Developmental Mechanism - Apoptosis: Difference between revisions
| Line 17: | Line 17: | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* <ref><pubmed></pubmed></ref> | * '''Who lives and who dies: Role of apoptosis in quashing developmental errors'''<ref><pubmed>21966582</pubmed></ref> "Apoptosis is essential for normal development. Large numbers of cells are eliminated by apoptosis in early neural development and during the formation of neural connections. However, our understanding of this life-or-death decision is incomplete, because it is difficult to identify dying cells by conventional strategies. Live imaging is powerful for studying apoptosis, because it can trace a death-fated cell throughout its lifetime. The Drosophila sensory organ development is a convenient system for studying neural-cell selection via lateral inhibition. We recently showed that about 20% of the differentiating neuronal cells die during sensory organ development, which results in the characteristic spatial patterning of the sensory organs. The eliminated differentiating neurons expressed neurogenic genes and high levels of activated Notch. Thus, live imaging allowed us to document the role of apoptosis in neural progenitor selection, and revealed that Notch activation is the mechanism determining which cells die during sensory organ development." | ||
|} | |} | ||
Revision as of 05:30, 14 November 2011
Introduction
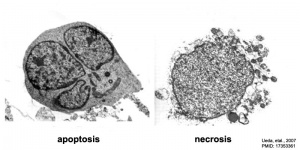
This single term "apoptosis" describes the way in which the majority of cells die within our adult body are removed every day, "Programmed Cell Death". In development, apoptosis begins in the early blastocyst and is a developmental mechanism found throughout tissues in the embryo and fetus developmental stages. In addition to the many developmental roles this process is used in multicellular organisms to remove cells that are: aged, superfluous, infected, contain genetic errors or are transformed.
While the cellular morphological changes associated with this process are the same in all cells, there are many different signaling pathways that can "trigger" this process. They fall generally into two signalling classes either intrinsic or extrinsic to the cell. To describe "programmed cell death" as apoptosis was originally used in 1972 by Kerr, Wyllie and Currie.[2]
| Cell Biology Lecture - Cell Death
Some Recent Findings
|
Developmental Examples
Digit development
Removing cells between digits (fingers and toes) of the upper and lower limbs.
Bone development
During ossification removing chondrocytes.
Neural development
Removing excess or inappropriately connected neurons.
Ovary development
Removing excess primordial follicles from the ovary cortex.
Nobel Prize 2002
The 2002 Nobel Prize in Physiology or Medicine went to three researchers who originally identified this mechanism in the genetic regulation of organ development and programmed cell death.
- Sydney Brenner (b 1927), Berkeley, CA, USA, established C. elegans as a novel experimental model organism. This provided a unique opportunity to link genetic analysis to cell division, differentiation and organ development – and to follow these processes under the microscope. Brenner's discoveries, carried out in Cambridge, UK, laid the foundation for this year's Prize.
- John Sulston (b 1942), Cambridge, England, mapped a cell lineage where every cell division and differentiation could be followed in the development of a tissue in C. elegans. He showed that specific cells undergo programmed cell death as an integral part of the normal differentiation process, and he identified the first mutation of a gene participating in the cell death process.
- Robert Horvitz (b 1947), Cambridge, MA, USA, has discovered and characterized key genes controlling cell death in C. elegans. He has shown how these genes interact with each other in the cell death process and that corresponding genes exist in humans.
- Links: Nobel Prize 2002
Apoptotic Cell Morphology
The following cellular changes occur in sequence during apoptosis.
- loss of cell membrane phospholipid asymmetry
- Condensation of chromatin
- Reduction in nuclear size JCB - Nucleus changes
- Internucleosomal DNA cleavage TUNEL staining
- DNA ladder
- shrinkage of the cell
- Cleavage of cytoskeletal proteins PNAS - Actin cleavage by ICE-like proteases
- note actin also binds DNase 1, cleavage may release this enzyme to further cleave DNA
- membrane blebbing
- breakdown of the cell into membrane-bound apoptotic bodies (apoptosomes)
- bodies then phagocytosed by other cells
Experimentally a number of different techniques have been developed and are now used to identify these changes.
- Links:
Apoptosis Regulators
Regulators can initiate or block apoptosis, the regulators shown block apoptosis.
| Regulator → | Adaptor → | Effector | |||
|---|---|---|---|---|---|
| C. elegans | Ced-9 → | Ced-4 → | Ced-3 → | Death | |
| Vertebrates | Bcl-2 → | Apaf-1 → | Caspase-9 → | Caspase-3 → | Death |
References
- ↑ <pubmed>17353361</pubmed>| PMC2064059 | J Cell Biol.
- ↑ <pubmed>4561027</pubmed>
- ↑ <pubmed>21966582</pubmed>
Textbooks
Reviews
<pubmed></pubmed>
Articles
Search PubMed
Search Pubmed: developmental apoptosis | developmental cell death | apoptosis
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- UNSW Cell Biology - Lecture - Cell Death
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 19) Embryology Developmental Mechanism - Apoptosis. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Developmental_Mechanism_-_Apoptosis
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G