Y Chromosome: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| (18 intermediate revisions by the same user not shown) | |||
| Line 2: | Line 2: | ||
==Introduction== | ==Introduction== | ||
[[File:Human_Y_chromosome_SRY_region.jpg|thumb|alt=Human Y chromosome showing SRY region|Human Y chromosome showing SRY region]] | [[File:Human_Y_chromosome_SRY_region.jpg|thumb|alt=Human Y chromosome showing SRY region|Human Y chromosome showing SRY region]] | ||
Humans have 23 pairs of chromosomes, 22 autosomes and a pair of sex chromosomes. Males have one | Humans have 23 pairs of chromosomes, 22 autosomes and a pair of sex chromosomes. Males have one {{ChrX}} and one {{ChrY}} chromosomes (46, XY) and females have a pair of [[X_Chromosome|X chromosomes]] (46, XX). The Y chromosome is much smaller than the X chromosome and contains 50 million base pairs encoding approximately 200+ genes. The Sry gene (location Yp11.3), found in 1990, encodes is responsible for male sex determination. SRY mutations lead to XY sex reversal in humans, and XX mice with an SRY transgene develop as fertile males. Interestingly, the laboratory rat, ''Rattus norvegicus'', has at least 6 full length copies of the Sry gene.{{#pmid:17408480|PMID17408480}} | ||
{{male}} | |||
| Line 13: | Line 13: | ||
The male-specific region of the human Y chromosome was originally called the non-recombining portion of the Y chromosome (NRY). This region consists of three different classes of euchromatic sequences: | The male-specific region of the human Y chromosome was originally called the non-recombining portion of the Y chromosome (NRY). This region consists of three different classes of euchromatic sequences: | ||
# X-transposed | |||
# X-degenerate | # X-transposed - 3-4 Ma transposition 99% identical to human {{ChrX}}q21 | ||
# ampliconic sequences | # X-degenerate - 14 single copy or 13 pseudogene homologs of X-linked genes | ||
# ampliconic sequences - Sequences with strong identity to other regions of Y | |||
The Y and X chromosome are both transcriptionally silenced during spermatogenesis, at primary spermatocyte stage onward, by a process called meiotic sex chromosome inactivation (MSCI). (More? [[Cell Division - Meiosis|Meiosis]]) | The Y and X chromosome are both transcriptionally silenced during spermatogenesis, at primary spermatocyte stage onward, by a process called meiotic sex chromosome inactivation (MSCI). (More? [[Cell Division - Meiosis|Meiosis]]) | ||
{{ | {{Y Chromosome Links}} | ||
[[File:Human_idiogram.jpg|400px]] | |||
Human idiogram | |||
== Some Recent Findings == | == Some Recent Findings == | ||
[[File: | [[File:Male XY karyotype.jpg|thumb|X and Y chromosomes]] | ||
{| | {| | ||
|-bgcolor="F5FAFF" | |-bgcolor="F5FAFF" | ||
| | | | ||
* ''' | * '''XY oocytes of sex-reversed females with a Sry mutation deviate from the normal developmental process beyond the mitotic stage'''{{#pmid:30289439|PMID30289439}} "The fertility of sex-reversed XY female mice is severely impaired by a massive loss of oocytes and failure of meiotic progression. This phenomenon remains an outstanding mystery. We sought to determine the molecular etiology of XY oocyte dysfunction by generating sex-reversed females that bear genetic ablation of Sry, a vital sex determination gene, on an inbred C57BL/6 background. These mutant mice, termed XYsry- mutants, showed severe attrition of germ cells during fetal development, resulting in the depletion of ovarian germ cells prior to sexual maturation. Comprehensive transcriptome analyses of primordial germ cells (PGCs) and postnatal oocytes demonstrated that XYsry- females had deviated significantly from normal developmental processes during the stages of mitotic proliferation. The impaired proliferation of XYsry- PGCs was associated with aberrant β-catenin signaling and the excessive expression of transposable elements. Upon entry to the meiotic stage, XYsry- oocytes demonstrated extensive defects, including the impairment of crossover formation, the failure of primordial follicle maintenance, and no capacity for embryo development. Together, these results suggest potential molecular causes for germ cell disruption in sex-reversed female mice, thereby providing insights into disorders of sex differentiation in humans, such as "Swyer syndrome," in which patients with an XY karyotype present as typical females and are infertile." | ||
* '''Alterations of sex determination pathway in the genital ridges of males with limited Y chromosome genes'''{{#pmid:30285093|PMID30285093}} "We previously demonstrated that in the {{mouse}} only two Y chromosome genes are required for a male to produce an offspring with the help of assisted reproduction technologies (ART): testis determinant Sry and spermatogonial proliferation factor Eif2s3y. Subsequently, we have shown that the function of these genes can be replaced by transgenic overexpression of their homologues, autosomally encoded Sox9 and X-chromosome encoded Eif2s3x. Males with Y chromosome contribution limited to two (XEif2s3yOSry), one (XEif2s3yOSox9 and XOSry, Eif2s3x) and no genes (XOSox9, Eif2s3x) produced haploid germ cells and sired offspring after ART. However, despite successful assisted reproductive outcome, they had smaller testes and displayed abnormal development of the seminiferous epithelium and testicular interstitium. Here we explored whether these testicular defects originated from altered pro-testis and pro-ovary factor signaling in genital ridges at the time of sex determination." | |||
|} | |} | ||
{| class="wikitable mw-collapsible mw-collapsed" | {| class="wikitable mw-collapsible mw-collapsed" | ||
! More recent papers | ! More recent papers | ||
|- | |- | ||
| [[File:Mark_Hill.jpg|90px|left]] {{Most_Recent_Refs}} | | [[File:Mark_Hill.jpg|90px|left]] {{Most_Recent_Refs}} | ||
Search term: [http://www.ncbi.nlm.nih.gov/pubmed/?term=Y+Chromosome ''Y Chromosome''] | Search term: [http://www.ncbi.nlm.nih.gov/pubmed/?term=Y+Chromosome ''Y Chromosome''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=SRY ''SRY''] | [http://www.ncbi.nlm.nih.gov/pubmed/?term=DAZ1 ''DAZ1''] | ||
|} | |||
{| class="wikitable mw-collapsible mw-collapsed" | |||
! Older papers | |||
|- | |||
| {{Older papers}} | |||
* '''Dual nuclear import mechanisms of sex determining factor SRY: intracellular Ca2+ as a switch''' {{#pmid:21051653|PMID21051653}} "The sex-determining region on the Y chromosome (SRY) has 2 nuclear localization signals (NLSs) that flank the DNA binding high mobility group (HMG) domain; the β-NLS and the CaM-NLS, which mediate nuclear transport through importin β1 (Impβ1) and the calcium-binding protein calmodulin (CaM), respectively. ...The results imply mutual exclusivity of nuclear transport via the 2 NLSs with intracellular Ca(2+) as the switch between the 2.-Kaur, G. Jans, D. A. Dual nuclear import mechanisms of sex determining factor SRY: intracellular Ca(2+) as a switch." | |||
|} | |} | ||
==History== | ==History== | ||
* '''1916''' - Bridges | * '''1916''' - Bridges{{#pmid:17245850|PMID17245850}}{{#pmid:17245853|PMID17245853}} describes the sex chromosomes of ''Drosophila melanogaster'' and shows that sex is determined by the X:autosome ratio | ||
* 1'''923''' - Painter | * 1'''923''' - Painter{{#pmid:17748214|PMID17748214}} describes the Sex chromosomes in mammals - it is assumed that sex determining mechanism will be the same as ''Drosophila'' | ||
* '''1959''' - Ford et al. | * '''1959''' - Ford et al.{{#pmid:13642858|PMID13642858}} find a human XO and female - Turner's syndrome | ||
* '''1959''' - Jacobs and Strong | * '''1959''' - Jacobs and Strong{{#pmid:13632697|PMID13632697}} find an XXY male - Kleinfelter's syndrome | ||
** together these prove that the Y chromosome determines sex and not the X:autosome ratio | ** together these prove that the Y chromosome determines sex and not the X:autosome ratio | ||
* '''1989''' XX males found that carry a very small portion of Y chromosomal DNA (60kb) | * '''1989''' XX males found that carry a very small portion of Y chromosomal DNA (60kb) | ||
| Line 55: | Line 65: | ||
[[File:Human Y chromosome 01.jpg|800px]] | [[File:Human Y chromosome 01.jpg|800px]] | ||
* [https://omim.org/entry/490000 ZFY] ({{ChrY}}p11.2) zinc finger transcription factor, required in males for spermatozoa or testis maturation.{{#pmid:20028140|PMID20028140}} A similar sequence called ZFX, has been identified on the {{ChrX}} chromosome, that in humans escapes [[X inactivation]]. | |||
:'''Links:''' [https://omim.org/entry/490000 ZFY] | [http://www.ncbi.nlm.nih.gov/omim/300162 OMIM - acetylserotonin O-methyltransferase-like] | [http://www.ncbi.nlm.nih.gov/omim/400029 OMIM - heat shock transcription factor, Y-linked 1] | [http://www.ncbi.nlm.nih.gov/omim/400038 OMIM - testis-specific transcript, Y-linked 5] | [http://www.ncbi.nlm.nih.gov/omim/300531 OMIM - sprouty homolog 3 (Drosophila)] | |||
[[File:Human Y chromosome.jpg|thumb|Human Y chromosome]] | [[File:Human Y chromosome.jpg|thumb|Human Y chromosome]] | ||
Sry acts as a transcriptional activator (HMG type-high mobility group) binding to DNA and initiating male sex determination then regulating male development. The protein sequence is shown on this current page and the full genebank entry can also be seen. The sry protein has a HMG box that binds DNA by intercalating in the minor groove. Read about the mapping of the testis determining factor which is SRY. | ==SRY== | ||
{{Sry}} (sex-determining region on the Y chromosome) gene was found in 1990 on the Y chromosome and the first Sox gene identified, the sry gene encodes a "testis-determining factor" a 204aa protein (Mr 23884 Da). | |||
{{Sry}} acts as a transcriptional activator (HMG type-high mobility group) binding to DNA and initiating male sex determination then regulating male development. The protein sequence is shown on this current page and the full genebank entry can also be seen. The sry protein has a HMG box that binds DNA by intercalating in the minor groove. Read about the mapping of the testis determining factor which is SRY. | |||
The actual gene targets of SRY are still being determined but at least one downstream gene Sox9 has been identified. Another gene Dax1 (nuclear hormone-receptor superfamily member) when expressed as a transgene will antagonize Sry and also force dosage-sensitive sex reversal. | The actual gene targets of SRY are still being determined but at least one downstream gene Sox9 has been identified. Another gene Dax1 (nuclear hormone-receptor superfamily member) when expressed as a transgene will antagonize Sry and also force dosage-sensitive sex reversal. | ||
[[File:Mouse sex determination genes 01.jpg|600px]] | |||
Mouse sex determination genes{{#pmid:22439850|PMID22439850}} | |||
Sox is an acronym of "'''S'''ry-related HMG b'''ox'''" and there have now been identified at least eight groups of genes that belong to this family with different functions. (for review see | |||
:'''Links:''' {{Sry}} | {{Male}} | [http://www.ncbi.nlm.nih.gov/omim/480000 OMIM - Sry] | [http://www.ncbi.nlm.nih.gov/Structure/mmdb/mmdbsrv.cgi?uid=49520 3d Structure Of The Human Sry-Dna Complex] | |||
===SOX=== | |||
Sox is an acronym of "'''S'''ry-related HMG b'''ox'''" and there have now been identified at least eight groups of genes that belong to this family with different functions. (for review see{{#pmid:17625949|PMID17625949}}) The "HMG box" is a region that functions for DNA binding, DNA bending, protein interactions, and nuclear import or export. | |||
The high mobility group (HMG) domain is a 79 amino acid protein region. | The high mobility group (HMG) domain is a 79 amino acid protein region. | ||
{{Sox family collapsetable}} | |||
===SRY Nuclear Import=== | ===SRY Nuclear Import=== | ||
| Line 76: | Line 102: | ||
[[File:SRY nuclear import model.jpg|600px]] | [[File:SRY nuclear import model.jpg|600px]] | ||
A model for nuclear import of | A model for nuclear import of {{Sry}} from normal males and XY females.{{#pmid:12764225|PMID12764225}} | ||
The distinct nuclear localization signals (NLSs) of SRY use different import pathways. | The distinct nuclear localization signals (NLSs) of SRY use different import pathways. | ||
| Line 87: | Line 112: | ||
===Sry Target Genes=== | ===Sry Target Genes=== | ||
'''Cerebellin 4 precursor''' (Cbln4) - encodes a secreted protein expressed in Sertoli cells in the developing gonad. | '''Cerebellin 4 precursor''' (Cbln4) - encodes a secreted protein expressed in Sertoli cells in the developing gonad.{{#pmid:19211811|PMID19211811}} | ||
'''Androgen receptor''' - SRY interacts with and negatively regulates this receptor transcriptional activity.{{#pmid:11585838|PMID11585838}} | |||
==DAZ== | |||
DELETED IN AZOOSPERMIA 1 (DAZ1) gene encodes an RNA-binding protein that has a role in spermatogenesis.{{#pmid:10857750|PMID10857750}} | |||
The DAZ cluster arose during primate evolution by: | |||
# transposition of the autosomal gene (on chromosome {{Chr3}) }to the Y | |||
# amplification and pruning of exons within the transposed gene | |||
# amplification of the modified gene | |||
''' | :'''Links:''' [https://www.omim.org/entry/400003 OMIM - DAZ1] | ||
==Testis-specific Protein Y Chromosome== | ==Testis-specific Protein Y Chromosome== | ||
[[File:Testis-specific_protein_on_Y_chromosome.jpg|thumb|Testis-specific protein on Y chromosome]] | [[File:Testis-specific_protein_on_Y_chromosome.jpg|thumb|Testis-specific protein on Y chromosome]] | ||
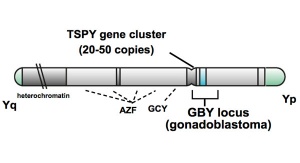
Testis-specific Protein Y Chromosome (TSPY) is an ampliconic gene on the Y chromosome with an unknown function, though the protein that has been shown to interact with gonadoblastoma. A recent study suggests that TSPY serves as a repressor in androgen-induced tumor development in testicular germ-cell tumours (TGCTs). | Testis-specific Protein Y Chromosome (TSPY) is an ampliconic gene on the Y chromosome with an unknown function, though the protein that has been shown to interact with gonadoblastoma. A recent study suggests that TSPY serves as a repressor in androgen-induced tumor development in testicular germ-cell tumours (TGCTs).{{#pmid:21041627|PMID21041627}} | ||
==Pseudoautosomal Regions== | ==Pseudoautosomal Regions== | ||
{| | {| | ||
| [[File:Sex_chromosomes_pseudoautosomal_regions.jpg|160px|Sex chromosomes pseudoautosomal regions.]] | | [[File:Sex_chromosomes_pseudoautosomal_regions.jpg|160px|Sex chromosomes pseudoautosomal regions.]] | ||
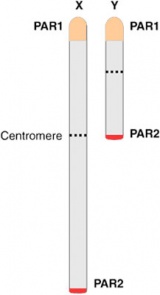
| These are two intervals of sequence identity at the tips of both the Y and X chromosomes, the human pseudoautosomal regions PAR1 and PAR2. | | These are two intervals of sequence identity at the tips of both the Y and X chromosomes, the human pseudoautosomal regions PAR1 and PAR2.{{#pmid:18398439|PMID18398439}} Loss of PAR1 has been associated with male sterility. | ||
|} | |} | ||
| Line 110: | Line 149: | ||
===Reviews=== | ===Reviews=== | ||
{{#pmid:20005972}} | |||
{{#pmid:19699459}} | |||
{{#pmid:18062864}} | |||
{{#pmid:17237341}} | |||
{{#pmid:15949865}} | |||
===Articles=== | ===Articles=== | ||
{{#pmid:21039604}} | |||
{{#pmid:2030730}} | |||
{{#pmid:1695712}} | |||
===Search PubMed=== | ===Search PubMed=== | ||
| Line 143: | Line 188: | ||
{{Footer}} | {{Footer}} | ||
[[Category:Male]] [[Category:Genital]] | [[Category:Male]][[Category:Genital]][[Category:Sry]] | ||
Latest revision as of 12:05, 27 April 2020
| Embryology - 21 May 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
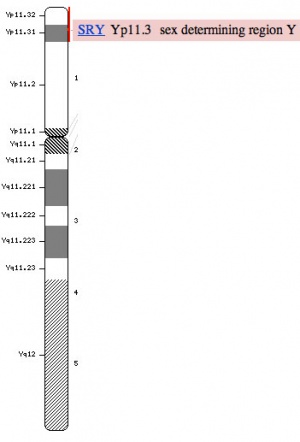
Introduction
Humans have 23 pairs of chromosomes, 22 autosomes and a pair of sex chromosomes. Males have one X and one Y chromosomes (46, XY) and females have a pair of X chromosomes (46, XX). The Y chromosome is much smaller than the X chromosome and contains 50 million base pairs encoding approximately 200+ genes. The Sry gene (location Yp11.3), found in 1990, encodes is responsible for male sex determination. SRY mutations lead to XY sex reversal in humans, and XX mice with an SRY transgene develop as fertile males. Interestingly, the laboratory rat, Rattus norvegicus, has at least 6 full length copies of the Sry gene.[1]
Placental male mammals have SRY protein transcription factor that activates Sox9 for testes formation. SRY evolved as a hybrid gene of Dgcr8 expressed in the developing placenta and Sox3 expressed in hypothalamic development.
SRY encodes a 204 amino acid protein that is a member of the HMG (High mobility group) box class of DNA-binding proteins. Transcription factors bind to specific sites of DNA and regulates the transcription (expression) of other genes.
The male-specific region of the human Y chromosome was originally called the non-recombining portion of the Y chromosome (NRY). This region consists of three different classes of euchromatic sequences:
- X-transposed - 3-4 Ma transposition 99% identical to human Xq21
- X-degenerate - 14 single copy or 13 pseudogene homologs of X-linked genes
- ampliconic sequences - Sequences with strong identity to other regions of Y
The Y and X chromosome are both transcriptionally silenced during spermatogenesis, at primary spermatocyte stage onward, by a process called meiotic sex chromosome inactivation (MSCI). (More? Meiosis)
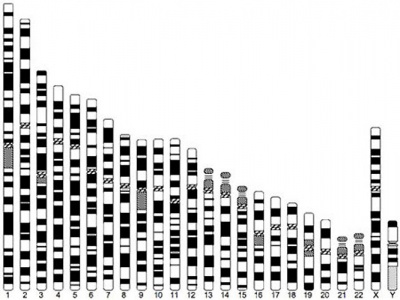
Human idiogram
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Y Chromosome | SRY | DAZ1 |
| Older papers |
|---|
| These papers originally appeared in the Some Recent Findings table, but as that list grew in length have now been shuffled down to this collapsible table.
See also the Discussion Page for other references listed by year and References on this current page.
|
History
- 1916 - Bridges[5][6] describes the sex chromosomes of Drosophila melanogaster and shows that sex is determined by the X:autosome ratio
- 1923 - Painter[7] describes the Sex chromosomes in mammals - it is assumed that sex determining mechanism will be the same as Drosophila
- 1959 - Ford et al.[8] find a human XO and female - Turner's syndrome
- 1959 - Jacobs and Strong[9] find an XXY male - Kleinfelter's syndrome
- together these prove that the Y chromosome determines sex and not the X:autosome ratio
- 1989 XX males found that carry a very small portion of Y chromosomal DNA (60kb)
- using DNA from these males a search for the testis determining gene, TDF or TDY led to the isolation of the gene
- gene called SRY because it comes from the Sex-determining Region of the Y chromosome. It is the molecular "switch" that determines sex in humans
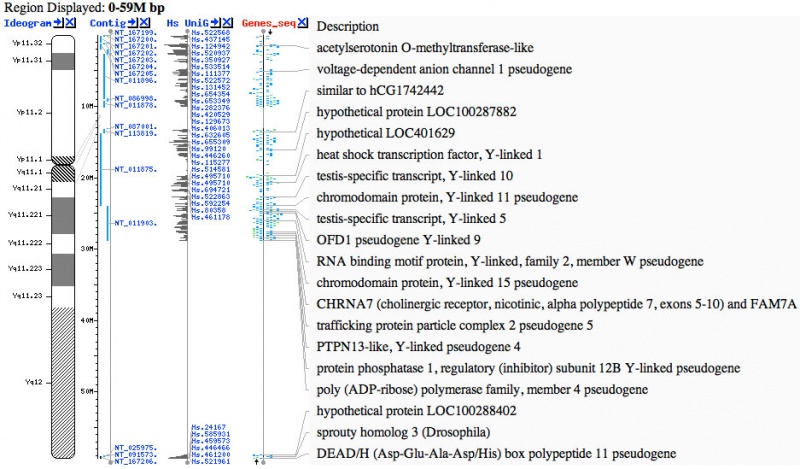
Y Chromosome Genes
- ZFY (Yp11.2) zinc finger transcription factor, required in males for spermatozoa or testis maturation.[10] A similar sequence called ZFX, has been identified on the X chromosome, that in humans escapes X inactivation.
- Links: ZFY | OMIM - acetylserotonin O-methyltransferase-like | OMIM - heat shock transcription factor, Y-linked 1 | OMIM - testis-specific transcript, Y-linked 5 | OMIM - sprouty homolog 3 (Drosophila)
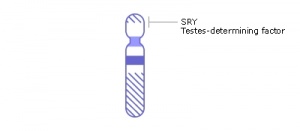
SRY
Sry (sex-determining region on the Y chromosome) gene was found in 1990 on the Y chromosome and the first Sox gene identified, the sry gene encodes a "testis-determining factor" a 204aa protein (Mr 23884 Da).
Sry acts as a transcriptional activator (HMG type-high mobility group) binding to DNA and initiating male sex determination then regulating male development. The protein sequence is shown on this current page and the full genebank entry can also be seen. The sry protein has a HMG box that binds DNA by intercalating in the minor groove. Read about the mapping of the testis determining factor which is SRY.
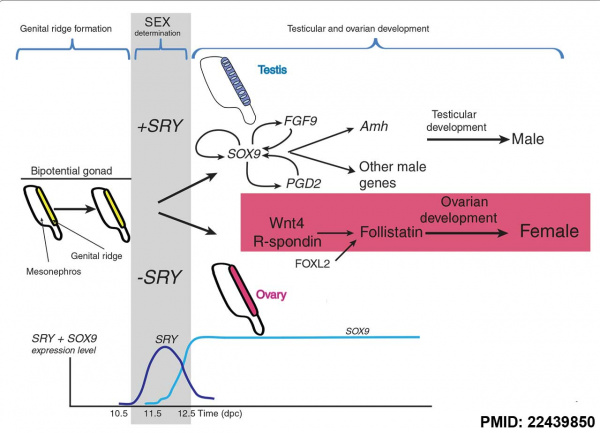
The actual gene targets of SRY are still being determined but at least one downstream gene Sox9 has been identified. Another gene Dax1 (nuclear hormone-receptor superfamily member) when expressed as a transgene will antagonize Sry and also force dosage-sensitive sex reversal.
Mouse sex determination genes[11]
- Links: Sry | Male | OMIM - Sry | 3d Structure Of The Human Sry-Dna Complex
SOX
Sox is an acronym of "Sry-related HMG box" and there have now been identified at least eight groups of genes that belong to this family with different functions. (for review see[12]) The "HMG box" is a region that functions for DNA binding, DNA bending, protein interactions, and nuclear import or export.
The high mobility group (HMG) domain is a 79 amino acid protein region.
| Human SOX Family | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
SRY Nuclear Import
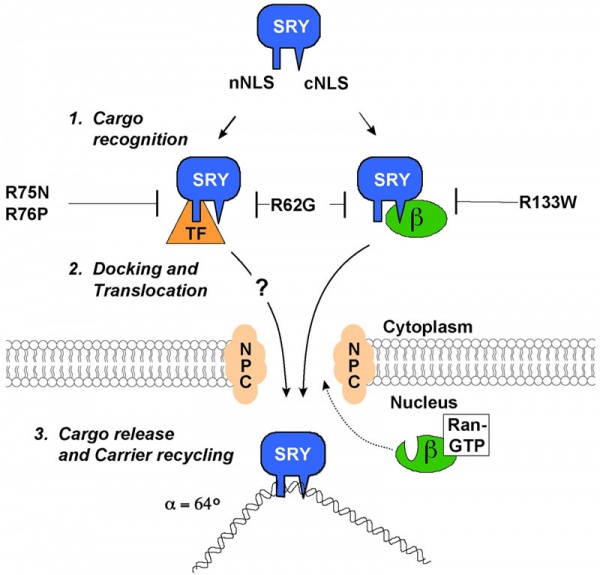
A model for nuclear import of Sry from normal males and XY females.[13]
The distinct nuclear localization signals (NLSs) of SRY use different import pathways.
- cNLS - recognized by IMPβ, docks the transport complex at the nuclear pore complex (NPC) and is then translocation through. After nuclear entry of the complex, RanGTP binds to IMPβ to trigger release of SRY; DNA binding by SRY may also facilitate the release process.
- nNLS - mediates nuclear import through a novel pathway not utilizing conventional nuclear import factors such as IMPs but an unidentified "transport factor" (TF) suggested to be calcium-calmodulin.
Sry Target Genes
Cerebellin 4 precursor (Cbln4) - encodes a secreted protein expressed in Sertoli cells in the developing gonad.[14]
Androgen receptor - SRY interacts with and negatively regulates this receptor transcriptional activity.[15]
DAZ
DELETED IN AZOOSPERMIA 1 (DAZ1) gene encodes an RNA-binding protein that has a role in spermatogenesis.[16]
The DAZ cluster arose during primate evolution by:
- transposition of the autosomal gene (on chromosome {{Chr3}) }to the Y
- amplification and pruning of exons within the transposed gene
- amplification of the modified gene
- Links: OMIM - DAZ1
Testis-specific Protein Y Chromosome
Testis-specific Protein Y Chromosome (TSPY) is an ampliconic gene on the Y chromosome with an unknown function, though the protein that has been shown to interact with gonadoblastoma. A recent study suggests that TSPY serves as a repressor in androgen-induced tumor development in testicular germ-cell tumours (TGCTs).[17]
Pseudoautosomal Regions
|
These are two intervals of sequence identity at the tips of both the Y and X chromosomes, the human pseudoautosomal regions PAR1 and PAR2.[18] Loss of PAR1 has been associated with male sterility. |
References
- ↑ Turner ME, Martin C, Martins AS, Dunmire J, Farkas J, Ely DL & Milsted A. (2007). Genomic and expression analysis of multiple Sry loci from a single Rattus norvegicus Y chromosome. BMC Genet. , 8, 11. PMID: 17408480 DOI.
- ↑ Sakashita A, Wakai T, Kawabata Y, Nishimura C, Sotomaru Y, Alavattam KG, Namekawa SH & Kono T. (2019). XY oocytes of sex-reversed females with a Sry mutation deviate from the normal developmental process beyond the mitotic stage†. Biol. Reprod. , 100, 697-710. PMID: 30289439 DOI.
- ↑ Ortega EA, Salvador Q, Fernandez M & Ward MA. (2018). Alterations of sex determination pathway in the genital ridges of males with limited Y chromosome genes. Biol. Reprod. , , . PMID: 30285093 DOI.
- ↑ Kaur G & Jans DA. (2011). Dual nuclear import mechanisms of sex determining factor SRY: intracellular Ca2+ as a switch. FASEB J. , 25, 665-75. PMID: 21051653 DOI.
- ↑ Bridges CB. (1916). Non-Disjunction as Proof of the Chromosome Theory of Heredity. Genetics , 1, 1-52. PMID: 17245850
- ↑ Bridges CB. (1916). Non-Disjunction as Proof of the Chromosome Theory of Heredity (Concluded). Genetics , 1, 107-63. PMID: 17245853
- ↑ Painter TS. (1923). FURTHER OBSERVATIONS ON THE SEX CHROMOSOMES OF MAMMALS. Science , 58, 247-8. PMID: 17748214 DOI.
- ↑ FORD CE, JONES KW, POLANI PE, DE ALMEIDA JC & BRIGGS JH. (1959). A sex-chromosome anomaly in a case of gonadal dysgenesis (Turner's syndrome). Lancet , 1, 711-3. PMID: 13642858
- ↑ JACOBS PA & STRONG JA. (1959). A case of human intersexuality having a possible XXY sex-determining mechanism. Nature , 183, 302-3. PMID: 13632697
- ↑ Grants J, Flanagan E, Yee A & Romaniuk PJ. (2010). Characterization of the DNA binding activity of the ZFY zinc finger domain. Biochemistry , 49, 679-86. PMID: 20028140 DOI.
- ↑ de Lau WB, Snel B & Clevers HC. (2012). The R-spondin protein family. Genome Biol. , 13, 242. PMID: 22439850 DOI.
- ↑ Lefebvre V, Dumitriu B, Penzo-Méndez A, Han Y & Pallavi B. (2007). Control of cell fate and differentiation by Sry-related high-mobility-group box (Sox) transcription factors. Int. J. Biochem. Cell Biol. , 39, 2195-214. PMID: 17625949 DOI.
- ↑ Harley VR, Layfield S, Mitchell CL, Forwood JK, John AP, Briggs LJ, McDowall SG & Jans DA. (2003). Defective importin beta recognition and nuclear import of the sex-determining factor SRY are associated with XY sex-reversing mutations. Proc. Natl. Acad. Sci. U.S.A. , 100, 7045-50. PMID: 12764225 DOI.
- ↑ Bradford ST, Hiramatsu R, Maddugoda MP, Bernard P, Chaboissier MC, Sinclair A, Schedl A, Harley V, Kanai Y, Koopman P & Wilhelm D. (2009). The cerebellin 4 precursor gene is a direct target of SRY and SOX9 in mice. Biol. Reprod. , 80, 1178-88. PMID: 19211811 DOI.
- ↑ Yuan X, Lu ML, Li T & Balk SP. (2001). SRY interacts with and negatively regulates androgen receptor transcriptional activity. J. Biol. Chem. , 276, 46647-54. PMID: 11585838 DOI.
- ↑ Tsui S, Dai T, Roettger S, Schempp W, Salido EC & Yen PH. (2000). Identification of two novel proteins that interact with germ-cell-specific RNA-binding proteins DAZ and DAZL1. Genomics , 65, 266-73. PMID: 10857750 DOI.
- ↑ Akimoto C, Ueda T, Inoue K, Yamaoka I, Sakari M, Obara W, Fujioka T, Nagahara A, Nonomura N, Tsutsumi S, Aburatani H, Miki T, Matsumoto T, Kitagawa H & Kato S. (2010). Testis-specific protein on Y chromosome (TSPY) represses the activity of the androgen receptor in androgen-dependent testicular germ-cell tumors. Proc. Natl. Acad. Sci. U.S.A. , 107, 19891-6. PMID: 21041627 DOI.
- ↑ Flaquer A, Rappold GA, Wienker TF & Fischer C. (2008). The human pseudoautosomal regions: a review for genetic epidemiologists. Eur. J. Hum. Genet. , 16, 771-9. PMID: 18398439 DOI.
Online Textbooks
Search NLM Online Textbooks "Y chromosome" : Developmental Biology | Endocrinology | Molecular Biology of the Cell | The Cell- A molecular Approach
Reviews
Sekido R. (2010). SRY: A transcriptional activator of mammalian testis determination. Int. J. Biochem. Cell Biol. , 42, 417-20. PMID: 20005972 DOI.
Bianchi NO. (2009). Y chromosome structural and functional changes in human malignant diseases. Mutat. Res. , 682, 21-7. PMID: 19699459 DOI.
Osborne EC, Lynch M, McLachlan R, Trounson AO & Cram DS. (2007). Microarray detection of Y chromosome deletions associated with male infertility. Reprod. Biomed. Online , 15, 673-80. PMID: 18062864
Wilhelm D, Palmer S & Koopman P. (2007). Sex determination and gonadal development in mammals. Physiol. Rev. , 87, 1-28. PMID: 17237341 DOI.
Koopman P. (2005). Sex determination: a tale of two Sox genes. Trends Genet. , 21, 367-70. PMID: 15949865 DOI.
Articles
Antonelli A, Marcucci L, Elli R, Tanzi N, Paoli D, Radicioni A, Lombardo F, Lenzi A & Gandini L. (2011). Semen quality in men with Y chromosome aberrations. Int. J. Androl. , 34, 453-60. PMID: 21039604 DOI.
Koopman P, Gubbay J, Vivian N, Goodfellow P & Lovell-Badge R. (1991). Male development of chromosomally female mice transgenic for Sry. Nature , 351, 117-21. PMID: 2030730 DOI.
Sinclair AH, Berta P, Palmer MS, Hawkins JR, Griffiths BL, Smith MJ, Foster JW, Frischauf AM, Lovell-Badge R & Goodfellow PN. (1990). A gene from the human sex-determining region encodes a protein with homology to a conserved DNA-binding motif. Nature , 346, 240-4. PMID: 1695712 DOI.
Search PubMed
Search November 2010 "Y Chromosome" - All (11590) Review (1052) Free Full Text (2688)
Search Pubmed: Y Chromosome | SRY
External Links
External Links Notice - The dynamic nature of the internet may mean that some of these listed links may no longer function. If the link no longer works search the web with the link text or name. Links to any external commercial sites are provided for information purposes only and should never be considered an endorsement. UNSW Embryology is provided as an educational resource with no clinical information or commercial affiliation.
- Ensembl Y Chromosome
- Genome View Y chromosome
- OMIM SRY
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, May 21) Embryology Y Chromosome. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Y_Chromosome
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G