Talk:2016 Group Project 5
|
Signalling: 1 Wnt | 2 Notch | 3 FGF Receptor | 4 Hedgehog | 5 T-box | 6 TGF-Beta | |||
| Here are some starting places for the topic. Can be patterning, differentiation, etc. as long as a developmental signal process/pathway. |
|
Assessment
| Group Assessment Criteria |
|---|
 Science Student Projects Science Student Projects
|
| More Information on Assessment Criteria | Science Student Projects |
Comments
- This is a well structured project with a good balance between text and graphic components.
- Several images needed to be deleted from the project due to copyright issues.
- Included quizzes were useful testing of topic knowledge. The quiz answers should have included additional descriptive information.
- Collapsible Timeline was useful and referenced.
- Animal model section was very well covered giving several animal examples.
- Glossary gave good brief descriptions of many but not all terms.
- References related to main project concepts and included current research findings.
- Project could have used a closing section like future research directions, etc.
- Project content language appropriate for university level student.
Edits
Media
- Z5039628 - copyright deletion.
- Z3516832 - copyright deletion.
- Video - Adult Tbx3;PrxCre mutant mouse is healthy and mobile despite forelimb deformities - good inclusion, relevant to project topic.
- Some image citations have not been included correctly, as shown in the class tutorial.
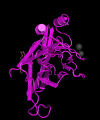
Tbx5 3D structure Z5020373 Student template and reference. The original journal J Mol Biol. has copyright, you should cite MMDB as shown by the citation link
- Ums.jpg
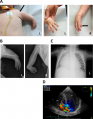
ums patient and d-f: mother of patient, normal Z5039628 This image cannot be reused online without permission and has been deleted from the project.
- Evolution of T box gene Family.jpg
Z3516832 Copyright and student template included. Image relevant to project, description from original figure legend. The only change to the original image is to change the colour of the boxes, this is not sufficient to qualify as "modified", therefore this image cannot be reused online without permission and has been deleted from the project. Citation - <pubmed>25294936</pubmed>
- T box in chick.jpg
Z3516832 Copyright (incorrect) and student template included. This image cannot be reused online without Journal permission and payment (32.20 USD) and has been deleted from the project.
Peer Review
Group 5 Peer Assessment
To be honest, this page is really good and already near completion in many aspects and there are very few negative points I could think of while going through the page. The first thing that appealed to many which was missing in a few of the other pages was the inclusion of an introduction. The page immediately stood out because of the inclusion of well formatted tables and images all related to the sub headings they were placed under. The research as well was extremely comprehensive and covered all topics required to gain a complete understanding of the T-box genes and their signalling. I particularly found the history aspect of it to be quite interesting. I also noticed that a wide variety of resources were used and the referencing for them was done with proper and consistent formatting making it easy to navigate through various articles and sources.
Some minor edits that can be done are that a few of a the PMID links remain under the headings which should be removed and included at the end with appropriate in text citation. Some of the abbreviations were not given in their full form when first used on the page.
This page is really great already and all the criteria have been carefully integrated and followed. Just a few changes here and there and you guys should have a top notch project!
Group 5 Peer Review
I have to say this page is the best across all groups. Very impressive work. Lots of references have been utilised and they are very supportive. The pictures cited are beautiful and are closely related to the contents. The molecular basis has been well explained and specific components of the pathway were pointed out in each signalling events related in embryo development. Moreover, you guys have also shed light on the ancient origin (evolutionary aspect) of this pathway which is really interesting.
Only a few points to mention. More pictures will make your page look better. Maybe one picture per section? Secondly, the origin and the ancient origin are a little bit confusing, maybe change their names and combine all the background things like those together? A few sections need to be completed and there are some structural problems need to be adjusted.
This is a really impressive web page. Good job Group 5!
Group 5 Peer Review
Congrats Group 5 on producing a really great looking page so far! Straightaway I like that you have an introduction paragraph and that I can immediately see inclusion of tables, images, and some correct referencing. Explaining the origin of the T-box name is a great piece of background info to include (maybe make it a proper subheading though?). You’ve also touched on some of the history of the gene/signalling pathway in your paragraphs but it might be good to also present that in a brief timeline/table. Your table for T-box family features is really great - especially because of inclusion of main expression sites and the related abnormalities. Also, you have references to primary research articles so it’s good to see you’re backing up your content with accurate and relevant sources. You’ve done a really great job on exploring the animal models as well.
Ultimately you guys have done great work so my suggestions for improving your page are mostly minor! You have a lot of PMID links just left at the end of some paragraphs so make sure you get those properly listed at the end in your reference section. If you change your pictures to thumbnails then I think they would integrate into your text better (because at the moment where they are placed breaks up the text). Also, I’m not sure if there’s a particular reason why you did this in the first place but I wouldn’t capitalise all the subheadings in the abnormalities section. And I suggest moving the ‘Ancient origins and evolution of the T-box gene family’ near the top of your page, because at the moment it seems out of place and doesn’t really flow on from discussing the abnormalities. The other thing I would say is that - if it’s possible - it would be great if you could explain more about the actual molecular pathway and include a picture of the molecules/factors involved, because at the moment you’ve only described it in the context of different TBX genes being expressed in each developmental role. I think especially if you have an image of the structure of some of the different proteins, transcription factors, etc. then it would really help to visualise the molecular aspects of the pathway. Hope my comments are helpful!
Group 5 Review
Your team has a very impressive wiki page, well done! The key points relating to T-Box as well as your choice of subheadings and headings are very good, however I would advise removing 'Good places to look'. In terms of diagrams, tables and graphs, these are present and augment the information presented quite well. The content presented is cited mostly correctly however care must be taken with pictures, which have to be checked for copyright reuse as well as ensuring that they are cited correctly in the first place, I would advise that your team checks each of your pictures to make sure that they are correctly cited.
In the context of peer level education, your content is understandable and written well even though the topic is complex. What is lacking however are using your own explanations as well as interesting hand drawn visual stimuli to present information, this can be easily remedied. Also, completion of the glossary section so that someone can understand complex terms would be useful. With the information that has been provided and the depth of research that has went into the meticulous presentation of information regarding T-box, it is clear that your team has went beyond formal teaching activities, however, perhaps the inclusion of some interactive features on your page such as a video with voice over or a quiz would help augment this criterion. The learning aims of the Embryology course are mostly in line with the information on the wiki page, but there is no section for current research/technologies, which is important to address the second criteria of the course aims.
Overall, you guys did a very nice job that requires only minor touch ups and the addition of a few pieces of information. Don't forget the current research section though, that is pretty important to include in my opinion. Well done!
Group 5 Peer Review
Positive Factors
Group 5 have introduced their topic really well, I think it could be improved by putting the second half of their intro under the ‘History’ subheading though and maybe it could be moved up so it is straight after the introduction. The table they have included shows they have considered addressing criteria 4 as I think it makes it easy for students to quickly take in a lot of information. Furthermore, they have addressed criteria 1 and 2 by organising the subheadings and sub-subheadings in a way that gives the page a logical flow. The amount of references already incorporated in their project shows that they have already completed extensive research on the topic area, which addresses criteria 5. Criteria 6 has clearly been addressed in the ‘development’ subsections.
Points for Improvement
Some improvements that Group 5 could make to their already extensive effort include: changing some of the headings in the table to bold so that they are clearer/easier to read; they could also uncapitalise the subheadings under ‘Abnormalities’ to make the page more uniform; and also formatting the images to incorporate them around the text (rather than breaking up the page each time) would improve the flow of information.
Overall
Overall Group 5 have already done extensive research as evidenced by the volume of information and various images included in their page, I can see that they have made an effort to address most of the criteria already. The improvements they need to make mainly involve formatting to make the page more student-friendly.
Group 5:
Positive aspects of the project and suggested improvements:
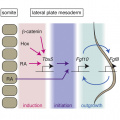
Upon reviewing this page, it is clear that group 5 has provided numerous headings and subheadings related to Tbx-genes ranging from origins of the genes, their function in embryonic development, abnormalities, history and animal models (criteria 1 and 6). In doing so, the group has also ventured to provide an in-depth explanation of each subheading. Take for example the subheading named, “limb development”, the authors have provided an in-depth description into the role of T-box transcription factors in limb development whilst utilising a diagram to reinforce this description (criteria 2). It also appears that in-text citations have been correctly used to reference the sources of data in most cases (criteria 3). The authors have utilised diagrams and a table to describe various components of the T-box gene ranging from the different types of T-box genes to its mechanisms in embryonic development (criteria 4). The extensive use of diagrams allows the audience to develop a holistic understanding of the various subheadings included, as these diagrams convey the description provided in a visual manner (criteria 5). It is also evident that the group has conducted research into animal models and evolution of the T-box gene, thus demonstrating that the group has investigated areas of research beyond formal teaching activities (criteria 5).
Improvements which may be made to this page would be to include a timeline regarding the history of the T-box family, as this will display the information in a much more organised and appealing manner. Another improvement which may be made would be to include a YouTube video to introduce the signalling process in development, such as in cardiac and limb development for example. In order to make the wikipage interactive, a further improvement which may be made would be to include a set of multiple choice questions at the end of the page which ask questions about the content covered.
Negative aspects of the project and suggested improvements:
Alongside the various positive aspects of this project, there are few negative aspects. A negative aspect identified includes the use of images from Wikipedia pages more than once. It was stated that only one Wikipedia page was allowed to be included as a source. Therefore a suggestion would be to obtain images and data from research articles rather than from Wikipedia pages, as research articles are often a more reliable source of data. It was also noticed that images were not utilised to describe different abnormalities associated with the TBX gene, hence a possible improvement would be to include images depicting such abnormalities. These images may make this section of the page more appealing and engaging to audiences.
It was also noticed that the image titled “Evolution of the T box gene family”, was incorrectly referenced. Therefore, it is suggested that the authors of the project ensure that the original author of the image are correctly referenced to ensure that copyright laws are not breached. The final negative aspect of the project was that the “Ancient origins and evolution of the T-box gene family” subheading appeared out of place in the page. Therefore a possible improvement would be to include evolution of the T-box gene under the “Origins of the T-box gene” subheading at the beginning of the page as this will create a sense of consistency in the page.
Group 5
First of all, well done on making significant progress on your project! You have addressed all aspects of the pathway involving T-Box genes through subdivision into various headings and subheadings. I particularly liked how there was an inclusion of the specific T-Box gene affected in each of the abnormalities in the subheading itself. The only suggestion I would make is to combine the ‘Ancient origins and evolution of the T-Box gene family’ section with the origins of the ‘T-Box genes’ section to provide a more coherent description of the history of these pathway. You could even form a table to create a timeline of events. In addition, I think it would be beneficial to include the ‘What does T-Box mean?’ as an introduction to the ‘origins of the T-box genes’ section as there is overlap between these sections.
The use of a table to describe the main T-box genes was helpful in providing a brief overview of the components of the pathway and their influence in embryological development. In addition, the link between T-Box genes and embryonic development has been explored considerably. In saying this, greater attention to detail must be paid to explaining abbreviations to aid one’s understanding of the concepts being discussed. For example, what is NKX2-5, Shh and OFT? Though you’ve explained that RA stands for retinoic acid in the ‘Organisms used in animal models for T-Box’ section, this same explanation is not provided in the ‘Limb development’ section where you have discussed that ‘RA and Shh both induced Tbx2’. These small changes will significantly improve the quality of your work.
The inclusion of abnormalities provides great insight into the role of T-Box genes in development. In saying this, though you have explored the effect of the mutation of these genes in animal models, more information is required to explain the effect of these mutations in humans and how they come about. Furthermore, under the heading of ‘Animal models’ there has been discussion mainly of the ‘brachyury gene’ which seems unrelated to animal models due to the lack of a proper introduction. I found the following section (organisms used in animal models for T-Box) to be a better introduction to the topic of animal models. In addition, there has been mention of a number of animal models ‘Drosophila, Xenopus, zebrafish, avians, and mice’ yet only marsupials and amphioxus has been discussed. This could be potentially misleading to readers.
Overall, a fantastic effort has been made. Not only have you touched upon nearly every section of the project, but have included some excellent diagrams and tables which aid understanding of this pathway. In saying this, it is noted that two Wikipedia images have been used though it has been suggested that only one of the images utilised can be from Wikipedia. All information provided was also appropriately referenced and cited. In addition, I think it would be useful to utilise the discussion page to encourage interaction between group members as it allows individuals to provide feedback and suggestions. Hope this helps!
Group 5 Peer Review
This group’s project has been very well done they have provided a substantial amount of information as well as media to supplement all written concepts they have explained. They have provided self-edited/modified images however they have not produced any of their own illustrations/flowcharts, this should be added to fulfil marking guidelines and demonstrate their own understanding of subject matter.
Unlike many other group they have provided well thought out tables which provide a brief summary of a collection of the main T-box genes. In this table they have outlined abnormalities however they want to consider adding visual aids for visible conditions such as clef palate, this would allow readers to easily understand what mutation in a certain T-box gene would manifest as phenotypically. In addition to this the image referencing is inconsistent throughout the page, in some areas there will be hyperlinked text “image from here” where as in other locations there will be a superscripted reference. There is nothing wrong with this however by accustoming the viewers to a consistent form of referencing will improve readability and allow for an easier flow.
Overall there is little to critique about this project and the group has put in a lot of work to ensure that majority of the boxes are ticked, they may however want to look into ways of making their page more engaging by adding small quizzes or true or false questions scattered along the span of the article to ensure readers are understanding the conveyed information properly.
Group 5 Critical Assessment
Well done on constructing a thorough Wiki page on the topic of T-box Genes and their Signalling! Viewing the page it is evident numerous headings and subheadings have been provided to accommodate for the large amount of information gathered. Starting off with the introduction I like how you have included a section on what T’-box exactly means, however the information provided in this section talks about the history significantly, hence to turn this into a positive I would suggest adding a table or timeline outlining the major events and discoveries in the past to present this information in a complete, meaningful way. This issue is also seen with the section ‘Origins of the T-box genes’ where major discoveries are highlighted and in which year they occurred. This information can also merge with the history timeline/table.
Within the ‘What does T-box mean?’ section you have also added information on which animal studies were undertaken for the discoveries. To avoid spreading of information and causing confusion for the reader, you could either construct a table to show which animal study was completed in which year, and what discovery it led to as 3 columns, or bring this information down to the section ‘Animal Models’. In saying that, you have attempted to utilize a table and the table works very well with the topic of the different T-box genes, and would prove great help for the viewer.
It is great to see T-box genes and Signalling has been explored further in the field of embryonic development. Extensive information is provided with good use of in- text citations, allowing the reader to navigate to relevant articles. The ability to navigate could be further improved by providing an accessible link to the ‘Abnormalities’ section in a case where you are directing the reader to the section for further information, instead of plain text. Beneath each section for e.g. ‘Limb Development’ Pubmed links have been provided to relevant articles, which is a fantastic idea, however the links have no indication whatsoever of what the article is about. You could add a sentence each next to the links briefly stating what the article is exploring in relation to limb development.
Overall with a few more images, possibly some interactive components such as clips, and a knowledge testing short exam or quiz this Wiki page will stand out. Remember to ensure your information flows well by placing it within appropriate sections!
Group 5 Peer Assessment
It is quite clear that what has been provided in your wiki page is extensive and well researched. The inclusion of tables summarizing the different T-box genes although extensive, is very concise and easy to read. I feel that this table really links all the elements of your page together, where you have included its function and related it to embryological development and abnormalities which you go on later to elaborate in other sections. I feel this really complements the introduction and gives a good feel for what’s to come in the rest of the page. The addition of what the term T-box means also is a nice touch, giving context and some history regarding the name.
Your origins section of T-box is quite well outlined, but as mentioned in your page, having a timeline with critical points of discovery with regards to the genes would probably be more beneficial as it would be a lot easier to read a see the time points as a whole. That being said, having the timeline alongside your outline would probably work well, as your outline can serve to elaborate on the timeline. With regards to your subheadings, it seems to they are quite extensive and cover practically all the key components of the T-box genes, and it is also good to see that there is a glossary subheading in place. Content wise there seems to be limited to no issues, but with regards to abbreviations, I have found that the usage hasn’t always been after the fact of providing the full name first. For example, bone morphogenic protein’s abbreviation is used consistently throughout the first part of the wiki page, but it is only described by its full name and then abbreviation later on. This is something you should check out and fix by either adding the full name the first time the abbreviation is used, or adding all these terms to the glossary.
With regards to the pictures they all seem to compliment the sections well and are quite plentiful. That being said though the picture in the “Marsupial forelimb development” does not appear to have the copyright information regarding to its usage, and referencing does not appear to be in full. This is also the same for the picture under the subheading “Organisms used in animal models for T-box”. Other than that the referencing is perfectly fine within the text.
Overall this project is really good and without any major flaws when it comes to the content. A few touch ups here and there with regards to my suggestion above, and your project should be good to go along as the quality is kept at this level.
Group 5 – T-Box
First impressions alone it is extremely clear that Group 5 has thoroughly researched this topic have tried hard to include many diagrams and tables to help separate their information up in order to more successfully convey the information across to the reader. Positive aspects of this project include the well-defined subheadings, making the navigation through the page very easy. The introduction is informative and introduces the following subheadings of the project well. The inclusion of what does T-Box mean is also interesting, setting you apart from the other projects. One of the best aspects of the project would have to be the summary table of the main T-box genes, which includes its main expression sites, its function and abnormalities relevant to the specific gene. You have made a note to include a timeline for the history of the T-Box gene, which I think would be successful in summarizing the scientific advances since its discovery, and also help to break up paragraphs of writing. The project appears to be referenced correctly using in-text citations, only query is whether the links to the PMID articles say in the bottom of cardiac and limb development are references or just articles in which you haven’t written on yet and will be referenced appropriately when you do later. The inclusion of a glossary is also a good idea just to help define and explain some of the more difficult terms mentioned.
As for negative aspects of the project, there wasn’t too many. Like for every project, in terms of making it more interactive it might be a good idea to include a YouTube video or animation of the signaling pathway or its role in a specific developmental process, as well as your own hand-drawn image just to fulfill the necessary criteria of this assignment. Furthermore, with some of the smaller images that don’t go the full width of the page, it might be a nice idea to align them to the right as a thumbnail next to their relevant text, so readers see them whilst reading about it. Also remember to make a reference the image you have chosen in your text to emphasise its importance to what you are actually talking about. Although the subheading “good places to look” might just be something for you guys while researching, I think that you could utilize this by including various links with more information on the relevant topics of which you have discussed. This would help to make you page more interactive as well.
This project appears to be extremely well done and is definitely one of the strongest. Most of the criticisms are regarding the formatting of the page and making it more interactive for the reader. All in all this is very well researched project!
Group 5 Peer Assessment
Positive aspects of the project and improvements:
Upon reviewing the page, it is evident that there has been a lot of research put in this project. Initially, there is evidence of a range of headings and subheadings which allowed the navigation from one aspect of the project to another extremely easy. This allowed me to confirm that the project is about T-box genes and their signalling. Secondly, it was excellent to see a range of images, tables and graphs as they provided visual aids to learn more about the topic and in general made it easier to accumulate information. Also it was good to see that these tables and images were correctly cited and referenced at the end which meant that there was no breach of copyright laws.
Also, throughout the project there was sufficient amount of information in each subheading which meant that the reader gained all relevant information pertaining to the section that they are reading. It was also great to see a range of abnormalities being added to the project. This meant that you have went above and beyond the scope of the assessment and researched that extra bit to provide additional information about the signalling pathway and complications arising from any mutations. This meant that you successfully satisfied criteria 5 and thus a more rounded project.
Negative aspects of the project and improvements:
This project certainly contains a range of positives but there were minimal negatives that can easily be amended in order to achieve a very high mark. I noticed that there was more than 1 image being used from Wikipedia and the criterion clearly says that a maximum of 1 was allowed. This is not a big deal but just in case there is harsh marking and penalties, it is advised to replace the additional image with another image. In addition, it would be useful to add a glossary of all the terms that one may find confusing such as “homologues”, “heterozygous”, “homology”, “notochord” etc. This in turn will provide the reader with enough information to understand the context of the project and in turn keep them engaged.
Another negative aspect of the project was that the subheading “Ancient origins and evolution of the T-box gene family” randomly appearing nearing the end of the project. This looked a bit out of place and not flowing with the rest of the passage. To correct this it would be advised to add this to the start of the page with the “Origin of the T-box genes” section just so the information clearly flows from one topic to another without creating confusion. Overall, this project is coming along quite nicely. It is evident that a lot of research has been put into constructing a coherent and succinct project but also have the visual cues to back up the main aspects. To maximise marks, it is recommended to reflect on the feedback and correct the minor mistakes.
Group 5 Peer Assessment
Positive feedback:
This is a very well, put together and organised page. Everything is very simple and straight-forward make it extremely student friendly and something I would definitely use to learn about T-Box genes.
The introduction along with explanation of the actual meaning of T-box is both informative and also interesting and gives students a good chance to take a break from the heavy load of information and actually indulge in some interesting facts.Following this, the table is one of the most useful things on the entire page and is extremely concise and structurally pleasing. It provides the key and relevant information and allowed me to make connections with T-box genes and their functions straight away.
There is also the use of many pictures throughout the page which definitely aids in visual learning and the more the used the better. The chronological structuring of the page is also very impressive. As I was reading the page I felt like the information that I was gathering was carrying on and helping me understand what was talked about in the next sections.
Critical Assessment:
The chronological structuring although very impressive did fall a little out of place when the sub-headings “Ancient origins and evolution of the T-box gene family” appeared at the end of the page when it seems like this is something that should be included in the start. It would be thoroughly recommended to utilised hand drawings to explain some of the concepts, especially when introducing the signalling because this would really compliment your already easy to understand introductions and really enhance learning/understanding.
Lastly I think it would be a good addition to your project to include some information on the current research that is being conducted on t-box genes and also if there are treatments for the abnormalities. This page is looking amazing so far so keep up the good work!
GROUP 5
From reading through Group 5’s project I can see that they having clearly and effectively talked about topics relating to Tbx genes. Their subheadings include animal models, history, function in embryonic development, as well as abnormalities. This provides evidence to me that they have obviously considered and addressed both criteria’s 6 and 1. Group 5 has done a fantastic job with criteria 2, in that they provide many visual representations of information with different forms. For examples they provide a table of the main T-box genes, but then also have many visual diagrams relating to the Tbx development. Even though criteria 2 is very well done, it could be improved by having a timeline of the history of the T-box family. Also, I have seen in some other groups that they have included multiple choice questions to test the audience on their knowledge, I think this is a good idea since it makes the learning experience more interactive.
Group 5 Peer Review
At first glance I can see that you’ve done an amazing job and a lot of time and effort has been put into your page. There are a lot of subheading that clearly divide the page into groups and makes it easier to navigate for the reader. Your introduction is excellent, it is very detailed and explains a lot about T-box genes and signalling. The use of the table to clarify the different T-box genes is very useful, it clearly explains the differences between them in a very simple way. You have also covered quite a lot of information on the function of T-box in development. This is quite an impressive section and also very detailed. Another great addition to your page is the abnormalities section, this is also very detailed and extremely interesting to read. You have also clearly discussed research related to your signalling pathway and several animal models, great addition to the page. The referencing throughout is also a great bonus to the reader and you have used an extensive number of sources.
One fault I can see on your page is the referencing added at the end of the paragraphs, maybe move them to the bottom of the page since you already have in-text referencing. You have included a section on the origins and evolution of the T-box gene, you should probably move it to the beginning of the page to give the reader a basic and initial understanding of what is known. Consider adding a glossary section to the end of your page to better explain some key terms you have used.
Overall, this is a wonderful page and you have paid a great attention to detail and covered a lot of content. All your information was relevant and very interesting to read. There is not much to fault on your page, only some minor changes to make the page more appealing and more presentable.
Group 5 Peer Review
Group 5, your project is the best I have seen out of all the groups! The amount of time and effort you have put in is notably evident in how much information your page holds. There are so many elements to the page, such as the Features, Origins, Functions of the T-box genes and many more. I am very impressed with what you guys have created.
As I was scrolling down your page, I love how you have used a variety of ways to present your information. For instance, you used a table to display the T-box genes. In this table, you have included the main expression sites during embryogenesis, the function and the abnormalities. The organised structure of the table provides consistency throughout the T-box genes and makes it simple to read and understand. Great work! You have also added multiple pictures throughout the page that supports the content beside it. It is correctly referenced and all is of good sizing and location. The amount of information you have included in your page is balanced out well with the table and the pictures.
There is very little to improve with your project. Considering how much information you have put in your project, maybe you can add a quiz to test the reader’s understanding? Also, I noticed that the glossary is still empty. Words and definitions should be added, but this is only a minor area of improvement. In regards to the references, I was very impressed with the amount of articles you have used for this project, shown through the correct use of in-text citation and collectively at the reference list at the bottom of the page.
Overall Group 5, you have done an awesome job so far. There are only a few minor improvements that can be done to this project. Great work!
Group 5 Peer Assessment
Extremely good page that looks almost up to completion. The information is well-detailed with appropriate headings, subheadings and referencing. The visual aids used are great in helping understand the topic and they too are also cited correctly. The specific pathway is described along with its history. The embryonic development information is cited with relevant articles and went well beyond the scope of the assessment with additional information about the mutations. Some points to improve (although minor) would be the use of a glossary and perhaps re-organising the structure of some subheadings such as the “Ancient origins..” section which seems like it should belong to the start. Overall, the page is really good and covers a lot of content whilst keeping it relevant and intriguing to the reader. Great work!
Some searches to get us started: T-box tbx
Z5020373 (talk) 14:44, 26 August 2016 (AEST) PMID 25294936 - a relatively recent article that provides background info on the T-box gene family PMID 16285859
Z3516832 (talk) 14:52, 26 August 2016 (AEST) http://www.columbia.edu/itc/hs/medical/humandev/2007/HD15/HD15.pdf
Z5020373 (talk) 11:32, 16 September 2016 (AEST) Does anyone know how to draw up a table on the page? Thanks.