Medaka Development: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 18: | Line 18: | ||
| | | | ||
* '''foxl3 is a germ cell-intrinsic factor involved in sperm-egg fate decision in medaka'''<ref name="PMID26067255"><pubmed>26067255</pubmed>| [http://www.sciencemag.org/content/early/2015/06/10/science.aaa2657.abstract Science.]</ref> "Sex determination is an essential step in the commitment of a germ cell to a sperm or egg. However, the intrinsic factors that determine the sexual fate of vertebrate germ cells are unknown. Here we show that foxl3, which is expressed in germ cells but not somatic cells in the gonad, is involved in sperm-egg fate decision in medaka fish. Adult XX medaka with disrupted foxl3 developed functional sperm in the expanded germinal epithelium of a histologically functional ovary. In chimeric medaka, mutant germ cells initiated spermatogenesis in female wild-type gonad. These results indicate that a germ cell-intrinsic cue for the sperm-egg fate decision is present in medaka and that spermatogenesis can proceed in a female gonadal environment." ( Forkhead Transcription Factor foxl3 is an ancient duplicated copy of [http://omim.org/entry/605597?search=foxl2&highlight=foxl2 foxl2] | [[Oocyte Development]] | [[Spermatozoa Development]]) | * '''foxl3 is a germ cell-intrinsic factor involved in sperm-egg fate decision in medaka'''<ref name="PMID26067255"><pubmed>26067255</pubmed>| [http://www.sciencemag.org/content/early/2015/06/10/science.aaa2657.abstract Science.]</ref> "Sex determination is an essential step in the commitment of a germ cell to a sperm or egg. However, the intrinsic factors that determine the sexual fate of vertebrate germ cells are unknown. Here we show that foxl3, which is expressed in germ cells but not somatic cells in the gonad, is involved in sperm-egg fate decision in medaka fish. Adult XX medaka with disrupted foxl3 developed functional sperm in the expanded germinal epithelium of a histologically functional ovary. In chimeric medaka, mutant germ cells initiated spermatogenesis in female wild-type gonad. These results indicate that a germ cell-intrinsic cue for the sperm-egg fate decision is present in medaka and that spermatogenesis can proceed in a female gonadal environment." ( Forkhead Transcription Factor foxl3 is an ancient duplicated copy of [http://omim.org/entry/605597?search=foxl2&highlight=foxl2 foxl2] | [[Oocyte Development]] | [[Spermatozoa Development]]) | ||
* '''YAP is essential for tissue tension to ensure vertebrate 3D body shape'''<ref name="PMID25778702"><pubmed>25778702</pubmed></ref> "Vertebrates have a unique 3D body shape in which correct tissue and organ shape and alignment are essential for function. For example, vision requires the lens to be centred in the eye cup which must in turn be correctly positioned in the head. Tissue morphogenesis depends on force generation, force transmission through the tissue, and response of tissues and extracellular matrix to force. Although a century ago D'Arcy Thompson postulated that terrestrial animal body shapes are conditioned by gravity, there has been no animal model directly demonstrating how the aforementioned mechano-morphogenetic processes are coordinated to generate a body shape that withstands gravity. Here we report a unique medaka fish (Oryzias latipes) mutant, hirame (hir), which is sensitive to deformation by gravity. hir embryos display a markedly flattened body caused by mutation of YAP, a nuclear executor of Hippo signalling that regulates organ size. We show that actomyosin-mediated tissue tension is reduced in hir embryos, leading to tissue flattening and tissue misalignment, both of which contribute to body flattening. By analysing YAP function in 3D spheroids of human cells, we identify the Rho GTPase activating protein ARHGAP18 as an effector of YAP in controlling tissue tension. Together, these findings reveal a previously unrecognised function of YAP in regulating tissue shape and alignment required for proper 3D body shape." | |||
* '''The Effects of Rearing Density, Salt Concentration, and Incubation Temperature on Japanese Medaka (Oryzias latipes) Embryo Development'''<ref name="PMID23244689"><pubmed>23244689</pubmed></ref> "When testing variations in temperature (24°, 28°, and 32°C) and salinity (0.3, 10, 15, and 20 ppt), the onset of heartbeat and onset of retina pigmentation were observed. The original hypotheses were not all supported: as rearing density increased, success of hatch decreased; as salinity increased, only the rate of development for heartbeat increased; as temperature increased, the rate of development for both onset of the heartbeat and retina pigmentation also increased." | * '''The Effects of Rearing Density, Salt Concentration, and Incubation Temperature on Japanese Medaka (Oryzias latipes) Embryo Development'''<ref name="PMID23244689"><pubmed>23244689</pubmed></ref> "When testing variations in temperature (24°, 28°, and 32°C) and salinity (0.3, 10, 15, and 20 ppt), the onset of heartbeat and onset of retina pigmentation were observed. The original hypotheses were not all supported: as rearing density increased, success of hatch decreased; as salinity increased, only the rate of development for heartbeat increased; as temperature increased, the rate of development for both onset of the heartbeat and retina pigmentation also increased." | ||
*''' Osterix-mCherry transgenic medaka for in vivo imaging of bone formation'''<ref name="PMID19097055"><pubmed>19097055</pubmed></ref> | *''' Osterix-mCherry transgenic medaka for in vivo imaging of bone formation'''<ref name="PMID19097055"><pubmed>19097055</pubmed></ref> | ||
Revision as of 11:12, 22 July 2015
| Embryology - 20 May 2024 |
|---|
| Google Translate - select your language from the list shown below (this will open a new external page) |
|
العربية | català | 中文 | 中國傳統的 | français | Deutsche | עִברִית | हिंदी | bahasa Indonesia | italiano | 日本語 | 한국어 | မြန်မာ | Pilipino | Polskie | português | ਪੰਜਾਬੀ ਦੇ | Română | русский | Español | Swahili | Svensk | ไทย | Türkçe | اردو | ייִדיש | Tiếng Việt These external translations are automated and may not be accurate. (More? About Translations) |
Introduction
Medaka Oryzias latipes or Japanese rice fish is a member of the killifish family first described in 1846 and has been widely used as a aquarium fish. A modified aquarium version with a genetically modified fluorescent (GFP) version also now available in some countries.
A 2004 study by Iwamatsu[1] has characterised the stages of normal fish development.
Medaka fish were also the first for the first vertebrate animal to mate in space (The International Microgravity Laboratory IML-2/STS-65 mission in 1994) as a developmental model for space experiments. The fish has also been used in studies of pigmentation development.
| Fish Links: Zebrafish Development | Medaka Development | Salmon Development | Movie - Zebrafish Heart | Student Group Project - Zebrafish | Recent References | Category:Zebrafish | Category:Medaka |
Some Recent Findings
|
| More recent papers |
|---|
|
This table allows an automated computer search of the external PubMed database using the listed "Search term" text link.
More? References | Discussion Page | Journal Searches | 2019 References | 2020 References Search term: Medaka Embryology <pubmed limit=5>Medaka Embryology</pubmed> |
Taxon
cellular organisms; Eukaryota; Fungi/Metazoa group; Metazoa; Eumetazoa; Bilateria; Coelomata; Deuterostomia; Chordata; Craniata; Vertebrata; Gnathostomata; Teleostomi; Euteleostomi; Actinopterygii; Actinopteri; Neopterygii; Teleostei; Elopocephala; Clupeocephala; Euteleostei; Neognathi; Neoteleostei; Eurypterygii; Ctenosquamata; Acanthomorpha; Euacanthomorpha; Holacanthopterygii; Acanthopterygii; Euacanthopterygii; Percomorpha; Smegmamorpha; Atherinomorpha; Beloniformes; Adrianichthyoidei; Adrianichthyidae; Oryziinae; Oryzias
Development Overview
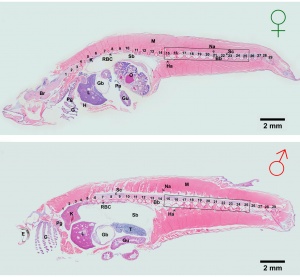
Development has been characterised by light microscope observation into 39 prehatch stages and 6 posthatch stages.[1]
Prehatch features observed included: number and size of blastomeres, form of the blastoderm, extent of epiboly, central nervous system, number and form of somites, optic and otic, notochord, heart, blood circulation, the size and movement of the body, tail, membranous fin (fin fold), viscera (liver gallbladder, gut tube), spleen and swim (air) bladder.
Posthatch features observed included: fins, scales and secondary sexual characteristics.
Developmental Stages
| Stage | Time | Event |
| 0 | Unfertilized eggs | |
| 1 | 3 min | Activated egg |
| 2 | Blastodisc | |
| 3 | 1 h 5 min | 2 cell |
| 4 | 1 h 45 min | 4 cell |
| 5 | 2 h 20 min | 8 cell |
| 6 | 2 h 55 min | 16 cell |
| 7 | 3 h 30 min | 32 cell |
| 8 | 4 h 5 min | Early morula |
| 9 | 5 h 15 min | Late morula |
| 10 | 6 h 30 min | Early blastula |
| 11 | 8 h 15 min | Late blastula |
| 12 | 10 h 20 min | Pre-early gastrula |
| 13 | 13 h | Early gastrula |
| 14 | 15 h | Pre-mid-gastrula |
| 15 | 17 h 30 min | Mid-gastrula |
| 16 | 21 h | Late gastrula |
| 17 | 1 day 1 h | Early neurula (head formation) |
| 18 | 1 day 2 h | Late neurula (optic bud formation) |
| 19 | 1 day 3 h 30 min | 2 somite |
| 20 | 1 day 7 h 30 min | 4 somite |
| 21 | 1 day 10 h | 6 somite (brain regionalization and otic vesicle formation) |
| 22 | 1 day 14 h | 9 somite (appearance of heart anlage) |
| 23 | 1 day 17 h | 12 somite (formation of tubular heart) |
| 24 | 1 day 20 h | 16 somite (start of heart beating) |
| 25 | 2 days 2 h | 18–19 somite (onset of blood circulation) |
| 26 | 2 days 6 h | 22 somite (development of guanophores and vacuolization of the notochord) |
| 27 | 2 days 10 h | 24 somite (appearance of pectoral fin bud) |
| 28 | 2 days 16 h | 30 somite (onset of retinal pigmentation) |
| 29 | 3 days 2 h | 34 somite (internal ear formation) |
| 30 | 3 days 10 h | 35 somite (blood vessel development) |
| 31 | 3 days 23 h | Gill blood vessel formation |
| 32 | 4 days 5 h | Somite completion (formation of pronephros and air bladder) |
| 33 | 4 days 10 h | at which notochord vacuolization is completed |
| 34 | 5 days 1 h | Pectoral fin blood circulation |
| 35 | 5 days 12 h | at which visceral blood vessels form |
| 36 | 6 days | Heart development |
| 37 | 7 days | Pericardial cavity formation |
| 38 | 8 days | Spleen development (differentiation of caudal fin begins) |
| 39 | 9 days | Hatching |
| 40 | 1st fry | |
| 41 | ||
| 42 | ||
| 43 | ||
| 44 | ||
| 45 |
National BioResource Project Medaka
The National BioResource Project Medaka (NBRP Medaka): an integrated bioresource for biological and biomedical sciences[8]"The Japanese government has supported the development of Medaka Bioresources since 2002. The second term of the Medaka Bioresource Project started in 2007. The National Institute for Basic Biology and Niigata University were selected as the core organizations for this project. More than 400 strains including more than 300 spontaneous and induced mutants, 8 inbred lines, 21 transgenic lines, 20 medaka-related species and 66 wild stock lines of medaka are now being provided to the scientific community and educational non-profit organizations. In addition to these live fish, NBRP Medaka is also able to provide cDNA/EST clones such as full-length cDNA and BAC/fosmid clones covering 90% of the medaka genome."
References
Articles
<pubmed>21094700</pubmed> <pubmed>15210169</pubmed>
Search Pubmed
Search Pubmed: Medaka Development
External Links
- National BioResource Project Medaka (NBRP Medaka) http://shigen.lab.nig.ac.jp/medaka/
Animal Development
| Animal Development: axolotl | bat | cat | chicken | cow | dog | dolphin | echidna | fly | frog | goat | grasshopper | guinea pig | hamster | horse | kangaroo | koala | lizard | medaka | mouse | opossum | pig | platypus | rabbit | rat | salamander | sea squirt | sea urchin | sheep | worm | zebrafish | life cycles | development timetable | development models | K12 |
Glossary Links
- Glossary: A | B | C | D | E | F | G | H | I | J | K | L | M | N | O | P | Q | R | S | T | U | V | W | X | Y | Z | Numbers | Symbols | Term Link
Cite this page: Hill, M.A. (2024, May 20) Embryology Medaka Development. Retrieved from https://embryology.med.unsw.edu.au/embryology/index.php/Medaka_Development
- © Dr Mark Hill 2024, UNSW Embryology ISBN: 978 0 7334 2609 4 - UNSW CRICOS Provider Code No. 00098G

