User:Z5015686
| Student Information (expand to read) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Individual Assessments | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Please leave this template on top of your student page as I will add your assessment items here. Beginning your online work - Working Online in this course
Click here to email Dr Mark Hill | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 1 Assessment - Researching a Topic | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
In the lab I showed you how to find the PubMed reference database and search it using a topic word. Lab 1 assessment will be for you to use this to find a research reference on "fertilization" and write a brief summary of the main finding of the paper.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 2 Assessment - Uploading an Image | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
OK you are now in a group
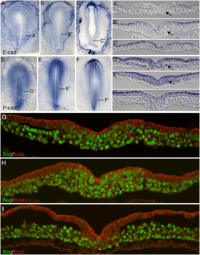
Initially the topic can be as specific or as broad as you want. Chicken embryo E-cad and P-cad gastrulation[1] References
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 4 Assessment - GIT Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
ANAT2341 Quiz Example | Category:Quiz | ANAT2341 Student 2015 Quiz Questions | Design 4 quiz questions based upon gastrointestinal tract. Add the quiz to your own page under Lab 4 assessment and provide a sub-sub-heading on the topic of the quiz. An example is shown below (open this page in view code or edit mode). Note that it is not just how you ask the question, but also how you explain the correct answer. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 5 Assessment - Course Review | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Complete the course review questionnaire and add the fact you have completed to your student page. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 6 Assessment - Cleft Lip and Palate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 7 Assessment - Muscular Dystrophy | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 8 Assessment - Quiz | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| A brief quiz was held in the practical class on urogenital development. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 9 Assessment - Peer Assessment | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 10 Assessment - Stem Cells | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
As part of the assessment for this course, you will give a 15 minutes journal club presentation in Lab 10. For this you will in your current student group discuss a recent (published after 2011) original research article (not a review!) on stem cell biology or technology.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lab 11 Assessment - Heart Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Read the following recent review article on heart repair and from the reference list identify a cited research article and write a brief summary of the paper's main findings. Then describe how the original research result was used in the review article.
<pubmed>26932668</pubmed>Development | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Lab Attendance
Z5015686 (talk) 14:34, 5 August 2016 (AEST)
Z5015686 (talk) 14:40, 12 August 2016 (AEST)
Z5015686 (talk) 13:15, 19 August 2016 (AEST)
Z5015686 (talk) 13:10, 26 August 2016 (AEST)
Z5015686 (talk) 13:07, 2 September 2016 (AEST)
Z5015686 (talk) 13:07, 9 September 2016 (AEST)
Z5015686 (talk) 13:33, 16 September 2016 (AEST)
Z5015686 (talk) 13:23, 7 October 2016 (AEDT)
Z5015686 (talk) 14:37, 14 October 2016 (AEDT)
Z5015686 (talk) 13:10, 21 October 2016 (AEDT)
Z5015686 (talk) 13:09, 28 October 2016 (AEDT)
Lab 1 Assessment
<pubmed> PMC4770082 </pubmed>
Short Summary Of Findings
The results of this paper suggest that human oocyte developmental potential can be predicted by the quality and maturation of the oocyte prior to fertilisation, (at the 2PN, pronucleus phase). The experiential design outlined in the paper involved measuring the mechanical properties of mouse and human zygotes using minimally invasive technologies (such as micropipette aspiration) to determine which were most predictive of viability (viability was defined as embryos that would most likely survive to blastocyst stage of development.) The results showed that individual parameters had limited predictive power on viability, however when considered together there was a greater distinction between viable and non-viable embryos. Through the use of statistical analysis it was found that their method of classification to predict embryo blastocyst formation that was based on these mechanical properties had >90% precision, 95% specificity and 75% sensitivity. Mice received embryos that were predicted to be either viable or non-viable based on their mechanical properties, which positively correlated to the mice who later had live births. The experimenters then investigated firstly whether there was a correlation between the viable and non-viable embryos and their gene expression, and secondly how/why these mechanical parameters correlated with viability. Interestingly they found that non-viable embryos had a reduced/different expression of some genes that are important for processes including, but not limited to, regulating cell cycle, oocyte maturation, chromosome segregation, DNA repair and telomere maintenance, thus suggesting that zygote gene expression correlates with viability. They also found that non-viable oocytes might undergo suboptimal fertilisation. Some genes that were identified to be differentially expressed in viable and non-viable embryos are important for fertilisation, including some whose products are found on the oocyte plasma membrane and in its zona pellucida, where if expressed incorrectly could potentially inhibit sperm-egg binding. Additionally, a reduced expression for a gene coding for a sperm protein was identified in non-viable zygotes, as well as a receptor that is involved in initiating the calcium oscillations that leads to cortical granule release and zona-hardening (which assists in the prevention of polyspermy.) Therefore in conclusion, this research demonstrates a way to accurately predict embryo viability early on in development, at the pronucleus stage, suggesting that embryo developmental potential is determined pre-fertilisation. This research has relevant applications in embryo selection process in IVF clinics.
| Mark Hill 18 August 2016 - You have added the citation correctly and written a good summary of the article's main findings. I guess the question is what provides the zygote viscoelastic properties and sperm gene expression? | Assessment 5/5 |
Lab 2 Assessment
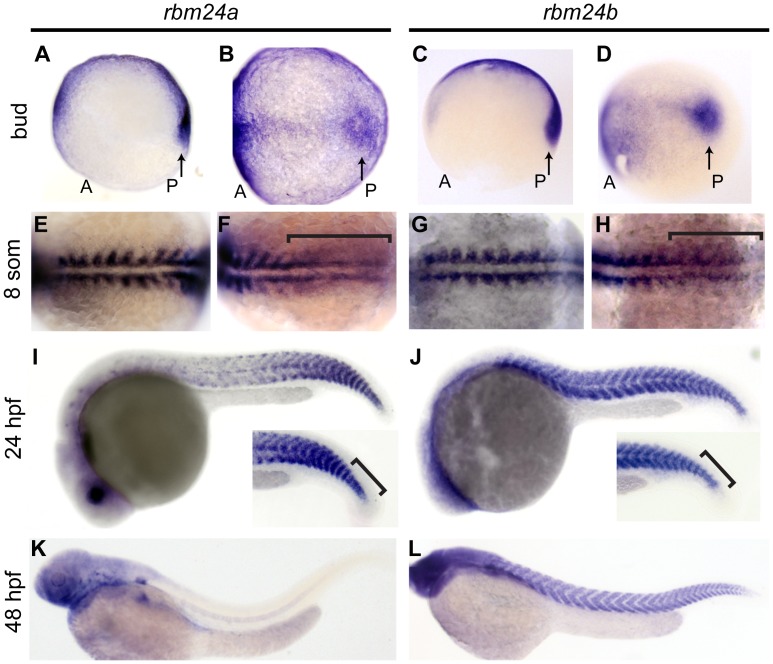
Rbm24a and rbm24b are expressed throughout somitogenesis[1]
| Mark Hill 29 August 2016 - All information Reference, Copyright and Student Image template correctly included with the file and referenced on your page here. | Assessment 5/5 |
Lab 3 Assessment
| Mark Hill 31 August 2016 - Lab 3 Assessment Quiz - Mesoderm and Ectoderm development. | Assessment 4/5 |
Lab 4 Assessment
Gastrointestinal Quiz
| Mark Hill 13 October 2016 - These seem good quiz questions, with some minor suggestions for improvement. Question 1 needs an explanation. Question should explain multiple answers are correct. Question 3 has a number of different topics mixed together, not good in MCQs. Question 4 is complicated for most correct type. | Assessment 5/5 |
Lab 5 Assessment
Completed Course Feedback Questionnaire
Z5015686 (talk) 13:15, 9 September 2016 (AEST)
| Mark Hill 11 October 2016 - Questionnaire on course structure. | Assessment 5/5 |
Lab 6 Assessment
Identify a known genetic mutation that is associated with cleft lip or palate: Mutations in the Interferon Regulatory Factor 6 (IRF6) protein-coding gene (located on chromosome 1) account for the majority of cases of Van der Woude syndrome (VDWS), an autosomal dominant genetic disorder, which has been found to be associated with both cleft lip and cleft palate.
Identify a recent research article on this gene: PMID 23029012
How does this mutation affect developmental signaling in normal development: For the most part the underlying mechanism behind the mutation of the IRF6 gene and the development of cleft lip and palate is largely unknown. However, animal studies involving Irf6 mutant mice have offered an explanation to why this gene could contribute to the development of these abnormalities. These mice presented with hyper-proliferative epidermis failing to undergo terminal differentiation, leading to epithelial adhesions that are able to occlude the oral cavity. IRF6 is also thought to be involved in keratinocyte proliferation and differentiation as well as the formation of the oral periderm. [2] Recent research suggests that IRF6 gene interacts with other genes, specifically the Transforming Growth Factor Alpha (TGFA) gene (involved in activating a signalling pathway responsible for cell proliferation, differentiation and development) and may account for up to 10% of cleft lip and cleft palate cases. Interestingly, IRFA knockout mice didn’t express Tgfa in tissues in the palate. [3] In summary it is thought that mutations in the IRF6 gene are thought to affect developmental signalling directly or through associations with other genes, however more research is required.
| Mark Hill 13 October 2016 - OMIM IRF6 is a good example. It would have been good to describe the full signaling pathway in the last part of the answer. | Assessment 5/5 |
Lab 7 Assessment
What is/are the dystrophin mutation(s)? The dystrophin gene is the largest known human gene and is located on locus Xp21. Mutations of this gene (such as selections, point mutations and duplications) affect the structure/function of the protein dystrophin, and is responsible for causing both Duchenne (DMD) and Becker (BMD) muscular dystrophies[4], which have a prevalence 4.78 and 1.53 per 100,000 males respectively.[5] Although they have similar signs and symptoms, they vary in their severity, onset age and rate at which the disease develops - with DND being in general, the more common and severe of the two, appearing earlier in childhood in the from of muscle weakness and rapidly develops, affected individuals have impaired development of normal motor functions. Both are associated with the heart condition cardiomyopathy (weakened cardiac muscles).[6]
What is the function of dystrophin?
Dystrophin is an important cytoskeletal protein, and is a crucial component of the larger dystrophin-glycoprotein complex (DGC) which functions to both stabilise and signal interactions between the cytoskeleton, membrane and extracellular matrix, essentially have a central role in mediating muscle stability. Dystrophin has four main functional domains (actin binding amino terminal, central rod, cysteine-rich domains and carboxyl terminus) which help mediate the complexes interactions with cellular components, for example mediates interactions with actin filaments through the actin binding domain, and interactions with microtubules through the rod domain.[7]
What other tissues/organs are affected by this disorder? This disorder is known to result in both cardiac failure and respiratory failure due to the weakening of muscles (as healthy muscle fibres are lost and replaced by fibrosis and fat, and thus have reduced function.)[8]
What therapies exist for DMD? Diagnosis of DMD can be confirmed through DNA tests, muscle biopsy (testing for presence or relative size of dystrophin) and even prenatal tests, and although there is no current cure for this disease, some treatments are available to help control age of onset in the hope to maximise affected individuals quality of life. Pharmacological treatments include corticosteriods (including prednisolone and deflazacort) which have shown some benefits in patients such as an improvement in strength, pulmonary function, timed motor function and delaying age at loss of ambulation and cardiomyopathy onset - however, these medications are not without their own set of side effects.[9] Current research is looking into the possibility gene therapies which aim to restore dystrophin expression such as the use of viral vectors (acting as vehicles for DMD gene)[10], and antisense oligonucleotide mediated exon skipping. [11]
What animal models are available for muscular dystrophy? Historically the most popular animal model for muscular dystrophy over the years has been the MDX mouse, the results of which have been shown to be promising and now a larger animal model of canine DMD (cDMD) is being used.[12]
| Mark Hill 13 October 2016 - Very good. | Assessment 5/5 |
Lab 8 Assessment
Absent from lab class due to illness
| Mark Hill 13 October 2016 - This was an in class quiz on urogenital development. Please see me and you can attempt this assessment. |
Lab 9 Assessment
‘’Critical assessment of group projects’’ Z5015686 (talk) 01:04, 7 October 2016 (AEDT)
These are excellent reviews of the project pages, with some specific examples to illustrate your assessments. They include a balanced critical assessment and feedback, given the existing status of these pages. Hopefully they have taken your advice. 9/10
Group 1 – Wnt Signalling Pathway
Positive aspects of the project include that fact that this group has included detailed information of the different WnT signaling pathways. It does seem however, that this information would perhaps be better conveyed to the audience if it were accompanied with images (either sourced from the internet or hand drawn) and/or videos/animations, as well as some information on the role of each signaling molecule/receptor subtype (perhaps in a table) just to provide a more thorough explanation of this pathway. Furthermore, this group has made a conscious decision to include a glossary, although they have not yet started this, it is going to be something the group can add to whilst finishing the project and will help the reader better understand the concepts they discuss. This group has included a large amount of references throughout their project, including a significant amount of recent primary articles, which shows the reader that their information is well researched and very current. However, the only criticism here is that they aren't appropriately formatted for the purpose of this assignment. I would suggest that in text citations would be more appropriate, so the reader can clearly identify where this specific information is from and then go directly to said source if need be.
Alternatively negative aspects of the project, which may need some revising before submitting the final version of this assignment, would be the formatting of the project as it appears relatively incomplete. Although there are some subheadings, which are helpful, it may be useful to add additional ones to these to make it a little clear for the reader. For example perhaps use a similar scaffold to the other group projects, which have included ones such as introduction, history, outline of the signaling pathway, its specific roles in embryonic development and then abnormalities specifically relating to embryonic development, as this would help break up the information better and make the projects more consistent for readers. Most of the work on this project seems to focus on explaining the signaling pathway so I assume its more the case of the group hasn’t got around to it yet, but I think more information on the role this signaling pathway specifically has in embryonic development is required, like the paragraph on early stages of skin formation, in order to tie in the assignment with what we have been learning in the labs and lectures. As mentioned I think the subheadings may need some revision, and the current ‘What can go wrong’ may be better described as ‘abnormalities’ that way you could also include a discussion of abnormalities to Wnt that specifically influence normal embryonic development, as well as still include the paragraphs on its influence on tumor cells which could perhaps be found using the ‘omim’ site searching by a receptor subtype or pathway. Also, although you have included more of a discussion of abnormalities that occur later in development, it is interesting for the reader and does go beyond our understanding from class, but the main focus probably should be on abnormalities in embryonic development.
In conclusion this project is definitely on its way to being really good, the information on the signaling pathways appears to be well research. The major criticisms were mostly focused on presentational aspects of the project like subheadings, references and the inclusion of images/tables. With some more research on its role in early embryonic development and abnormalities this will be very successful.
Group 2 – Notch Signaling Pathway
First impressions of Group 2’s page on the notch-signaling pathway are all positive. Subheadings are very well defined. They have chosen to include a brief yet informative introduction on the pathway, a simple table outlining the major scientific developments over the last 100 years, the molecular mechanisms of the pathway, its specific role in embryonic development (which they have further defined as cardiovascular and CNS), role in animal development, abnormalities relating to this pathway and a glossary. I think another positive aspect of this project, is that they have identified additional subheadings for which they are still to do research on; a particularly important one is current areas of research which not many groups have included. Furthermore, additional positive aspects of this project include the addition of images on the canonical notch signaling pathway and its role in cardiovascular development (which both appear also to be appropriately added to the website), which support the text nicely. It might also be useful to find a relevant video to include just to break up some of the text, and help make the page more interactive. It appears this group has widely researched their topic using both primary and review articles, which are all appropriately referenced using in-text citations. All of these aspects help to clearly convey the necessary information to the reader, and fulfill much of the required criteria of this project. In terms of their written information, Group 2 has included really detailed information on its role in embryonic cardiovascular development, as well as identifying some of the major research articles that have lead to these discoveries and a little bit about them (which then the reader if they are interested it can go read thanks to the inclusion of the in-text citations.) They do include a section of the roles of this pathway in animal development, which is really interesting and goes beyond the normal scope of this course.
Some negative aspects of the project include that, as part of the criteria being that the project has an “element of teaching at a peer level using the student's own innovative diagrams, tables or figures and/or using interesting examples or explanations” perhaps it would be useful to consider including a hand drawn image when researching the non-canonical pathway or transcriptional regulation of notch signaling, or even of some of the receptor/ligands involved in this signaling pathway. Furthermore, on a similar note it may be important to summarise the receptor subtypes involved in the different pathways, their role in embryonic development and abnormalities of the receptor subtype specifically relating to embryonic development in a table or dot point format. Additionally perhaps more information on its role in the CNS (or other systems during embryonic development) even if its not as detailed as cardiovascular, may help to inform the reader of all of its various roles.
In conclusion, it appears that this project is one of the strongest, it has very clear and informative subheadings separating well researched written material, supported by images sourced from the Internet. The main criticisms were just including your own innovative diagrams or explanations, videos to help make it more interactive and table or dot points summarizing the different receptor subtypes involved in each pathway. Following the completion of this, and the subheadings yet to be researched (and glossary) it appears that this project is going to be very successful in informing peers about the said pathway.
Group 4 – Hedgehog Pathway
Positive aspects of this project include that Group 4 appear to have well defined subheadings, which function well to help the reader navigate through the page. The information is appropriately referenced using in-text citations, appearing to be from both primary and review articles. There is a significant amount of research on the mechanisms of the pathway but less of a focus on the role of this pathway in embryonic development, which I think is really important in order to relate it back to what we are leaning in both the lectures and tutorials. I think the inclusion of current research is a very important aspect to include in this project, as it identifies the current direction in which this research is heading. This might be also interesting to link to its clinical significance and abnormalities in the signaling pathway.
However, some negative aspects of the page include the lack of an introduction as this essentially establishes your page. You need to include a brief outline of the signaling pathway, a summary of its role in development and the other aspects of it you are looking to discuss. Furthermore, the inclusion of an image outlining the signaling pathway without any information inducing or explaining it should be corrected. The project appears to be very informative but isn’t very interactive and lacks images. Perhaps sourcing images of results from some of the primary articles, which you have referenced or include videos outlining the signaling pathway, might be a useful addition. It might be a good idea to include a glossary at the bottom of the page to help readers to better understand some of these more difficult terms. Also under the subheading of history, like in some of the other projects, a table could be a useful addition, just summarizing all the scientific advances regarding this pathway since it was first discovered, this helps set up how far we have come and then may be helpful when talking about the direction in which we are heading under current research.
In conclusion, this looks like it’s on its way to being a successful project. In summary though, a greater emphasis on its role in embryonic development and conscious effort to make the page more interactive and engaging for the reader will go a long way.
Group 5 – T-Box
First impressions alone it is extremely clear that Group 5 has thoroughly researched this topic have tried hard to include many diagrams and tables to help separate their information up in order to more successfully convey the information across to the reader. Positive aspects of this project include the well-defined subheadings, making the navigation through the page very easy. The introduction is informative and introduces the following subheadings of the project well. The inclusion of what does T-Box mean is also interesting, setting you apart from the other projects. One of the best aspects of the project would have to be the summary table of the main T-box genes, which includes its main expression sites, its function and abnormalities relevant to the specific gene. You have made a note to include a timeline for the history of the T-Box gene, which I think would be successful in summarizing the scientific advances since its discovery, and also help to break up paragraphs of writing. The project appears to be referenced correctly using in-text citations, only query is whether the links to the PMID articles say in the bottom of cardiac and limb development are references or just articles in which you haven’t written on yet and will be referenced appropriately when you do later. The inclusion of a glossary is also a good idea just to help define and explain some of the more difficult terms mentioned.
As for negative aspects of the project, there wasn’t too many. Like for every project, in terms of making it more interactive it might be a good idea to include a YouTube video or animation of the signaling pathway or its role in a specific developmental process, as well as your own hand-drawn image just to fulfill the necessary criteria of this assignment. Furthermore, with some of the smaller images that don’t go the full width of the page, it might be a nice idea to align them to the right as a thumbnail next to their relevant text, so readers see them whilst reading about it. Also remember to make a reference the image you have chosen in your text to emphasise its importance to what you are actually talking about. Although the subheading “good places to look” might just be something for you guys while researching, I think that you could utilize this by including various links with more information on the relevant topics of which you have discussed. This would help to make you page more interactive as well.
This project appears to be extremely well done and is definitely one of the strongest. Most of the criticisms are regarding the formatting of the page and making it more interactive for the reader. All in all this is very well researched project!
Group 6 – TGF-beta
You guys have made a good start to the project identifying some important subheadings introducing the TGF-beta signaling pathway, outlining its history, current research and limitations (which may be more appropriately labeled as abnormalities.) However, I do think the structure of these should be revised, what I mean by this is that you should create more levels of headings (as currently all the headings are located under the larger heading of introduction.) Furthermore, it terms of the headings, I think you need to introduce the signaling pathway, then discuss the history of its discovery, then discuss the specific mechanisms behind the pathway, its role in embryonic development (which is a very important aspect in order to relate your project back to what we are learning in the lectures and tutorials), then animal models and abnormalities. You have chosen to include some images which appear to be useful for explaining the signaling pathway, however I think it is important to refer to them in your text, as well as appropriately referencing them with the copyright from the original source (as the larger one is missing this information.)
Some negative aspects of this project are the lack of appropriate references, there are no in-text citations and the identified sources that have been used appear to be websites. Remember that most of the information, if not all should be acquired from primary research articles (supplemented with the occasional review article.) Furthermore, similar to other projects, in order to make your page more engaging you could look into including tables (say for the history or summary of receptor subtypes), more images, YouTube links or animations, or an interactive quiz.
In conclusion it seems that there is still a lot of work to be completed on this page before it is to be submitted, however you have made a successful start. The main criticisms are regarding revisiting the subheadings and including the role of embryonic development as I think this is really critical to the project, as well as adding more information to the page in general. In saying that it appears you guys are heading in the right direction!
Lab 10 Assessment
| Lab 10 - Stem Cell Presentations 2016 | |
|---|---|
| Group Mark | Assessor General Comments |
|
Group 1: 15/20 Group 2: 19/20 Group 3: 20/20 Group 4: 19/20 Group 5: 16/20 Group 6: 16/20 |
The students put great effort in their presentation and we heard a nice variety of studies in stem cell biology and regenerative medicine today. The interaction after the presentation was great.
As general feedback I would like to advise students to:
|
Lab 12 Assessment
5/5
Identify a cited research article: PMID 25848746
Write a brief summary on the papers findings: The researches in this article acknowledge that the administration of neuregulin-1 (NRG1) has been previously proposed as a method to promote cardiac regeneration, and their experiment specifically looked at the role of NRG1 co-receptor ERBB2 in cardiac regeneration (through implementing both loss and gain of function experimental methods.) They first found that NRG1-induced cardiomyocyte proliferation diminished one week after birth as a result of a reduction in ERBB2, and through knockout studies showed that ERBB2 is required for cardiomyocyte proliferation at embryonic and neonatal stages. They also did a series of experiments activating ERBB2 in cardiomyocytes from mice of different ages. Their experiments as a whole, suggest that (1) ERBB2 is needed for cardiomyocyte proliferation and (2) ERBB2 is able to reactivate postnatal cardiomyocyte proliferation and regenerative potentials. They acknowledge that further research and a deeper understanding of this signalling pathway (as well as others) could lead to major advances in regenerative medicines.
Describe how the original research result was used in the review article (PMID 26932668): Broadly speaking, this review article summaries the current knowledge the regulation of cardiomyocyte proliferation during both heart development and regeneration. This research article contributed to this review article by highlighting the role that the regulation of the Nrg1/ErbB2 pathway has in controlling postnatal cardiac growth. It was a basis one of their illustrations (Figure 3) which shows (1) that the transition of cardiomyocytes from hyperplastic to hypertrophic growth (during neonatal periods) is correlated with reductions in NRG1 co-receptor ERBB2, and (2) that in cardiomyocytes constitutively active ERBB2 (caERBB2) expression can extend/reactivate cell division (continued hyperplasic growth) as well as hypertrophic growth, which results in cardiomegaly (which is the abnormal enlargement of the heart.)
References
- ↑ <pubmed>25170925</pubmed> PLOSONE
- ↑ <pubmed>21331089</pubmed> [1]
- ↑ <pubmed>23029012</pubmed> [2]
- ↑ <pubmed> 4767260</pubmed> [3]
- ↑ <pubmed>24780148</pubmed>[4]
- ↑ <pubmed> 4767260</pubmed> [5]
- ↑ <pubmed> 4767260</pubmed> [6]
- ↑ <pubmed>4767260</pubmed> [7]
- ↑ <pubmed>26833937</pubmed> [8]
- ↑ <pubmed>27215286</pubmed> [9]
- ↑ <pubmed>23829870</pubmed> [10]
- ↑ <pubmed> 25740330</pubmed> [11]
fertilization PMID 27486480