Blastocyst Development
Introduction
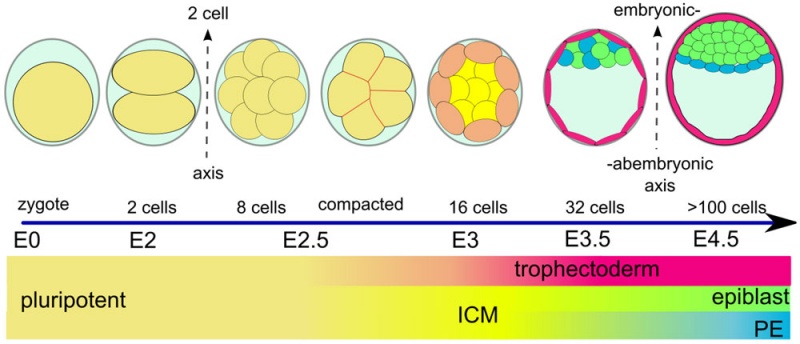
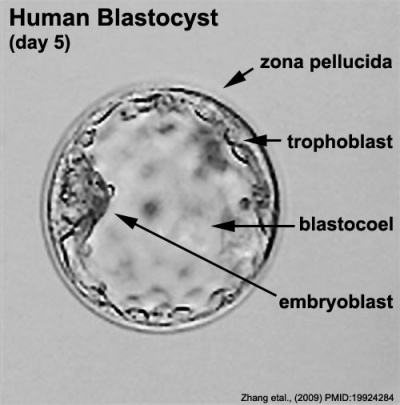
(Greek, blastos = sprout + cystos = cavity) or blastula, the term used to describe the hollow cellular mass that forms in early development. The blastocyst consists of cells forming an outer trophoblast layer, an inner cell mass and a fluid-filled cavity. The blastocyst inner cell mass is the source of true embryonic stem cells capable of forming all cell types within the embryo. In humans, this stage occurs in the first and second weeks after the zygote forms a solid cellular mass morula stage) and before implantation.
- Links: Fertilization | Week 1 | Morula | Blastocyst Development
Some Recent Findings
|
Human Blastocyst Formation
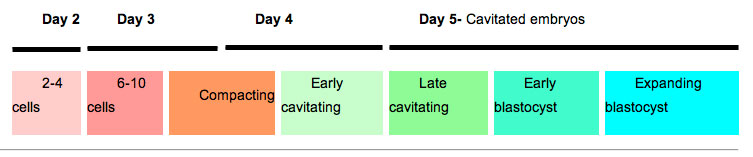
The table below shows human blastocyst in vitro development changes during week 1.[5]
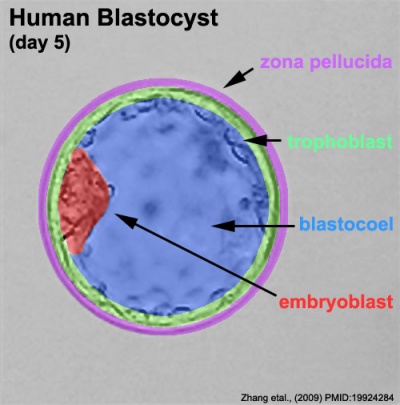
Labeled Blastocyst
Blastocyst Hatching
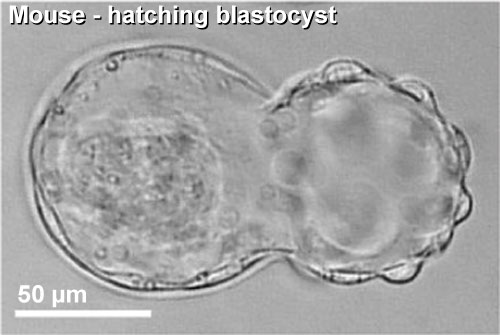
|
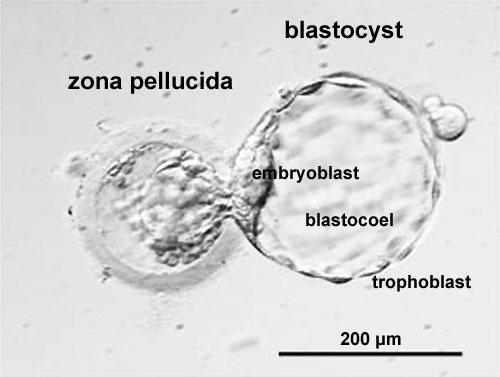
|
| Blastocyst hatching from zona pellucida (mouse) | Blastocyst hatching from zona pellucida (human) |
Movies
The following movies are edited from a recent paper on human blastocyst development (day 3-6).[2] Select the Quicktime or Flash format to view annotated movies.
|
||
| Blastocyst Development | Blastocyst Contractions | Blastocyst Hatching |
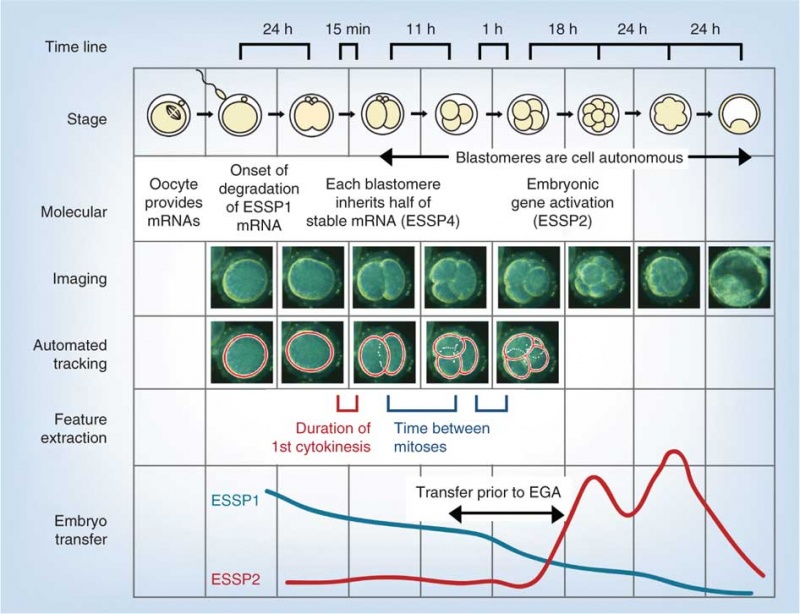
Model Human Blastocyst Development
The following figure is from a recent study[2] using video and genetic analysis of in vitro human development during week 1 following fertilization.
- EGA - embryonic genome activation
- ESSP - embryonic stage–specific pattern, four unique embryonic stage–specific patterns (1-4)
- Links: Figure with legend
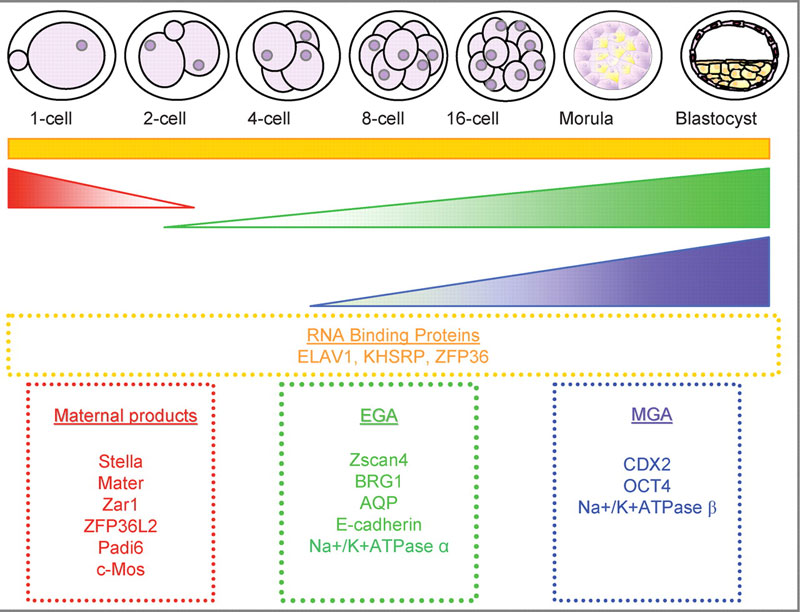
Mouse Blastocyst Gene Expression
General gene expression patterns are indicated from genomic profiling.[6]
- red - loss of maternal mRNAs
- green - activation of embryonic genome (EGA)
- purple - maternal gene activation (MGA)
- orange - continuous expression
Inner Cell Mass
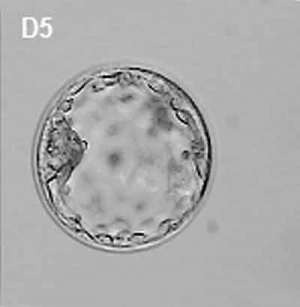
This outer layer of cells is also called the "embryoblast", a cluster of cells located and attached on one wall of the outer trophoblast layer.
Trophoblast Layer
This outer layer of cells is also called the "trophectoderm" (TE) epithelium. A key function is for the transport of sodium (Na+) and chloride (Cl-) ions through this layer into the blastocoel.
Differentiation of this layer has been shown to be regulated by the transcription factors Tead4[7] and then Caudal-related homeobox 2 (Cdx2).
- Links: Trophoblast | OMIM -Tead4 | OMIM - Cdx2
Blastocoel Formation
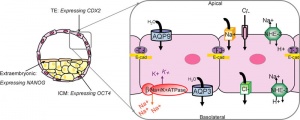
- trophectoderm transports of Na+ and Cl- ions through this layer into the blastocoel
- generates an osmotic gradient driving fluid across this epithelium
- distinct apical and basolateral membrane domains specific for transport
- facilitates transepithelial Na+ and fluid transport for blastocoel formation
- transport is driven by Na, K-adenosine triphosphatase (ATPase) in basolateral membranes of the trophectoderm [8]
Blastocyst Metabolism
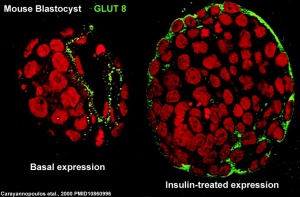
At the blastocyst stage, mammalian development metabolism switches on anaerobic glycolysis metabolism to satisfy metabolic demands of growing blastocyst and formation of the blastocoel. This is thought to be driven by the integral membrane protein family of facilitative glucose transporters (GLUT or SLC2A).
- aerobic - oxidation of lactate and pyruvate via the citric acid cycle (Krebs cycle) and oxidative phosphorylation
- glycolysis- converts glucose into pyruvate
- GLUT - GLUcose Transporter (divided into 3 classes I-III)
- SLC2 - Solute Carrier Family 2
Glucose Transporter Expression
- GLUT1 - from zygote to blastocyst. (all mammalian tissues, basal glucose uptake)
- GLUT2 and GLUT3 - from late eight cell stage to blastocyst. (GLUT2, liver and pancreatic beta cells; GLUT3, all mammalian tissues, basal glucose uptake)
- GLUT4 - not expressed. (muscle and adipose tissue)
- GLUT8 - up-regulated at blastocyst stage. (central nervous system and heart)
- (Data mainly from mouse development, adult tissue expression shown in brackets)
A mouse study,[9] has shown GLUT8 is up-regulated following insulin stimulation, though a more recent GLUT8 knockout mouse shows normal early embryonic development in the absence of this transporter.[10]
- Links: Biochemistry - glucose transporters | GLUT1 | GLUT2 | GLUT8
Blastula Cell Communication
Two forms of cellular junctions Figure 21-69. The blastula
- gap junctions, allow electrically couple cells of epithelium surrounding a fluid-filled cavity
- tight junctions, close to outer surface create a seal, isolates interior of embryo from external medium
Blastocyst Hatching - zona pellucida lost, ZP has sperm entry site, and entire ZP broken down by uterine secretions and possibly blastula secretions. Uterine Glands - secretions required for blastocyst motility and nutrition
Blastocyst Hatching
| At about day 5 the human blastocyst "hatches" out of the protective zona pellucida. This hatching allows increased growth, access to uterine nutrient secretions and blastocyst adhesion to the uterine lining. Associated with this hatching process are a series of physical contractions.
|
width=250px|height=240px|controller=true|autoplay=false</qt>
Human blastocyst contractions (day 5-6)[2] |
Molecular Factors
- TEA DNA- binding domain, these factors bind to the consensus TEA/ATTS cognate binding site[13]
- TEF-3 - renamed Tead1 and Tead4
- Tead3 - is expressed in the placental syncytiotrophoblasts
- E-cadherin - Calcium ion-dependent cell adhesion molecule, a cell membrane adhesive protein required for morula compaction
- epithin - A type II transmembrane serine protease, identified in mouse for compaction of the morula during preimplantation embryonic development. Expressed from 8-cell stage at blastomere contacts and co-localises in the morula with E-cadherin. PMID: 15848395
- Na, K-adenosine triphosphatase - A sodium potassium pump that generates an osmotic gradient for fluid flow into the blastocoel
- Zonula occludens-1 - (ZO-1) Tight junction protein involved in morula to blastocyst transformation in the mouse PMID: 18423437
Blastocyst in Other Species
Mouse Blastocyst
Sox2 expression[14]
Early gene expression[14]
Early gene expression[14]
Early gene expression[14]
Early gene expression[14]
Early mouse development model[15]
Bovine Blastocyst
References
- ↑ 1.0 1.1 <pubmed>19924284</pubmed>| PMC2773928 | PLoS One
- ↑ 2.0 2.1 2.2 2.3 <pubmed>20890283</pubmed>| Nat Biotechnol.
- ↑ <pubmed>20864103</pubmed>
- ↑ <pubmed>20147376</pubmed>
- ↑ <pubmed>10221713</pubmed>
- ↑ 6.0 6.1 <pubmed>19043080</pubmed>| Mol Hum Reprod.
- ↑ <pubmed>18083014</pubmed>
- ↑ <pubmed>16139691</pubmed>
- ↑ 9.0 9.1 <pubmed>10860996</pubmed>| PMC16542 | Proc Natl Acad Sci U S A.
- ↑ <pubmed>16705176</pubmed>
- ↑ <pubmed>14967891</pubmed>
- ↑ <pubmed>2708952 </pubmed>
- ↑ <pubmed>8702974</pubmed>
- ↑ 14.0 14.1 14.2 14.3 14.4 <pubmed>21103067</pubmed>| PMC2980489 | PLoS One.
- ↑ <pubmed>21573197</pubmed>| PMC3088645 | PLoS Comput Biol.
Reviews
<pubmed>20607796</pubmed> <pubmed>20364097</pubmed> <pubmed>17389140</pubmed>
Articles
<pubmed>20157423</pubmed> <pubmed>19289087</pubmed> <pubmed>18817772</pubmed> <pubmed>18083014</pubmed> <pubmed>16773657</pubmed>| Dev. Dyn.
Search PubMed
Search April 2010
Search Pubmed: blastocyst development | blastocoel development | inner cell mass development | trophectoderm |
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, April 24) Embryology Blastocyst Development. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Blastocyst_Development
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G
![Early gene expression[14]](/embryology/images/thumb/c/ca/Mouse-_preimplantation_gene_expression_02.jpg/120px-Mouse-_preimplantation_gene_expression_02.jpg)
![Early gene expression[14]](/embryology/images/thumb/d/d9/Mouse-_preimplantation_gene_expression_03.jpg/120px-Mouse-_preimplantation_gene_expression_03.jpg)
![Early gene expression[14]](/embryology/images/thumb/5/56/Mouse-_preimplantation_gene_expression_04.jpg/120px-Mouse-_preimplantation_gene_expression_04.jpg)
![Early gene expression[14]](/embryology/images/thumb/7/74/Mouse-_preimplantation_gene_expression_05.jpg/120px-Mouse-_preimplantation_gene_expression_05.jpg)