Developmental Signals - Fox: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 5: | Line 5: | ||
''Draft page.'' | ''Draft page.'' | ||
{{Factor Links}} | {{Factor Links}} | ||
==Some Recent Findings== | ==Some Recent Findings== | ||
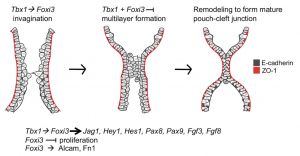
[[File:Pharyngeal arch segmentation model - Tbx and Fox.jpg|thumb|Pharyngeal arch segmentation model Tbx1 and Foxi3{{#pmid:31412026|PMID31412026}}]] | |||
{| | {| | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* '''{{Tbx}}1 and {{Fox}}i3 genetically interact in the pharyngeal pouch endoderm in a mouse model for 22q11.2 deletion syndrome'''{{#pmid:31412026|PMID31412026}} "We investigated whether Tbx1, the gene for 22q11.2 deletion syndrome (22q11.2DS) and Foxi3, both required for segmentation of the pharyngeal apparatus (PA) to individual arches, genetically interact. We found that all Tbx1+/-;Foxi3+/- double heterozygous mouse embryos had {{thymus}} and {{parathyroid}} gland defects, similar to those in 22q11.2DS patients.... Several genes expressed in the PA epithelia were downregulated in both {{Tbx}}1 and {{Fox}}i3 null mutant embryos including Notch pathway genes Jag1, Hes1, and Hey1, suggesting that they may, along with other genes, act downstream to explain the observed genetic interaction. We found Alcam and Fibronectin extracellular matrix proteins were reduced in expression in Foxi3 null but not Tbx1 null embryos, suggesting that some, but not all of the downstream mechanisms are shared." [https://www.omim.org/entry/6020547 OMIM - Tbx1] | [https://www.omim.org/entry/612351 OMIM - Foxi3] | |||
* '''Role of forkhead box gene family in bone metabolism'''{{#pmid:31549399|PMID31549399}} "Bone metabolism is associated with many bone diseases and regulated by multiple signal pathways. Over the past three decades, the functions of a superfamily of evolutionarily conserved transcriptional regulators, known as forkhead box (Fox) family, has been demonstrated to contribute to the bone metabolism. Genetic analysis studies have demonstrated that Fox gene family participate in bone metabolism and that their expression can be regulated by multiple factors. The deregulation of Fox gene family can lead to a series of bone metabolic diseases. In this manuscript, we sketched the biology of the Foxs family, summarized its function of regulating bone metabolism and maintaining bone homeostasis to estimate its potential therapeutic effects in bone diseases, and suggested directions for future exploration in this important field." | * '''Role of forkhead box gene family in bone metabolism'''{{#pmid:31549399|PMID31549399}} "Bone metabolism is associated with many bone diseases and regulated by multiple signal pathways. Over the past three decades, the functions of a superfamily of evolutionarily conserved transcriptional regulators, known as forkhead box (Fox) family, has been demonstrated to contribute to the bone metabolism. Genetic analysis studies have demonstrated that Fox gene family participate in bone metabolism and that their expression can be regulated by multiple factors. The deregulation of Fox gene family can lead to a series of bone metabolic diseases. In this manuscript, we sketched the biology of the Foxs family, summarized its function of regulating bone metabolism and maintaining bone homeostasis to estimate its potential therapeutic effects in bone diseases, and suggested directions for future exploration in this important field." | ||
Revision as of 01:52, 10 March 2020
| Embryology - 25 Apr 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
A protein transcription factor belonging to the evolutionarily conserved forkhead box (FOX) superfamily.
Draft page.
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Fox |
| Older papers |
|---|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
|
Transcription Factor
Abnormalities
Associated with defects in each Fox protein or their signaling pathway.
References
- ↑ 1.0 1.1 Hasten E & Morrow BE. (2019). Tbx1 and Foxi3 genetically interact in the pharyngeal pouch endoderm in a mouse model for 22q11.2 deletion syndrome. PLoS Genet. , 15, e1008301. PMID: 31412026 DOI.
- ↑ Huang J, Shen G, Ren H, Zhang Z, Yu X, Zhao W, Shang Q, Cui J, Yu P, Peng J, Liang Z, Yang X & Jiang. (2020). Role of forkhead box gene family in bone metabolism. J. Cell. Physiol. , 235, 1986-1994. PMID: 31549399 DOI.
- ↑ Castells-Nobau A, Eidhof I, Fenckova M, Brenman-Suttner DB, Scheffer-de Gooyert JM, Christine S, Schellevis RL, van der Laan K, Quentin C, van Ninhuijs L, Hofmann F, Ejsmont R, Fisher SE, Kramer JM, Sigrist SJ, Simon AF & Schenck A. (2019). Conserved regulation of neurodevelopmental processes and behavior by FoxP in Drosophila. PLoS ONE , 14, e0211652. PMID: 30753188 DOI.
- ↑ Shen J, Isaacson D, Cao M, Sinclair A, Cunha GR & Baskin L. (2018). Immunohistochemical expression analysis of the human fetal lower urogenital tract. Differentiation , 103, 100-119. PMID: 30287094 DOI.
- ↑ Maeda J, Yamagishi H, McAnally J, Yamagishi C & Srivastava D. (2006). Tbx1 is regulated by forkhead proteins in the secondary heart field. Dev. Dyn. , 235, 701-10. PMID: 16444712 DOI.
Search Bookshelf Fox
Reviews
Golson ML & Kaestner KH. (2016). Fox transcription factors: from development to disease. Development , 143, 4558-4570. PMID: 27965437 DOI.
Ramezani A, Nikravesh H & Faghihloo E. (2019). The roles of FOX proteins in virus-associated cancers. J. Cell. Physiol. , 234, 3347-3361. PMID: 30362516 DOI.
Fortin J, Ongaro L, Li Y, Tran S, Lamba P, Wang Y, Zhou X & Bernard DJ. (2015). Minireview: Activin Signaling in Gonadotropes: What Does the FOX say… to the SMAD?. Mol. Endocrinol. , 29, 963-77. PMID: 25942106 DOI.
Thackray VG. (2014). Fox tales: regulation of gonadotropin gene expression by forkhead transcription factors. Mol. Cell. Endocrinol. , 385, 62-70. PMID: 24099863 DOI.
Articles
Search Pubmed
Search Pubmed Now: Fox
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- OMIM -
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 25) Embryology Developmental Signals - Fox. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Fox
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
