User:Z5020117
| Student Information (expand to read) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Individual Assessments | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Please leave this template on top of your student page as I will add your assessment items here. Beginning your online work - Working Online in this course
Click here to email Dr Mark Hill | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 1 Assessment - Researching a Topic | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
In the lab I showed you how to find the PubMed reference database and search it using a topic word. Lab 1 assessment will be for you to use this to find a research reference on "fertilization" and write a brief summary of the main finding of the paper.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 2 Assessment - Uploading an Image | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
OK you are now in a group
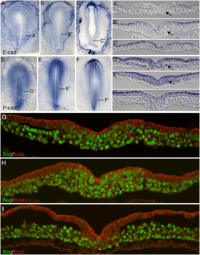
Initially the topic can be as specific or as broad as you want. Chicken embryo E-cad and P-cad gastrulation[1] References
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 4 Assessment - GIT Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
ANAT2341 Quiz Example | Category:Quiz | ANAT2341 Student 2015 Quiz Questions | Design 4 quiz questions based upon gastrointestinal tract. Add the quiz to your own page under Lab 4 assessment and provide a sub-sub-heading on the topic of the quiz. An example is shown below (open this page in view code or edit mode). Note that it is not just how you ask the question, but also how you explain the correct answer. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 5 Assessment - Course Review | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Complete the course review questionnaire and add the fact you have completed to your student page. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 6 Assessment - Cleft Lip and Palate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 7 Assessment - Muscular Dystrophy | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 8 Assessment - Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| A brief quiz was held in the practical class on urogenital development. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 9 Assessment - Peer Assessment | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 10 Assessment - Stem Cells | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
As part of the assessment for this course, you will give a 15 minutes journal club presentation in Lab 10. For this you will in your current student group discuss a recent (published after 2011) original research article (not a review!) on stem cell biology or technology.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 11 Assessment - Heart Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Read the following recent review article on heart repair and from the reference list identify a cited research article and write a brief summary of the paper's main findings. Then describe how the original research result was used in the review article.
<pubmed>26932668</pubmed>Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Lab Attendance
Z5020117 (talk) 14:34, 5 August 2016 (AEST) Z5020117 (talk) 14:41, 12 August 2016 (AEST) Z5020117 (talk) 14:07, 19 August 2016 (AEST) Z5020117 (talk) 14:11, 26 August 2016 (AEST) Z5020117 (talk) 13:21, 9 September 2016 (AEST) Z5020117 (talk) 15:01, 16 September 2016 (AEST) Z5020117 (talk) 13:27, 23 September 2016 (AEST) Z5020117 (talk) 13:54, 7 October 2016 (AEDT)
Lab 1 Assessment
'Preimplantation genetic screening for all 24 chromosomes by microarray comparative genomic hybridization significantly increases implantation rates and clinical pregnancy rates in patients undergoing in vitro fertilization with poor prognosis' Summary
<pubmed>27382234</pubmed>
The use of Preimplantation Genetic Screening (PGS) in association with IVF has not been prevalent due to its expensive and highly invasive nature, almost doubling the cost of IVF. Currently, morphology evaluation is predominantly used due to its non-invasive nature despite its variable efficacy. Majumdar et al. designed an experiment to evaluate an improved PGS system that analyses all 24 chromosomes. They believe the incorporation of chromosomal analysis will increase pregnancy and implantation rates in patients with poor prognosis. The twenty subjects of this study were classified into one of three groups, advanced maternal age (AMA), repeated miscarriage (RI) and recurrent implantation failure (RIF).
This study found that the transfer of only a few embryos, particularly euploid embryos, resulted in higher implantation rates in those receiving PGS in comparison to the control non-PGS group. Overall, it was found that in comparison to the traditional morphology evaluation previously used, the incorporation of PGS allows for improved outcomes following IVF even when no euploid embryos were transferred. With recent research establishing the correlation between high prevalence of aneuploidy embryos in patients with AMA and unsuccessful implantations, PGS can be used to successfully identify and eliminate the possibility of aneuploidy embryo transfer, thus allowing for increased implantation and pregnancy rates. In saying this, Majumdar et al. emphasise how successful implantation cannot be guaranteed with PGS as some pregnancy failures occur as a result of factors other than chromosomal abnormalities. Though further research is needed to consolidate the benefits of PGS, this study identified the possibility of achieving successful implantation and pregnancy in a shorter time period with fewer miscarriages when utilising PGS rather than morphology evaluation in association with IVF.
| Mark Hill 18 August 2016 - You have added the citation correctly and written a reasonable summary of the papers findings. Why would you thing PGD would improve implantation rates and clinical pregnancy rates? Note that this is not a high impact Journal, try those first for your article selections. | Assessment 5/5 |
Lab 2 Assessment
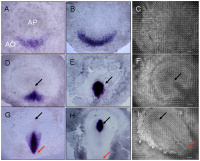
Primitive streak development in chick embryo[1]
| Mark Hill 29 August 2016 - All information Reference, Copyright and Student Image template correctly included with the file and referenced on your page here. The citation on the page here could also have appeared in the image legend as shown below. I have also added a reference sub-heading to fix the formatting issue. | Assessment 5/5 |
Lab 3 Assessment
| Mark Hill 31 August 2016 - Lab 3 Assessment Quiz - Mesoderm and Ectoderm development. | Assessment 2.5/5 |
References
Lab 4 Assessment
GIT Abnormalities Quiz
| Mark Hill 17 October 2016 - GIT quiz covers a range of topics. Question 1 tests clinical knowledge concerning intestinal malrotation. While a good question, may not test concepts on development. Question 3 tests prevalence knowledge, would be good to give more than just the original reference in the answer. Question 4 is OK. | Assessment 5/5 |
Lab 6 Assessment
Completed course questionnaire
| Mark Hill 17 October 2016 - Completed course questionnaire | Assessment 5/5 |
Cleft Palate
Genetic mutation of the transcription factor TBX22, which encodes for the DNA binding-domain, T-Box[1], causes cleft palate. [2]. It has been shown that during palatogenesis TBX22 is found within the tongue and palatal shelves, thus indicating its role in the development of both the tongue and palate. Various mutations of TBX22 can occur, including frameshift mutations resulting in the production of truncated proteins, as well as missense mutations causing a change of a single nucleotide. These mutations result in an inefficient or reduced capability of DNA to bind to the T-Box. In the case of missense mutations, it could also lead to the inability to activate transcription factors. As a result, the lack of formation of functional proteins leads to a dysfunctional palatogenesis process and thus, significantly affecting signalling in normal development to cause formation of cleft palate.
It was also found that TBX22 serves as a transcriptional repressor and modification of this repressor activity occurs through SUMO-1, a small ubiquitin-like modifier that binds upstream from the T-Box domain[3]. Therefore, mutations of SUMO-1 can also impair the function of TBX22 and can present as the craniofacial defect of X-linked cleft palate as found in many cases. Research shows that loss of function of SUMO-1 occurs as a result of exposure to an array of environmental and other factors during early pregnancy including, smoking, lack of nutritional supplements and maternal age and it is exposure to these factors that can have an effect on normal signalling in development encouraging the formation of cleft palate.
| Mark Hill 17 October 2016 - TBX22 is a factor identified by several students and your summary is useful. | Assessment 5/5 |
Lab 7 Assessment
1. What is/are the dystrophin mutation(s)?
Majority of the dystrophin mutations, approximately 60%[4], are due to deletions or insertions of nucleotides resulting in a downstream frameshift of the dystrophin gene. The remainder of dystrophin mutations are either point mutations, where there is a substitution of a single nucleotide or minor frameshift errors. These mutations can result in the complete absence or in milder forms, the alteration or reduction of the dystrophin protein.
2. What is the function of dystrophin?
The dystrophin protein found in both skeletal and cardiac muscle plays a structural role by linking the internal cytoskeleton of the muscle with the extracellular matrix. It is also responsible for protecting muscles during contraction and relaxation from injury by strengthening muscle fibres. Dystrophin also plays an additional role in cell signaling through interaction with other proteins involved in sending and receiving chemical signals. Research has shown that dystrophin may be present in minor amounts within the neurons of the brain. Thus, they may be involved in the formation of synapses[5] .
3. What other tissues/organs are affected by this disorder?
DMD can affect the respiratory muscles thus negatively impacting lung function. As the dystrophin protein is also found within cardiac muscle, the heart is also affected in DMD. Due to this dysrhythmia or arrhythmia, irregular heart rhythm and cardiomyopathy, abnormal pumping action can result. The presence of dystrophin in brain tissue can also result in learning and behavioural difficulties[6] .
4. What therapies exist for DMD?
Though there is no known cure for DMD, extensive research is being currently performed to treat DMD. The following are a few examples of research being conducted in the field of DMD.
Some of the therapies available include gene replacement therapy whereby plasmids or viruses are utilised to deliver dystrophin sequences but this is currently a work in progress. Myoblast transplantation where myoblasts are artificially delivered into the affected site can also be offered. This is because it has been shown that myoblasts can fuse to form new muscle fibres but upon exhaustion of the proliferative ability of the myoblasts, the skeletal muscle is converted into connective tissue. Unfortunately, studies have shown unsatisfactory results for such treatment. In saying this, stem-cell therapy has shown to be a good alternative to myoblast transplantation due to the extended proliferative life-span of stem cells.
Administration of aminoglycoside antibiotics is a potential therapy that targets DMD caused by premature stop codons. Results of such treatment have not been promising but have indicated that it may be more useful in only a select few DMD mutations. On the other hand, chimaeraplasts have been utilised as a vehicle to deliver the correct nucleotide to the site of dystrophin mutation. Unfortunately, the viability of such treatment is short-lived and requires further research.
Antisense oligonucleotides have been utilised to help redirect dystrophin splicing to exclude the inclusion of the premature stop codon, the most common cause of DMD. This will allow partial restoration of the reading frame and thus allow formation of the dystrophin, albeit shorter protein. Research has also shown that proteasome inhibitors can be used to improve the integrity of muscle. Lastly, upregulation therapy focuses on replacement of defective genes by increasing expression of alternative genes e.g. utrophin. These are promising areas of research in DMD therapy.
Currently, the only form of therapy available is management of DMD and this can be done through prescription of steroid medication to help maintain muscle integrity. Surgery can also be performed to release tightness of joints as well as treat scoliosis, the lateral curvature of the spine which can come as a result of DMD. Supportive equipment can also be provided including night splints, walking frames, wheelchairs, and other mobility aids. It is also important to regulate the patient’s diet and exercise routine. Muscle relaxants and anti-inflammatory medication can also be provided to help with any pain or discomfort .
5. What animal models are available for muscular dystrophy?
Currently there are two animal models that have been utilised to further understand Duchenne Muscular Dystrophy, the mdx mouse model and the golden retriever muscular dystrophy (GRMD) dog. These models are significant as both these species lack the dystrophin protein.
| Mark Hill 17 October 2016 - Very good cited answers (except the animal models?) | Assessment 5/5 |
Lab 8 Assessment
Completed the quiz on urogenital development
| Mark Hill 27 October 2016 - Very well done. Hard to give more feedback when you get all the questions correct. | Assessment 8/8 |
Lab 9 Assessment
These are very good reviews of the project pages, with some specific examples. They include a balanced critical assessment, given the existing status of some of these pages. Please use a more professional language when providing feedback. 8/10
Peer Reviews
Group 1
You guys have made significant progress on your project, managing to touch briefly on each section of your assignment. There have been some good choices of subheadings but I think some improvement can be made. For example, I think it would be useful to breakdown the general heading of ‘introduction’ into smaller subheadings so readers are made aware of what will be discussed in this section. It would also be useful to touch upon the importance of this pathway and thus, highlighting its significance in embryological development.
In terms of the content of the project, being only in the draft stage a considerable amount of editing is required. For example, there has been mention of the TCF/LEF family and though the use of this abbreviations is useful, I think it would be appropriate to initially include the full name and explain this term in brief detail. In addition, there has been discussion of the ‘canonical’ and ‘non-canonical’ pathways of WnT Signalling Pathway but you could consider discussing the significance of having these two separate pathways. Comparing and contrasting these two pathways may also assist in aiding one’s understanding of the topic.
Though it is great that you have made progress, I think more detail is required in each section, particularly in linking the effect of these pathways on embryological development. Also, greater attention needs to paid to referencing and utilisation of studies that have dissected this signalling pathway. For example, greater emphasis can be placed on studies performed on ‘embryos of Xenopus laevis’ or the in vitro experiments on mice. Instead of saying ‘a study’ or ‘another study’ acknowledge the researchers of this study as it will increase the validity of your argument while providing readers with the opportunity to refer back to these papers for more information if required or interested. More detail is also required on the effect of this pathway on skin formation. One way this could be done is by expanding on the information already provided, for example, explain how ‘WnT signalling inhibits the ectoderm’s responsiveness to FGFs’ and provide a detailed explanation of the feedback mechanism. Though your topic is focusing on ‘WnT Signalling pathway in the skin of fetus’ It would be beneficial to explore the roles of Wnt signalling in other areas of embryological development as this could provide insight into the abnormalities caused by mutations in this pathway. In terms of the ‘what can go wrong’ section, try breaking this segment into the various embryological deficiencies that can develop through disruption of the WnT pathway and try and make it relevant by providing statistics.
Overall, you guys have done a fantastic job! It was good to see that all group members had contributed to the project. The main thing that requires improvement is the lack of detail. Through editing and inclusion of appropriate references and citations you can significantly improve the quality of your work. It would be useful to add some diagrams or images to help explain the pathway. In addition, try utilising your discussion page and communicating with your other team members. By providing feedback and suggestions you can assist in efficiently producing an excellent project. I hope this helps!!
Group 2
Well done on the progress you have made thus far! You guys have chosen appropriate headings and subheadings that effectively break down the Notch signalling pathway. A coherent introduction has been provided, giving a taste of what is to be expected in this project. The use of a table to explore the history of this signalling pathway was particularly useful in making the information understandable and relevant. Though you have done an excellent job, was there any reason you stopped at 1989? It may even be useful to create a brief timeline of events, thus allowing you to better explore current areas of research by considering past studies that have been performed.
You’ve provided a good overview of the canonical pathway with the appropriate use of a diagram which aids reader’s understanding of the information provided. In saying this, I think it would be useful to expand on how this pathway is tightly controlled, is it through transcriptional regulation or through other means? In addition, it may be useful to explain the differences in the non-canonical and canonical pathways in terms of their significance and role in embryonic development. I’ve noticed that you have provided a general overview of the role of Notch signalling pathway in embryonic development, do these roles differ between the canonical and non-canonical pathways?
In addition, it’s good that you have included the role of the Notch signalling pathway in animal development as it explores the scope of this pathway beyond human embryology but it may also be useful to explore animal models in research, especially considering that the ‘first description of a “notch” defect’ was discovered in Drosophila. By combining the role of animal models in expanding our knowledge of the Notch signalling pathway with the effect of this pathway in animals, it provides a more rounded approach to explaining and discussing this signalling pathway.
I particularly like how you have included statistics in the ‘Abnormalities of Notch signalling’ section as it provides insight into the importance of this pathway in embryological development. You have successfully described the type of mutation that results in the particularly disease in most cases except for Alagille syndrome. More detail in how the mutation causes the syndrome would be useful with an explanation of how the mutation is brought about.
Overall, you guys have done a fantastic job! You have appropriately referenced and cited all the information provided and have included useful flowcharts, tables and diagrams that aid understanding of the text provided. Providing more detail to each of the sections and communicating with all your team members in the discussion page will ensure that you produce an excellent project! Good luck!
Group 3
You guys have made a good start on your project! I particularly liked how the headings were subdivided appropriately into smaller subheadings as it effectively broke down the FGFR pathway and made the page easy to navigate. Though you have included a short and succinct introduction, I think it should address all the sections being discussed to give the reader a better overview of your project. In addition, the use of a table to explore the timeline of research of the FGF pathway was an excellent idea but I think the text above the table could be incorporated into the table itself and a more extensive timeline could be provided.
Though it was good that you provided a brief overview of the FGFR pathway, you’ve only discussed the components of the pathway rather than the pathway itself. Furthermore, when discussing signal transduction, I think you should be more specific when explaining the process, for example when you mentioned ‘which leads to changes in gene transcription through interactions with DNA’, it causes changes in transcription in which genes and through interactions with which DNA? In saying this, it was wonderful to see the inclusion of a hand-drawn diagram which represents not only your understanding of the pathway but also aids readers understanding of the FGFR pathway.
A good overview has been provided to explain the role of FGFs in embryonic development. The only suggestion I can make is to provide explanations or full names of the abbreviations to aid understanding of the concepts explored. For example, what is ETV1 and EWSR1? By explaining what these abbreviations are the reader will gain better understanding on how they function to help maintain FGF10 expression. In terms of the section on abnormalities, a succinct and coherent introduction was provided. There was a good description of the morphological changes produced by these mutations along with the cause of these abnormalities. There isn’t much I would change in this section except for maybe explaining FGFR2 mutation.
Overall, you guys have done a fantastic job! I thought the inclusion of a quiz was particularly innovative as it makes your project interactive and thus, aids the learning process. Everything was well cited and referenced and it was wonderful to see the use of an original diagram. It was also good to see all groups members contributing to the discussion page which indicates effective communication within the team.
Group 4
A good start has been made to the project with the appropriate selection of headings and subheadings which provide a brief overview of what is to be discussed in terms of the Hedgehog signalling pathway. By breaking down the mechanism of the pathway, it made the foreign concept much easier to understand. In saying this, this section is quite text-heavy and may benefit with the relocation of the included diagram or even inclusion of other diagrams and flowcharts to engage readers. With the introduction of a fairly new concept, the inclusion of visual or audio stimuli and maybe even a short quiz may encourage interaction with readers.
The discussion of this pathway in mammals exposed readers to the diversity of the Hh signalling pathway but in saying this, the inclusion of a table may be useful to compare and contrast the differences between the pathways in mammals and insects. Overall, this section was well written. On the other hand, when considering the section on animal models, it provided insight into the role of Hh signalling pathway on embryological development and offered a brief introduction to the abnormalities caused by disruptions of this pathway. Once again, the inclusion of diagrams would be useful in this section to provide visual insight into the research being performed.
Though there has been significant exploration of the mechanism and animal models utilised in this pathway, more work is needed to link this pathway to embryological development and this could provide a good leeway into understanding the abnormalities associated with disruption of this pathway. This project can be significantly improved simply by focusing on making it more interactive ad engaging with the inclusion of a variety of stimuli like tables, diagrams, quizzes and even videos. In addition, all information has been well cited and referenced and there has been substantial communication between group members, allowing team members to provide feedback and suggestions thus, ultimately increasing the quality of the work produced.
Group 5
First of all, well done on making significant progress on your project! You have addressed all aspects of the pathway involving T-Box genes through subdivision into various headings and subheadings. I particularly liked how there was an inclusion of the specific T-Box gene affected in each of the abnormalities in the subheading itself. The only suggestion I would make is to combine the ‘Ancient origins and evolution of the T-Box gene family’ section with the origins of the ‘T-Box genes’ section to provide a more coherent description of the history of these pathway. You could even form a table to create a timeline of events. In addition, I think it would be beneficial to include the ‘What does T-Box mean?’ as an introduction to the ‘origins of the T-box genes’ section as there is overlap between these sections.
The use of a table to describe the main T-box genes was helpful in providing a brief overview of the components of the pathway and their influence in embryological development. In addition, the link between T-Box genes and embryonic development has been explored considerably. In saying this, greater attention to detail must be paid to explaining abbreviations to aid one’s understanding of the concepts being discussed. For example, what is NKX2-5, Shh and OFT? Though you’ve explained that RA stands for retinoic acid in the ‘Organisms used in animal models for T-Box’ section, this same explanation is not provided in the ‘Limb development’ section where you have discussed that ‘RA and Shh both induced Tbx2’. These small changes will significantly improve the quality of your work.
The inclusion of abnormalities provides great insight into the role of T-Box genes in development. In saying this, though you have explored the effect of the mutation of these genes in animal models, more information is required to explain the effect of these mutations in humans and how they come about. Furthermore, under the heading of ‘Animal models’ there has been discussion mainly of the ‘brachyury gene’ which seems unrelated to animal models due to the lack of a proper introduction. I found the following section (organisms used in animal models for T-Box) to be a better introduction to the topic of animal models. In addition, there has been mention of a number of animal models ‘Drosophila, Xenopus, zebrafish, avians, and mice’ yet only marsupials and amphioxus has been discussed. This could be potentially misleading to readers.
Overall, a fantastic effort has been made. Not only have you touched upon nearly every section of the project, but have included some excellent diagrams and tables which aid understanding of this pathway. In saying this, it is noted that two Wikipedia images have been used though it has been suggested that only one of the images utilised can be from Wikipedia. All information provided was also appropriately referenced and cited. In addition, I think it would be useful to utilise the discussion page to encourage interaction between group members as it allows individuals to provide feedback and suggestions. Hope this helps!
Group 6
It was good to see some progress being made on the project with the development of some subheadings and the inclusion of an image. In saying this, a better selection of headings and sub-headings could be developed to break down the topic of TGF beta signaling pathway. I think the sub-headings provided under the general heading of ‘Introduction’ could form the main headings of this research project and they could then be further broken down into various subheadings. In addition, the subheading of TGF-beta could be eliminated and this definition could be incorporated into the glossary or general introduction of the topic instead. Furthermore, more focus is needed on the influence of this pathway on embryological development and the abnormalities caused by mutations to the pathway and its components. For example, there has been mention of the effect of TGF-beta in ‘development of the embryo and adult organism, as well as cell growth, immune function and hormone secretion’ but further discussion has not been pursued.
Though a good description of the ‘process of TGF-beta signaling pathway’ has been provided, it could be further improved by referencing the images included in this section in your text (e.g. refer to Figure 1) to aid one’s understanding of the concept being explored. In addition, a timeline of events could be provided to explore the history of this pathway and it would be appropriate to begin with the discovery of TGF-beta. To a reader, information on ‘transformed or malignant cells’ seems unrelated to the TGF-beta signaling pathway even though it may be in fact be related, due to lack of discussion of this pathway or TGF-beta in this description. In regards to the section on ‘Limitations’, what types of limitations are you trying to explore? Limitations in research? This could be better defined by appropriately allocating subheadings to each of the sections.
Though you are heading in the right direction, spending time to produce a basic layout of your project by creating appropriate headings and subheadings would be useful in breaking down the concepts needed to be explored in this pathway. This could be achieved by communicating with group members through the discussion page and providing feedback and suggestions. In addition, greater focus is required in referencing and citing your work to ensure researchers and authors are acknowledged for their work. Also, by exploring animal models of the TGF-beta pathway and the effect of this research in understanding this pathway in humans and its influence on embryological development, you could greatly increase the quality of your work. You could also try and make your project more interactive and engaging through the inclusion of tables, images and diagrams. I hope this helps! Good luck!
Lab 10 Assessment
Presented our discussion of the research article <pubmed>21727907</pubmed>
| Lab 10 - Stem Cell Presentations 2016 | |
|---|---|
| Group Mark | Assessor General Comments |
|
Group 1: 15/20 Group 2: 19/20 Group 3: 20/20 Group 4: 19/20 Group 5: 16/20 Group 6: 16/20 |
The students put great effort in their presentation and we heard a nice variety of studies in stem cell biology and regenerative medicine today. The interaction after the presentation was great.
As general feedback I would like to advise students to:
|
- ↑ NIH U.S. National Library of Medicine,. (2016). TBX22. Genetics Home Reference. Retrieved 12 September 2016, from https://ghr.nlm.nih.gov/gene/TBX22
- ↑ <pubmed>14729838</pubmed>
- ↑ <pubmed>17846996</pubmed>
- ↑ <pubmed>15470384</pubmed>
- ↑ U.S. National Library of Medicine,. (2016). DMD gene. Genetics Home Reference. Retrieved 19 September 2016, from https://ghr.nlm.nih.gov/gene/DMD
- ↑ Muscular Dystrophy Australia,. (2015). Muscular Dystrophy - "The Home of MDA". Mda.org.au. Retrieved 19 September 2016, from http://www.mda.org.au/disorders/dystrophies/dmd-bmd.asp