Developmental Signals - Bone Morphogenetic Protein: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 70: | Line 70: | ||
Bmp2, Bmp4 and Bmp7 are co-required in the mouse AER for normal digit patterning but not limb outgrowth<ref name=PMID22662233><pubmed>22662233</pubmed>| [http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0037826 PLoS One.]</ref> | Bmp2, Bmp4 and Bmp7 are co-required in the mouse AER for normal digit patterning but not limb outgrowth<ref name=PMID22662233><pubmed>22662233</pubmed>| [http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0037826 PLoS One.]</ref> | ||
:"Outgrowth and patterning of the vertebrate limb requires a functional apical ectodermal ridge (AER). The AER is a thickening of ectodermal tissue located at the distal end of the limb bud. Loss of this structure, either through genetic or physical manipulations results in truncation of the limb. A number of genes, including Bmps, are expressed in the AER. Previously, it was shown that removal of the BMP receptor Bmpr1a specifically from the AER resulted in complete loss of hindlimbs suggesting that Bmp signaling in the AER is required for limb outgrowth. In this report, we genetically removed the three known AER-expressed Bmp ligands, Bmp2, Bmp4 and Bmp7 from the AER of the limb bud using floxed conditional alleles and the Msx2-cre allele. Surprisingly, only defects in digit patterning and not limb outgrowth were observed. In triple mutants, the anterior and posterior AER was present but loss of the central region of the AER was observed. These data suggest that Bmp ligands expressed in the AER are not required for limb outgrowth but instead play an essential role in maintaining the AER and patterning vertebrate digits." | :"Outgrowth and patterning of the vertebrate limb requires a functional apical ectodermal ridge (AER). The AER is a thickening of ectodermal tissue located at the distal end of the limb bud. Loss of this structure, either through genetic or physical manipulations results in truncation of the limb. A number of genes, including Bmps, are expressed in the AER. Previously, it was shown that removal of the BMP receptor Bmpr1a specifically from the AER resulted in complete loss of hindlimbs suggesting that Bmp signaling in the AER is required for limb outgrowth. In this report, we genetically removed the three known AER-expressed Bmp ligands, Bmp2, Bmp4 and Bmp7 from the AER of the limb bud using floxed conditional alleles and the Msx2-cre allele. Surprisingly, only defects in digit patterning and not limb outgrowth were observed. In triple mutants, the anterior and posterior AER was present but loss of the central region of the AER was observed. These data suggest that Bmp ligands expressed in the AER are not required for limb outgrowth but instead play an essential role in maintaining the AER and patterning vertebrate digits." | ||
[[File:Limb AER BMP expression01.jpg|800px]] | |||
Revision as of 08:37, 8 April 2016
| Embryology - 24 Apr 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
Belongs to the transforming growth factor-beta (TGFB) superfamily. The proteins are synthesized as prepropeptides, then cleaved, and then processed into dimeric proteins.
- TGFB family members: TGFB1, TGFB, TGFB3, bone morphogenetic proteins Bmp-2A, Bmp-2B, Bmp-3, and Bmp-6. mullerian inhibitory substance.
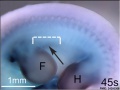
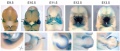
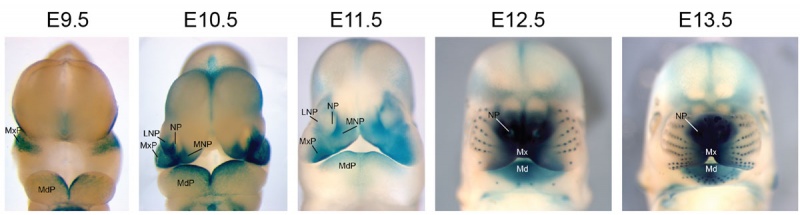
Mouse Bmp4 expression face.[1]
- BMP Mouse Links: Face and limb E9.5-13.5 | Face E9.5-13.5 | Body E11.0 | Body E11.5 | BMP | Mouse Development
Growth Differentiation Factor-6 (Gdf6) is a member of the Bone Morphogenetic Protein (BMP) family of secreted signaling molecules.
| Factor Links: AMH | hCG | BMP | sonic hedgehog | bHLH | HOX | FGF | FOX | Hippo | LIM | Nanog | NGF | Nodal | Notch | PAX | retinoic acid | SIX | Slit2/Robo1 | SOX | TBX | TGF-beta | VEGF | WNT | Category:Molecular |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Bone Morphogenetic Protein <pubmed limit=5>Bone Morphogenetic Protein</pubmed> |
Structure
Gene
Function
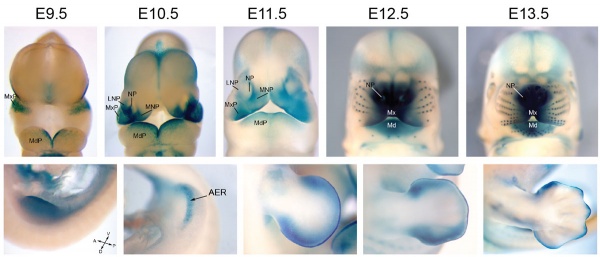
Mouse Bmp4 expression limb and face.[1] |
|
Neural Development
During gastrulation the BMP pathway is antagonised and involved with neural induction. Neural induction signaling through the BMP-regulated Smad1/5 proteins appears to be controlled by fibroblast growth factor (FGF)-regulated Ca2+ entry activating calcineurin (CaN) that in turn dephosphorylates Smad1/5 proteins.[6]
- Links: Neural Development
Oocyte Development
|
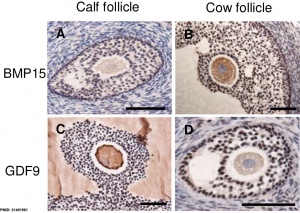
|
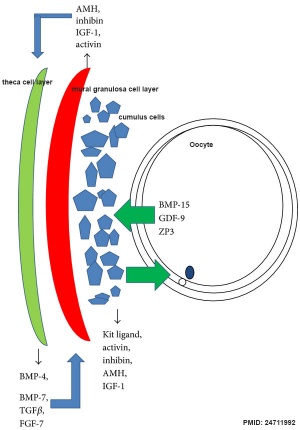
| Molecular paracrine interactions involving BMP15 signaling[7] | Localization of BMP15 in calf and cow follicles[8] |
- Links: Oocyte Development
Limb Development
Bmp2, Bmp4 and Bmp7 are co-required in the mouse AER for normal digit patterning but not limb outgrowth[9]
- "Outgrowth and patterning of the vertebrate limb requires a functional apical ectodermal ridge (AER). The AER is a thickening of ectodermal tissue located at the distal end of the limb bud. Loss of this structure, either through genetic or physical manipulations results in truncation of the limb. A number of genes, including Bmps, are expressed in the AER. Previously, it was shown that removal of the BMP receptor Bmpr1a specifically from the AER resulted in complete loss of hindlimbs suggesting that Bmp signaling in the AER is required for limb outgrowth. In this report, we genetically removed the three known AER-expressed Bmp ligands, Bmp2, Bmp4 and Bmp7 from the AER of the limb bud using floxed conditional alleles and the Msx2-cre allele. Surprisingly, only defects in digit patterning and not limb outgrowth were observed. In triple mutants, the anterior and posterior AER was present but loss of the central region of the AER was observed. These data suggest that Bmp ligands expressed in the AER are not required for limb outgrowth but instead play an essential role in maintaining the AER and patterning vertebrate digits."
- Links: Limb Development
Signaling Pathway
Identified BMP modulators:[10] Noggin, Chordin, Chordin-like 1, Chordin-like 2, Twisted gastrulation, Dan, BMPER, Sost, Sostdc1, Follistatin, Follistatin-like 1, Follistatin-like 5 and Tolloid.
Receptor
Intracellular Signaling
SNW-domain containing protein 1
(SNW1, SKI-INTERACTING PROTEIN; SKIIP)
A protein that interacts with nuclear receptors and enhances ligand-activated transcription, also called a nuclear receptor co-activator.
Regulator of Spatial BMP Activity, Neural Plate Border Formation, and Neural Crest Specification in Vertebrate Embryos[11]
- "Bone morphogenetic protein (BMP) gradients provide positional information to direct cell fate specification, such as patterning of the vertebrate ectoderm into neural, neural crest, and epidermal tissues, with precise borders segregating these domains. However, little is known about how BMP activity is regulated spatially and temporally during vertebrate development to contribute to embryonic patterning, and more specifically to neural crest formation. Through a large-scale in vivo functional screen in Xenopus for neural crest fate, we identified an essential regulator of BMP activity, SNW1. SNW1 is a nuclear protein known to regulate gene expression. Using antisense morpholinos to deplete SNW1 protein in both Xenopus and zebrafish embryos, we demonstrate that dorsally expressed SNW1 is required for neural crest specification, and this is independent of mesoderm formation and gastrulation morphogenetic movements. By exploiting a combination of immunostaining for phosphorylated Smad1 in Xenopus embryos and a BMP-dependent reporter transgenic zebrafish line, we show that SNW1 regulates a specific domain of BMP activity in the dorsal ectoderm at the neural plate border at post-gastrula stages. We use double in situ hybridizations and immunofluorescence to show how this domain of BMP activity is spatially positioned relative to the neural crest domain and that of SNW1 expression. Further in vivo and in vitro assays using cell culture and tissue explants allow us to conclude that SNW1 acts upstream of the BMP receptors. Finally, we show that the requirement of SNW1 for neural crest specification is through its ability to regulate BMP activity, as we demonstrate that targeted overexpression of BMP to the neural plate border is sufficient to restore neural crest formation in Xenopus SNW1 morphants. We conclude that through its ability to regulate a specific domain of BMP activity in the vertebrate embryo, SNW1 is a critical regulator of neural plate border formation and thus neural crest specification."
Additional Images
OMIM
About OMIM "Online Mendelian Inheritance in Man OMIM is a comprehensive, authoritative, and timely compendium of human genes and genetic phenotypes. The full-text, referenced overviews in OMIM contain information on all known mendelian disorders and over 12,000 genes. OMIM focuses on the relationship between phenotype and genotype. It is updated daily, and the entries contain copious links to other genetics resources." OMIM
- Links: OMIM300247
References
- ↑ 1.0 1.1 22701669</pubmed>| PLoS One
- ↑ <pubmed>26934293</pubmed>
- ↑ <pubmed>26113727</pubmed>
- ↑ <pubmed>24700857</pubmed>
- ↑ <pubmed>22084104</pubmed>
- ↑ <pubmed>24698271</pubmed>
- ↑ <pubmed>24711992</pubmed>| PMC3966335 | Biomed Res Int.
- ↑ <pubmed>21401961<pubmed>| Reprod Biol Endocrinol.
- ↑ <pubmed>22662233</pubmed>| PLoS One.
- ↑ <pubmed>23573253</pubmed>
- ↑ <pubmed>21358802</pubmed>
Reviews
<pubmed>26113727</pubmed> <pubmed>24174749</pubmed> <pubmed>22327483</pubmed> <pubmed>17029022</pubmed> <pubmed>15621726</pubmed>
Articles
<pubmed></pubmed> <pubmed>24698271</pubmed> <pubmed>23055827</pubmed> <pubmed>22693607</pubmed>
Online Textbooks
- Molecular Biology of the Cell. 4th edition. Alberts B, Johnson A, Lewis J, et al.New York: Garland Science; 2002. Signal Proteins of the TGF-β Superfamily Act Through Receptor Serine/Threonine Kinases and Smads
- Madame Curie Bioscience Database (Internet). Austin (TX): Landes Bioscience; 2000. TGFβ-dependent Epithelial-Mesenchymal Transition
Search PubMed
- Search PubMed: Bone Morphogenetic Protein
- All Databases: Bone Morphogenetic Protein
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- NCBI - Mouse BMP4 gene
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 24) Embryology Developmental Signals - Bone Morphogenetic Protein. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Signals_-_Bone_Morphogenetic_Protein
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G