|} | |} | ||
| Line 67: | Line 69: | ||
:'''Links:''' [[Blastocyst]] | :'''Links:''' [[Blastocyst]] | ||
==Bone== | |||
Hippo signaling pathway has recently been identified as a key regulator of osteoclast formation, see review.{{#pmid:29219182|PMID29219182}} | |||
* Hippo signaling pathway regulatory molecules - RASSF2, NF2, MST1/2, SAV1, LATS1/2, MOB1, YAP, and TAZ. | |||
* osteoclast differentiation - upon activation, MST and LAST, transcriptional co-activators YAP and TAZ bind to the members of the TEA domain (TEAD) family transcription factors | |||
** regulate expression of downstream target genes connective tissue growth factor (CTGF/CCN2) and cysteine-rich protein 61 (CYR61/CCN1). | |||
* RANKL-mediated signaling cascades including NF-κB, MAPKs, AP1, and NFATc1, Hippo-signaling molecules such as YAP/TAZ/TEAD complex, RASSF2, MST2, and Ajuba could also potentially modulate osteoclast differentiation and function. | |||
:'''Links:''' [[Musculoskeletal System - Bone Development|Bone Development]] | |||
==References== | ==References== | ||
Revision as of 12:04, 12 March 2018
| Embryology - 19 Apr 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
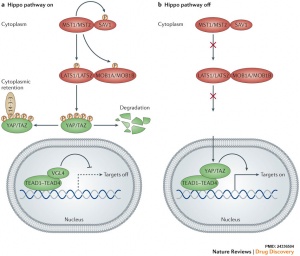
The Hippo (Hpo) pathway, first identified in Drosophila, controls organ size by regulating cell proliferation (inhibition) and apoptosis (induction). In contrast, the TOR signalling pathway regulates organ size by stimulating cell growth, thus increasing cell size.
Hippo is a protein kinase cascade pathway, getting its name from the “hippopotamus”-like fly phenotype.
| Fly Phenotype (dorsal view head thorax SEM) | |
|---|---|
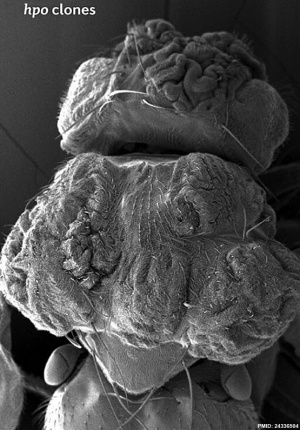
|

|
| Hippo-type (hpo) | Wild-type (WT) |
| Image source[1] |
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Embryo Hippo | Images <pubmed limit=5>Embryo Hippo</pubmed> |
Early Development
Zygote
Maternally inherited Yes-associated protein (Yap), a co-activator of TEAD family transcription factors, plays a key role in activating embryonic transcription following fertilization in the mouse. Lysophosphatidic acid (LPA) in the mouse tubal fluid binds to its G-protein coupled receptor at the plasma membrane, and induces the activation of YAP by inhibiting LATS1/2. [2]
- Links: Zygote | Mouse Development
Blastocyst
| Mouse blastocyst (32 cell stage) | |
|---|---|
| phosphorylation of angiomotin at adherens junctions | cell polarization sequesters Amot from basolateral adherens junctions |
| active Hippo signaling | inactive Hippo signaling |
| inner cells | outer cells |
| inner cell mass (ICM) fate | trophectoderm (TE) fate |
- (Table data see review[3] Notch signaling also has a role in blastocyst fate development)
- Links: Blastocyst
Bone
Hippo signaling pathway has recently been identified as a key regulator of osteoclast formation, see review.[5]
- Hippo signaling pathway regulatory molecules - RASSF2, NF2, MST1/2, SAV1, LATS1/2, MOB1, YAP, and TAZ.
- osteoclast differentiation - upon activation, MST and LAST, transcriptional co-activators YAP and TAZ bind to the members of the TEA domain (TEAD) family transcription factors
- regulate expression of downstream target genes connective tissue growth factor (CTGF/CCN2) and cysteine-rich protein 61 (CYR61/CCN1).
- RANKL-mediated signaling cascades including NF-κB, MAPKs, AP1, and NFATc1, Hippo-signaling molecules such as YAP/TAZ/TEAD complex, RASSF2, MST2, and Ajuba could also potentially modulate osteoclast differentiation and function.
- Links: Bone Development
References
- ↑ 1.0 1.1 <pubmed>24336504</pubmed>| Nat Rev Drug Discov.
- ↑ 2.0 2.1 <pubmed>26902285</pubmed>
- ↑ 3.0 3.1 <pubmed>25986053</pubmed>
- ↑ <pubmed>25778702</pubmed>
- ↑ Yang W, Han W, Qin A, Wang Z, Xu J & Qian Y. (2018). The emerging role of Hippo signaling pathway in regulating osteoclast formation. J. Cell. Physiol. , 233, 4606-4617. PMID: 29219182 DOI.
Reviews
<pubmed></pubmed> <pubmed></pubmed> <pubmed>26032720</pubmed> <pubmed>22575479</pubmed> <pubmed>21808241</pubmed> <pubmed>19517570</pubmed>
Articles
<pubmed></pubmed> <pubmed></pubmed> <pubmed>26416966</pubmed> <pubmed>25628125</pubmed> <pubmed>25918243</pubmed>
Search Pubmed
Search Bookshelf Hippo
Search Pubmed Now: Hippo
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 19) Embryology Developmental Signals - Hippo. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Hippo
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
